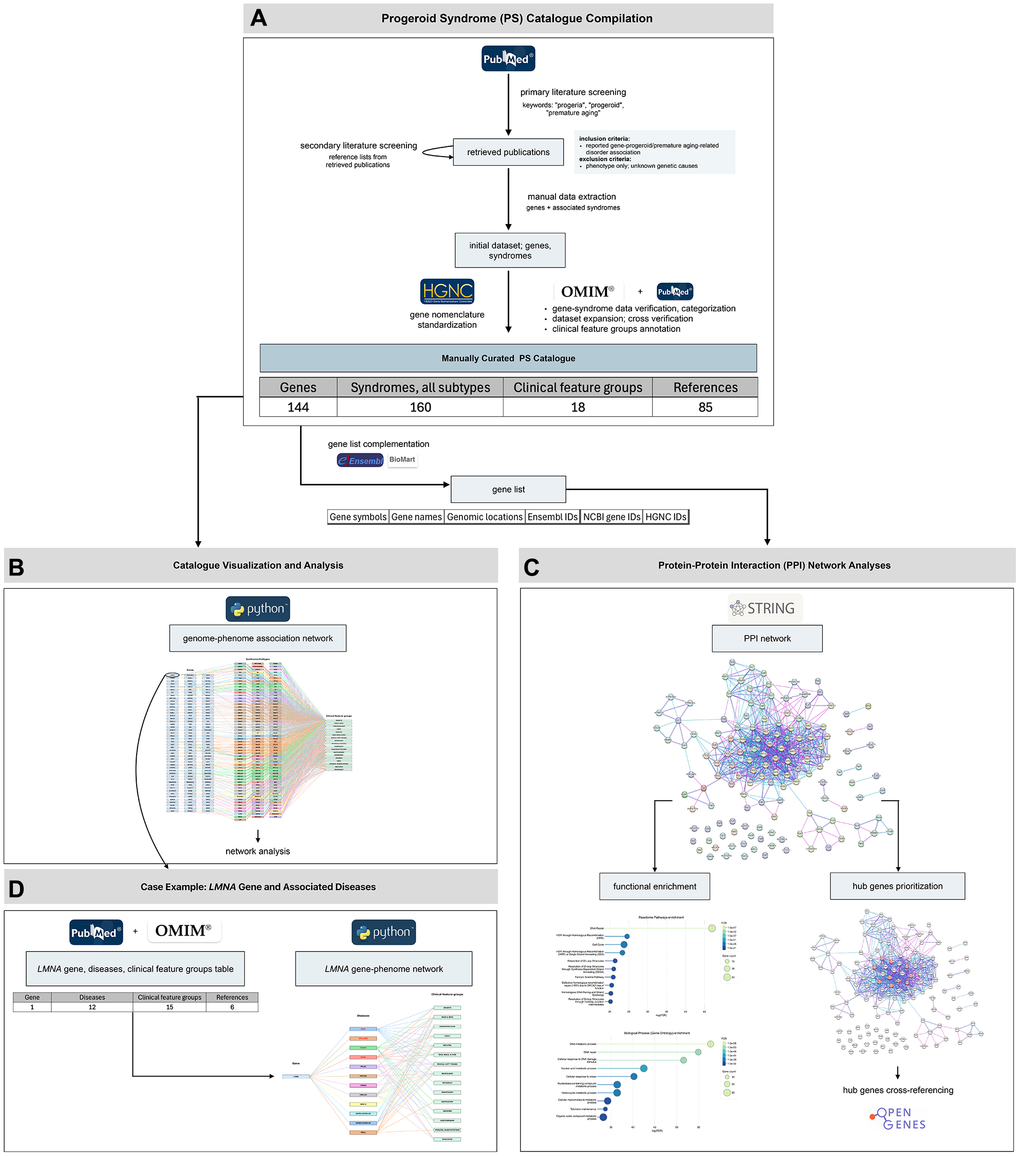
Figure 1.Overview of the study workflow and main results. The study proceeded through four major stages. (A) Progeroid syndrome (PS) catalogue compilation: We generated a manually curated PS catalogue following a structured workflow of primary and secondary literature screening, manual data extraction, data verification and categorization, dataset expansion, gene nomenclature standardization, and clinical feature groups annotation. The final dataset includes 144 genes, associated with a total of 160 clinical entities and 18 clinical feature groups, compiled from 84 publications and the OMIM database. (B) Catalogue visualization and analysis: The curated dataset was visualized through a genome–phenome association network and further explored though network analysis to assess genetic and phenotypic heterogeneity. (C) Protein–protein interaction (PPI) network analyses and functional characterization: The compiled gene set was analyzed through PPI network construction, functional enrichment of biological processes and pathways, and prioritization of highly connected hub genes, followed by cross-referencing with the Open Genes database. (D) Case example – LMNA gene: To illustrate single-gene pleiotropy, LMNA-associated diseases and their clinical feature groups were summarized in a dedicated table and a gene–phenotype network.