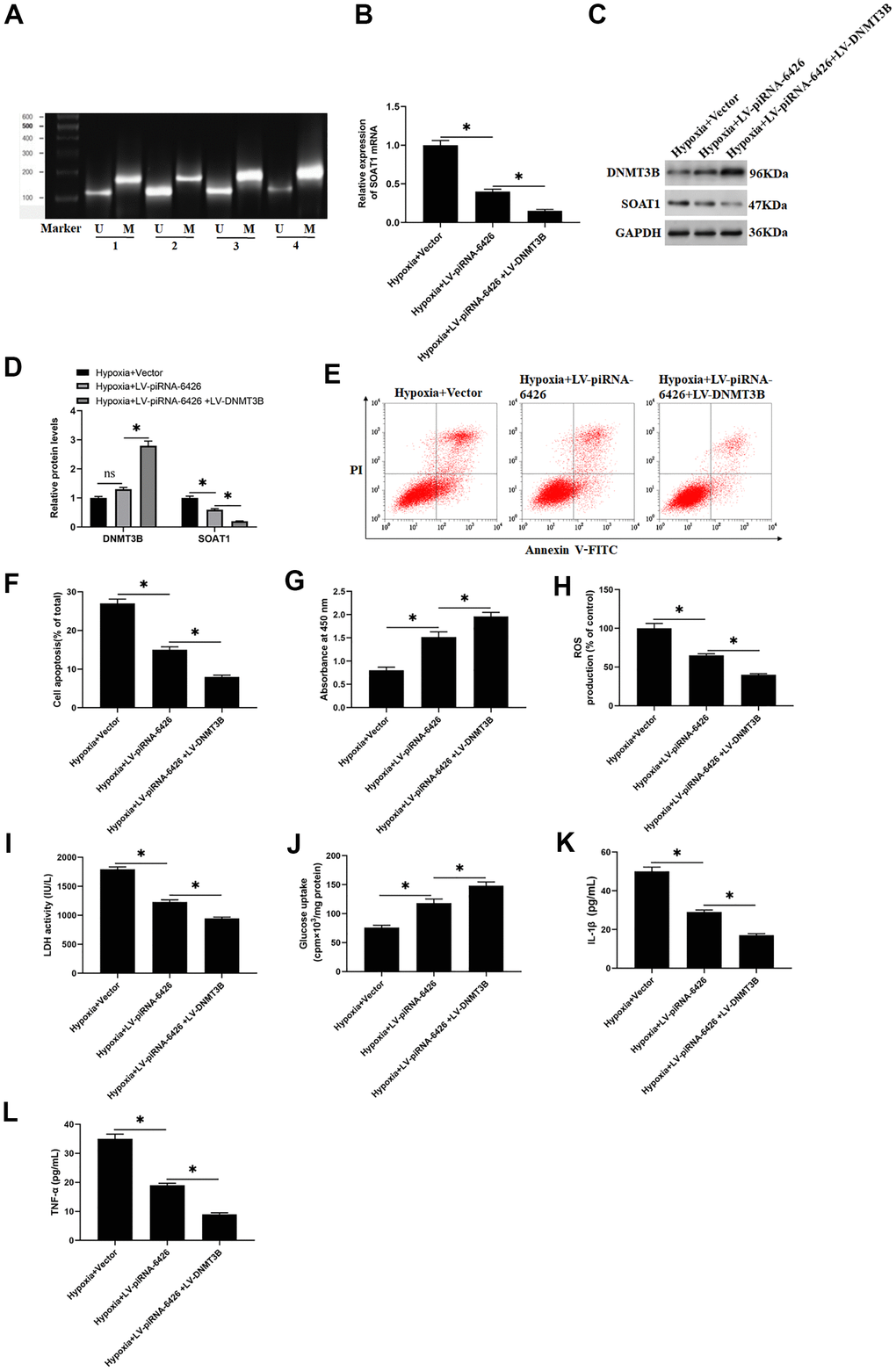
Figure 6.High methylation level of SOAT1 promoter alleviates hypoxia-induced dysfunction of rat cardiomyocytes. The cells were induced in a hypoxic incubator for 24 hours to establish a HF cell model after 48 h of per-incubation with LV-piRNA-6426 or together with LV-DNMT3B. (A) MSP assay was used to detect the methylation level of SOAT1 promoter. (B) RT-qPCR assay was used to detect the expression level of SOAT1 mRNA. (C, D) Western blotting was used to detect the expression levels of SOAT1 and DNMT3B proteins. (E, F) Flow cytometry was used to detect cell apoptosis. (G) MTT assay was used to identify cell viability. (H) The production of ROS was analyzed with DCFH-DA. (I) The LDH activity was detected with an ELISA kit. (J) D-(2-3H)-glucose uptake assay was used to perform glucose uptake on fully fused rat cardiomyocytes. (K, L) ELISA kits were used to detect the secretion of IL-1β and TNF-α. Values were expressed as mean ± SEM. ns P>0.05, *P<0.05, **P<0.01, n=6.