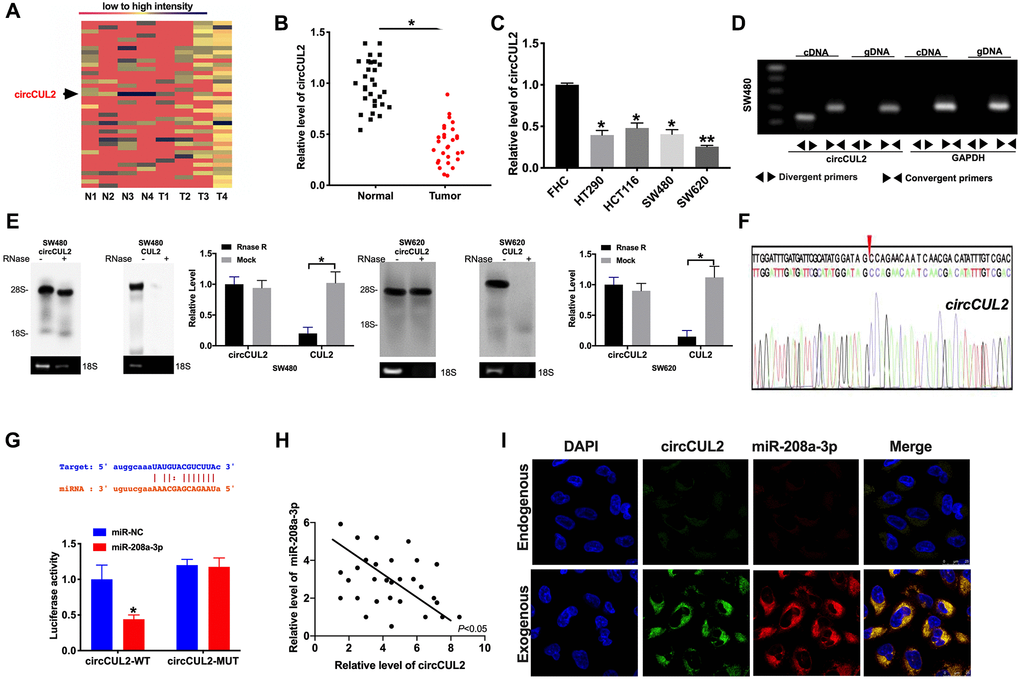
Figure 1.circCUL2 is downregulated in CRC tissues and cells. (A) The chip assay was performed in CRC tumor tissues and paired adjacent normal tissues. (n = 4). (B) The expression level of circCUL2 was detected in tumor tissues and paired adjacent normal tissues. n = 30. *P < 0.05. (C) The expression level of circCUL2 in the CRC cell line, FHC cell lines was indicated as control. n = 5. *P < 0.05, **P < 0.01. (D) The PCR products of circCUL2 and linear CUL2 were tested by gel electrophoresis. Divergent primers amplified circCUL2 in cDNA but not genomic DNA (gDNA). Convergent primers amplified linear CUL2 in both cDNA and gDNA. GAPDH was used as a linear control. (E) CircCUL2 was resistant to RNaseR digestion in SW480 and SW620 CRC cells. n = 3. *P < 0.05. (F) The result of Sanger sequencing. (G) The binding sites between circCUL2 and miR-208a-3p (upper), luciferase assay report verified the relationship between circCUL2 and miR-208a-3p (lower). n = 4. *P < 0.05. (H) The correlation analysis between circCUL2 and miR-208a-3p in 30 paired CRC tissues. n = 30. *P < 0.05. (I) Co-localization between circCUL2 and miR-208a-3p was revealed by fluorescence in situ hybridization.