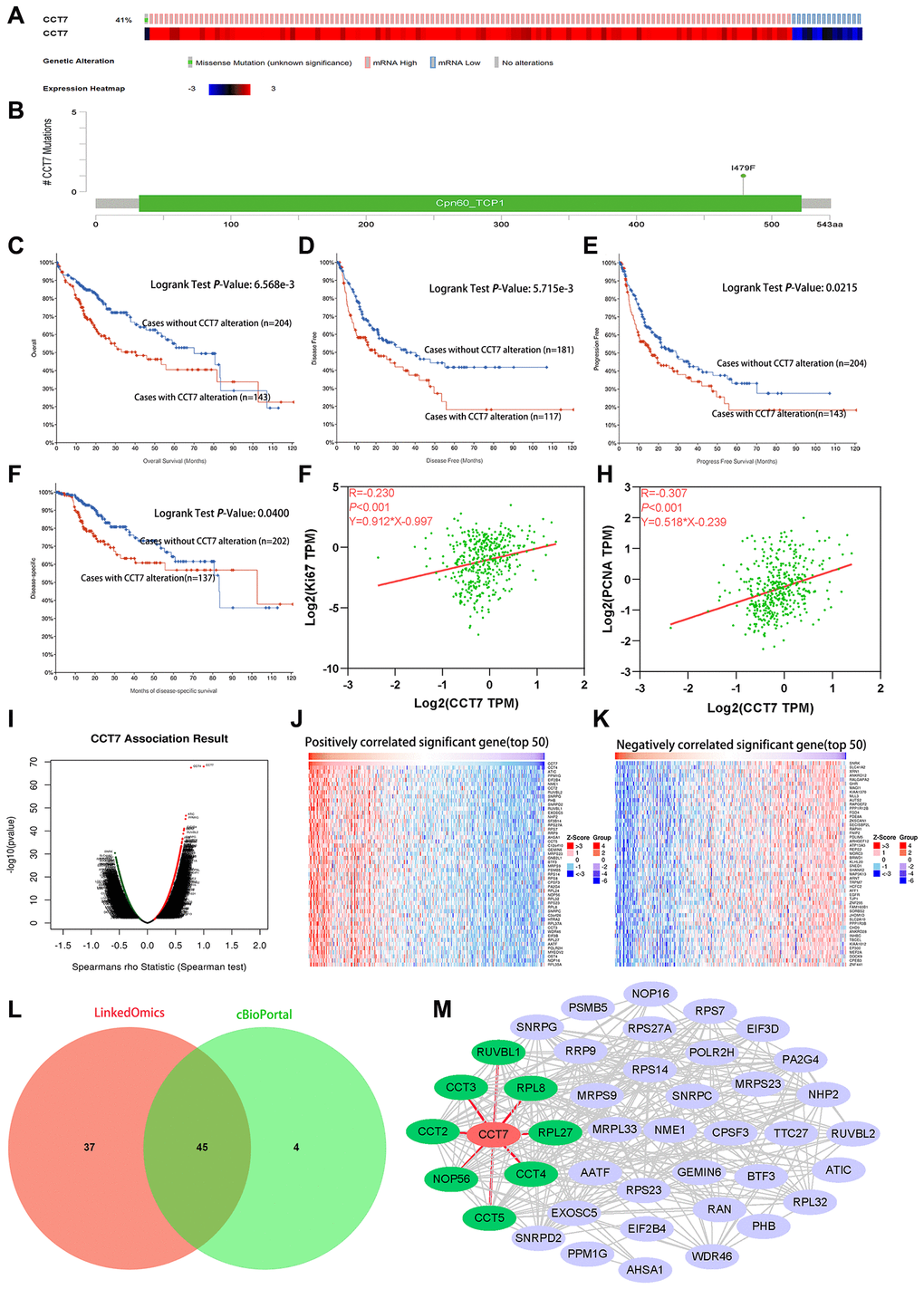
Figure 7.Genetic alterations of CCT7 are associated with poorer survival in HCC patients. (A) CCT7 was genetically altered in 143 patients (41%) from a cohort of 348 HCC patients. (B) A mutational hotspot of 1479F/Missense was found in 104 patients. (C–F) Survival analyses showing that HCC patients with CCT7 alterations had poorer OS (C), disease-free survival (D), progression-free survival (E) and disease-specific survival (F) than those without. (G, H) Spearman’s correlation analysis revealing that CCT7 mRNA expression correlated positively with Ki67 (G) and PCNA (H) expression. (I–K) CCT7 expression-associated target gene analysis in the LinkedOmics database. (I) Volcano chart exhibiting genes with significant positive/negative correlations with CCT7 expression. (J) Top 50 genes that were positively associated with CCT7 expression. (K) Top 50 genes that were negatively associated with CCT7 expression. (L) Venn plot showing the overlapping genes from the LinkedOmics and cBioPortal databases with Spearman’s values greater than 0.55. (M) PPI network for 45 genes co-expressed with CCT7. CCT2, CCT3, CCT4, CCT5, NOP56, RPL8, RPL27 and RUVBL1 were found to interact with CCT7.