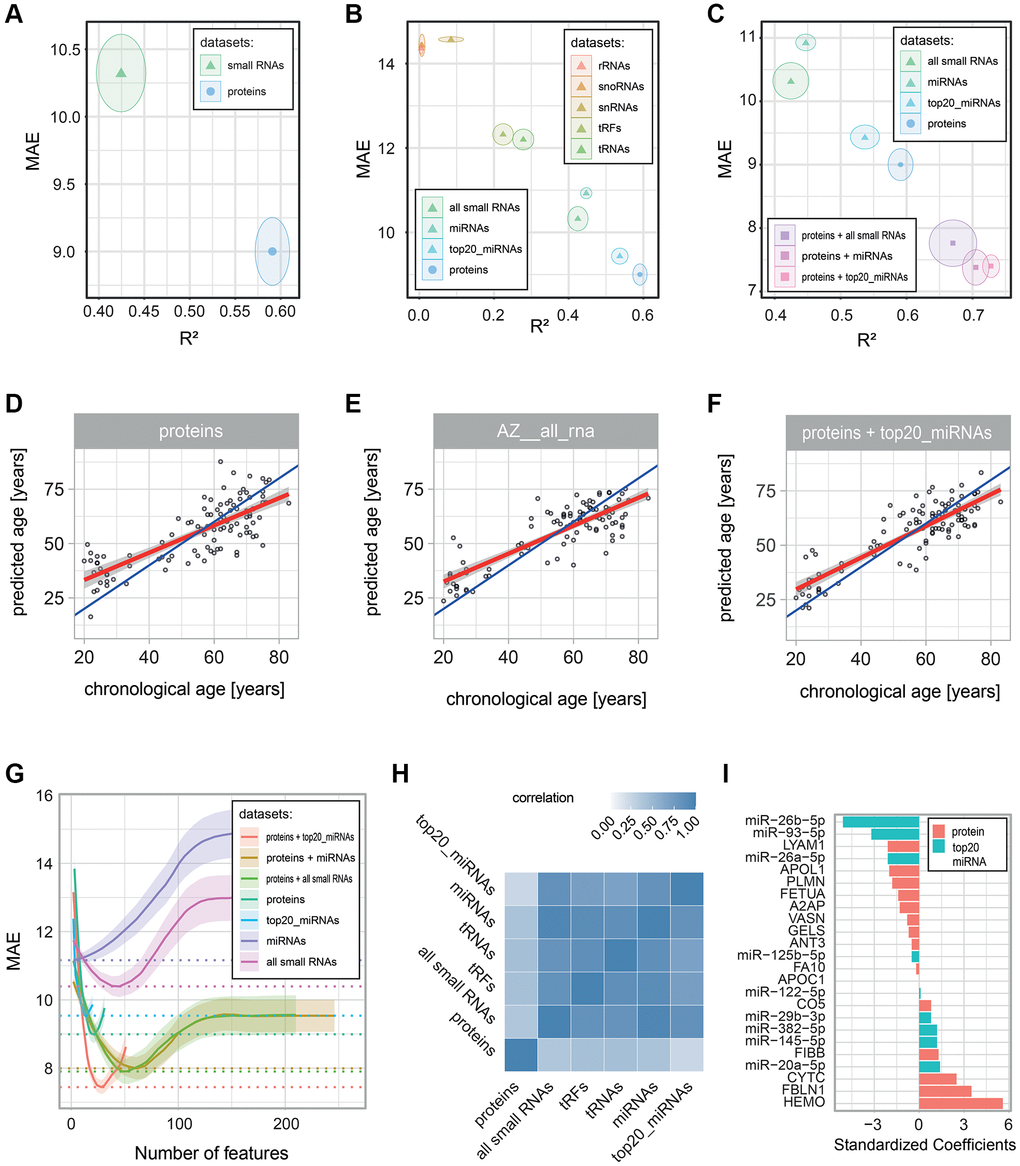
Figure 4.Performance of age-predictive models built on various data types. Age-predictive L1-norm penalized generalized linear models were built using protein and small RNA measurements, either separately or in combinations. Performance was estimated via 10-fold cross-validation with 100 repeats. Prediction errors were determined from predictions based on left-out data (data that was not used to build the model). (A–C) Performance of the built models: the mean (dot) and standard deviation (circle) of two error metrics are shown: the coefficient of determination (R2) on the x-axis and the Mean Absolute Error (MAE) on the y-axis. The panels compare (A) all small RNAs with all proteins, (B) the different classes of small RNAs, and (C) models combining proteins and small RNAs. (D–F) Scatter plots of chronological age vs. predicted age are shown for all individuals in the cohort for (D) the proteomics-based model, (E) the all small RNA-based model, and (F) the proteomics and top 20_miRNA-based model. Blue and red lines show, respectively, the identity and linear regression lines. (G) Plot of the number of predictive molecules kept in the model (with non-zero coefficients) on the x-axis vs. the mean (line) and standard deviation (shadow) MAE on the y-axis. MAE values were smoothed via a LOESS regression (R loess function with a span argument of 0.6). (H) Heatmap showing the correlation of the error in predictions (delta age) for the proteomics-based model and the small RNA-based models with R2 > 0.2. (I) Absolute standardized coefficients of the proteomics and top 20_miRNA-based models.