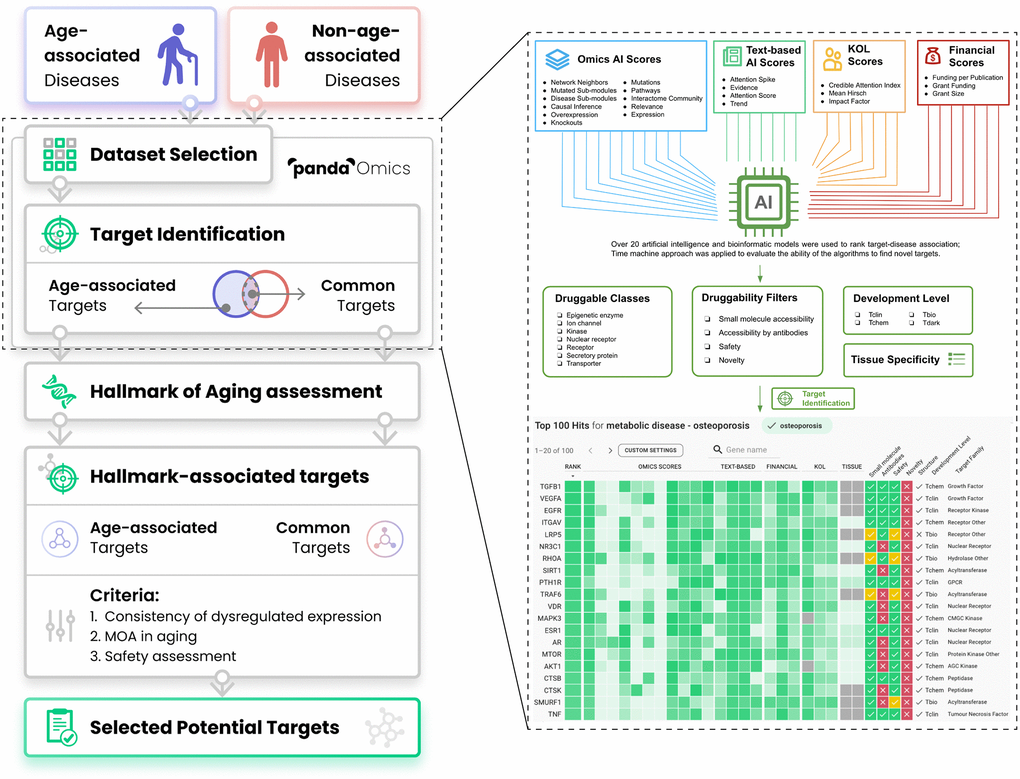
Figure 1.Workflow of the present study. Thirty-three diseases were separated into either age-associated diseases (AADs) or non-age-associated diseases (NAADs) based on the impact of age on the risk of the disease’s onset. Their corresponding transcriptomic datasets were retrieved from public repositories and processed by PandaOmics. Age bias between case and control groups has been considered during dataset selection. With multiple levels of novelty settings, targets implicated in AADs and NAADs were identified by ‘PandaOmics - target identification’. PandaOmics prioritized targets for one disease and refined the targets based on several flexible druggability filters. The target-disease associations were ranked according to over 20 artificial intelligence and bioinformatics models ranging from Omics AI scores, Text-based AI scores, Finance scores to KOL scores. Target identification was performed independently for each disease. Top-ranked targets shared by both disease categories were regarded as common targets, while targets unique to AADs were defined as age-associated targets (AAD targets). All common targets and AAD targets were subjected to the hallmarks of aging assessment by searching the literature for their evidence in modulating longevity or longevity pathways. To propose potential targets with a dual role in anti-aging and disease treatment, hallmark-associated targets were further evaluated based on their expression profiles across AADs, mechanism of action, and safety. A total of 9 targets were selected, with three levels of novelty. Abbreviation: KOL: Key opinion leader.