How to become immortal: let MEFs count the ways
Abstract
Understanding the molecular mechanisms and biological consequences of genetic changes occurring during bypass of cellular senescence spans a broad area of medical research from the cancer field to regenerative medicine. Senescence escape and immortalisation have been intensively studied in murine embryonic fibroblasts as a model system, and are known to occur when the p53/ARF tumour suppressor pathway is disrupted. We showed recently that murine fibroblasts with a humanised p53 gene (Hupki cells, from a human p53 knock-in mouse model) first senesce, and then become immortalised in the same way as their homologues with normal murine p53. In both cell types, immortalised cultures frequently sustain either a p53 gene mutation matching a human tumour mutation and resulting in loss of p53 transcriptional transactivation, or a biallelic deletion at the p19/ARF locus. Whilst these genetic events were not unexpected, we were surprised to find that a significant proportion of immortalised cell cultures apparently had neither a p53 mutation nor loss of p19/ARF. Here we consider various routes to p53/ARF disruption in senescence bypass, and dysfunction of other tumour suppressor networks that may contribute to release from tenacious cell cycle arrest in senescent cultures.
Culture
shock causes rapid replicative senescence in primary MEFs with functional p53,
but p53 deficient cells are resistant
Cellular senescence is an important
defence mechanism against tumour metastasis, growth and progression [1,2].
Furthermore, in both humans and in mouse models, cancers can respond to
chemotherapy by a massive senescence response followed by tumour cell clearance
[3-5]. MEFs are a classic model system for studying cell senescence and
immortalisation, with clear parallels to key genetic alterations during human
tumourigenesis whilst offering expedient advantages over human cell cultures
when exploring basic molecular mechanisms of senescence control and senescence
bypass [6,7]. When explanted in vitro MEFs initially continue to replicate,
but then rapidly undergo stress-associated senescence due to in vitro
conditions, especially oxidative stress elicited by standard culturing
conditions, which supply supraphysiological
levels of oxygen [8]. DNA damage from reactive oxygen species is clearly a key
factor in this senescence response [8,9]. As is well known from research in
several laboratories, cells that sustain spontaneous damage to the p53/p19ARF
pathway (p14 in humans) can overcome this replication block, leading to gradual
outgrowth of an immortalised cell population with unlimited growth potential
[10-14].
Our
MEF cell line library comprises immortalised cell lines from cultures of
fibroblasts derived from (a) embryos of a standard laboratory wild-type mouse
strain (129/Sv), and (b) embryos from human p53 knock-in (Hupki) mice, which we
constructed for our research on p53 biology [15-17]. The original Hupki mouse
strain [15] harbours normal human p53 gene sequences encoding the DNA binding
domain and the polyproline domain embedded in the endogenous murine p53 locus.
This strain is phenotypically normal, not tumour-prone, and displays classical
p53 wild-type responses, including DNA damage induced apoptosis,
transcriptional transactivation of p53 target genes and stress-induced cellular
senescence. Curiously, a mouse strain in which the entire p53 sequence was
replaced by the human counterpart lost wild-type p53 function due to abnormal
interactions with the p53 negative regulator Mdm2 [18]. The Hupki strain is not
p53-deficient, and can be used as a source of primary MEFs, thus allowing the
extensive literature on MEF senescence bypass, and the database of human tumour
suppressor mutations to be linked to specific mutations that support senescence
bypass. Using this approach, we have shown that the basic features of
stress-induced senescence and immortalisation are comparable in MEFs from
standard strain wild-type (WT) mice and Hupki mice. Studies from our laboratory
with several hundred immortalised Hupki MEF cell lines have shown that the
genetic alterations in p53 that lead to senescence bypass of MEFs are typical
of human tumours. Missense point mutations in p53 and p19/ARF silencing by
biallelic deletion, also common in human
tumours, are the two most common routes to spontaneous p53/p19ARF pathway
inactivation in immortalised MEFs identified thus far ([16,19,20] and
unpublished observations).
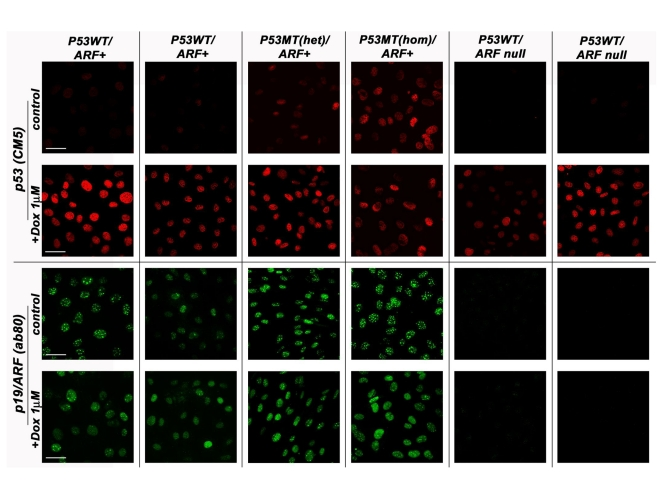
Figure 1. The p53/p19ARF status of various MEF lines derived from normal strain 129 mice. MEF cell lines
genotyped as either wild-type (WT) p53 or mutant (MT) p53 (both
heterozygous and homozygous) were compared against those carrying the
p19/ARF deletion in their response to doxorubicin (8h, 1μM) treatment.
Cells were methanol fixed and processed by indirect immunofluorescence
confocal microscopy with either the anti-p53 CM5 (Novacastra) or
anti-p19/ARF ab80 (AbCam) antibody. Scale bar represents 50μm. All samples
were processed at the same intensity and magnification.
Whilst
the prevalence of cell lines immortalised by p19/ARF biallelic deletion or p53
mutation was not unexpected (up to 50 % of cell lines, depending on
immortalisation protocols), we were surprised to find that a significant
fraction of cell lines (derived both from WT and Hupki primary cells) appeared
to have retained WT p53 and p19/ARF expression, as examined by DNA sequencing
or PCR amplification and promoter methylation analyses, respectively ([20] and
unpub-lished). Probing by immunofluorochemistry for p53 nuclear accumulation
and induction of p21/WAF1 following exposure of these cell lines to the DNA
damaging agent doxorubicin (Figure 1), as well as detection of p19/ARF protein
by immunoblotting [20] support the p53, p19/ARF wild-type status of these cell
lines. What might then be the genetic alterations that allowed these cells to
bypass senescence? One possibility concerns the phenomenon of tumour
suppressor haploinsufficiency in promoting cell growth (see insightful review
by Quon and Berns, 2001) [21,22]. Contrary to the original 2-hit paradigm
for tumour suppressor genes, where both alleles must be inactivated to elicit a growth promoting phenotype,
ample evidence from in vivo and in vitro studies and human tumour analyses
demonstrates that not only the absence, but also the moderate reduction of
tumour suppressor gene products can be sufficient to alter growth phenotype.
In the case of MEFs, as expected, cells from p53 null mice fail to senesce when
explanted in vitro under standard culture conditions, but cells from
heterozygous Hupki mice (progeny of Hupki and p53 null mice) also continue to
grow when explanted, with only a brief slowing of doubling time after the first
several passages (unpublished observations). Some of these immortal cultures
eventually do reveal loss of the WT allele with continued passaging, but others
retain the original unmutated p53 allele. This suggests that the presence of
only one WT allele in primary explanted MEFs may be sufficient to bypass
senescence initially. Conceivably, as passage number increases, so will the
chances that the WT allele is eventually discarded, possibly provoking a jump
in growth rate, as we have noted for some slow-growing MEF cell lines.
Some
of the MEF cell lines from (homozygous) WT or Hupki mice that we have examined
and tentatively classified as p53 and p19/ARF wild-type thus may in fact have
only one normal allele of these suppressor genes, having suffered loss of one
copy in vitro allowing unlimited growth in culture.
Beyond
p53/p19ARF
Given
that at least half of the spontaneously arising >100 MEF cell lines we
examined for p53 and p19/ARF aberrations appeared to have overcome senescence
block upon explanting in vitro by mechanisms other than the 2 canonical genetic
events (p53 mutation; p19 biallelic deletion), there could well be various
alternative pathways to immortalisation not directly involving damage to the immediate
p53/p19ARF axis.
Considerable effort is underway to
identify the key regulators of senescence and immortalisation. Since the
process of senescence bypass provides an intrinsic phenotypic readout of
functionality and automatically generates cell lines amenable to subsequent
analysis, reverse genetics is a powerful approach to deciphering the important
molecular events involved in senescence control [23]. Several large-scale
screens for genes involved in senescence/senescence bypass have been performed,
including both gain-of-function screens involving the ectopic expression of
cDNA libraries as well as loss-of-function screens involving the expression of
antisense cDNA libraries and shRNA libraries. These screens have been performed
in human and mouse cell models of both replicative and oncogene-induced senescence
and each approach has identified different genes involved in senescence,
providing novel and sometimes unexpected insights into the process (Table 1).
Reassuringly, well established players in the master regulatory pathways of
senescence (for example p53) have also been identified in these screens [24,25]. Indeed, inactivation of p53 is often used as a positive control in such
experiments [24,26]. While p53 knock-down in senescent MEFs has been shown to
reverse senescence [27], this may or may not be true of other genes involved in
senescence which may require inactivation or expression prior to the
acquisition of senescence.
The
role of p53 mutation/p19ARF deletion in senescence bypass in MEFs reflects the
importance of the p53/p19ARF axis as a master regulatory pathway of senescence
in these cells. What is apparent from genetic screening, as well as from other
complementary work, is that many of the novel senescence-associated genes
identified can also impact on this key pathway, both upstream and downstream of
p53 [24,26,28-31]. The p16/pRb pathway, another senescence master regulator,
is also commonly affected by novel senescence-associated molecules [28,32]. A
number of additional interacting signaling pathways have been implicated in the
induction or bypass of senescence including the RAS/MAPK pathway [25,33,34],
the AKT pathway [35-37] and the JNK pathway [38,39], although the relative
contribution of these to the senescent phenotype appears to be dependent on
species, cell type and the pro-senescence stimulus.
Concluding
remarks: Senescence - good or bad?
An entirely new aspect to the importance of cellular
senescence has recently surfaced from experiments to produce iPS (induced
pluripotent stem) cells from embryonic fibroblasts. It has been shown that
senescence provides a progressive barrier to conversion of primary MEFs (and
indeed other types of differentiated cells) to pluripotency. Crucially,
disruption of the p53/p19ARF signalling axis greatly increased efficiency of
their conversion [40,41]. Furthermore, genetic ablation of p53 in cells
normally considered refractory to reprogramming into pluripotent stem cells can
overcome this block [40]. Clearly, senescence is a key process to target in
optimising strategies to enhance somatic cell reprogramming. Identification of
factors influencing senescence will reveal novel genes/pathways to modulate
that could enhance conversion to pluripotency without compromising genetic integrity, expediting potential applications for iPS cells in
regenerative medicine.
Table 1. Novel genes identified in reverse-genetics senescence bypass screens.
The table shows the diversity of genes which either promote
senescence or its bypass as identified in cellular screens for
senescence bypass.
Genes well known to be important in cellular senescence such as p53, p21 and PAI-1 are not
included here.
aOther genes identified in this screen: BNIP3L, BIN1, HSPA9, IL1R1, PEA15, RAP1GAP, DMTF1, FOXA1, IRF1, MEN1, HIRA,
SMARCB1, FBXO31, NF2 [25].
bAdditional genes identified in this screen: RPS6KA6, HTATIP, HDAC4, SAH3, CCT2 [24].
| Gene | Promotion or inhibition of senescence | Potential
senescence-associated pathway/mechanism of action | Biological Function | Refe-rence | Cell type |
| BCL6 |
inhibition
|
Induces cyclin D1 expression
and
renders cells unresponsive
to antiproliferative signals from the p19(ARF)-p53 pathway
|
Transcription factor
|
[29]
|
MEFs, human B cells
|
| Bub1 |
inhibition
|
Bub1 RNAi induces
senescence. Bub1 expression does not extend lifespan
|
Mitotic checkpoint Ser/Thr
kinase
|
[25a, 28, 42]
|
Primary MEFs
|
| Csn2 |
promotion
|
Inactivation inhibits p53
transcriptional activity and confers resistance to both p53- and
p16INK4a-induced proliferation arrest
|
Component of the Cop9
signalosome
|
[28]
|
Primary MEFs
|
| Brf1 |
promotion
|
Inhibition of p53 transcription
and reduction p16ink4a-induced arrest
|
Subunit of the RNA
polymerase II complex
|
[28]
|
Primary MEFs
|
| Aldose Reductase |
promotion
|
Inhibition of p53
transcription and reduction p16ink4a-induced arrest
|
Metabolic enzyme - glucose
metabolism
|
[28]
|
Primary MEFs
|
| Tid1 |
induction
|
Tid1 is a repressor of
NF-κB signaling
|
DNA-J like protein which
functions as a co-chaperone
|
[43]
|
Rat embryo fibroblasts
|
| hDRIL1 |
inhibition
|
Renders primary MEFs
unresponsive to RAS(V12)-induced anti-proliferative signaling by
p19(ARF)/p53/p21(CIP1), as well as by p16(INK4a)
Binds E2F1 and
induces Cyclin E1
|
Transcription factor
|
[32]
|
MEFs
RASV12
induced senescence
|
| CBX7 |
inhibition
|
Controls cellular lifespan
through regulation of both the p16(Ink4a)/Rb and the Arf/p53 pathways
Represses INK4a-ARF locus
|
Transcription factor
|
[44]
|
Normal human prostate
epithelial cells
|
| LPA(2) |
inhibition
|
E2F induction
|
Phospholipid receptor
|
[45]
|
Mouse neuronal cells
|
| Dbs |
inhibition
|
E2F induction
|
Rho-specific guanine
nucleotide exchange factor
|
[45]
|
Mouse neuronal cells
|
| TBX2 |
inhibition
|
TBX2 represses the Cdkn2a
(p19(ARF)) promoter
|
Transcription factor
|
[31]
| Bmi1-/- MEFs
|
| TBX3 |
inhibition
|
TBX-3 potently represses
expression of both mouse p19(ARF) and human p14(ARF)
|
Transcription factor
|
[30]
|
Mouse neuronal cells
|
| Topo1 |
promotion
|
DNA damage-ATM-p53
|
Nuclear enzyme regulating
DNA structure
Relaxes positively and
negatively supercoiled DNA
|
[26]
|
Normal human cells
|
| IGFBP7 |
promotion
|
MEK, ERK pathway
In
Brafv600E-mediated senescence, IGFBP7 inhibits BRAF-MEK-ERK signaling by
inducing RKIP, which prevents BRAF from phosphorylating MEK
|
Ser/Thr protein kinase,
oncogene
Growth factor receptor
|
[25, 46]
|
Human primary fibroblasts,
melanocytes
|
| KLF4 |
promotion
|
p53 pathway
Suppresses the expression of
p53 by directly acting on its promoter
Induces p21
|
Transcription factor
|
[47]
|
Conditionally immortalized
MEFs
co-expressing RASV12 |
| SAHH |
promotion
|
p53 pathway
SAHH inactivation inhibits
p53 transcriptional activity
| |
[24b,
28, 48]
|
Primary human fibroblasts,
Primary MEFs
|
| CXCR2 (IL8RB) |
promotion
|
p53 pathway
CXCR2 knock-down alleviates
both replicative and oncogene-induced senescence and diminishes the
DNA-damage response.
|
Chemokine receptor
|
[49]
|
Primary human fibroblasts
|
Conflicts of Interest
The authors of this manuscript have no conflict of
interest to declare.
References
-
1.
Collado
M
, Blasco
MA
and Serrano
M.
Cellular senescence in cancer and aging.
Cell.
2007;
130:
223
-233.
[PubMed]
.
-
2.
Lowe
SW
, Cepero
E
and Evan
G.
Intrinsic tumour suppression.
Nature.
2004;
432:
307
-315.
[PubMed]
.
-
3.
Schmitt
CA
, Fridman
JS
, Yang
M
, Lee
S
, Baranov
E
, Hoffman
RM
and Lowe
SW.
A senescence program controlled by p53 and p16INK4a contributes to the outcome of cancer therapy.
Cell.
2002;
109:
335
-346.
[PubMed]
.
-
4.
Xue
W
, Zender
L
, Miething
C
, Dickins
RA
, Hernando
E
, Krizhanovsky
V
, Cordon-Cardo
C
and Lowe
SW.
Senescence and tumour clearance is triggered by p53 restoration in murine liver carcinomas.
Nature.
2007;
445:
656
-660.
[PubMed]
.
-
5.
Collado
M
and Serrano
M.
Senescence in tumours: evidence from mice and humans.
Nat Rev Cancer.
2010;
10:
51
-57.
[PubMed]
.
-
6.
Hahn
WC
and Weinberg
RA.
Modelling the molecular circuitry of cancer.
Nat Rev Cancer.
2002;
2:
331
-341.
[PubMed]
.
-
7.
Zuckerman
V
, Wolyniec
K
, Sionov
RV
, Haupt
S
and Haupt
Y.
Tumour suppression by p53: the importance of apoptosis and cellular senescence.
J Pathol.
2009;
219:
3
-15.
[PubMed]
.
-
8.
Parrinello
S
, Samper
E
, Krtolica
A
, Goldstein
J
, Melov
S
and Campisi
J.
Oxygen sensitivity severely limits the replicative lifespan of murine fibroblasts.
Nat Cell Biol.
2003;
5:
741
-747.
[PubMed]
.
-
9.
Busuttil
RA
, Rubio
M
, Dolle
ME
, Campisi
J
and Vijg
J.
Oxygen accelerates the accumulation of mutations during the senescence and immortalization of murine cells in culture.
Aging Cell.
2003;
2:
287
-294.
[PubMed]
.
-
10.
Kamijo
T
, Zindy
F
, Roussel
MF
, Quelle
DE
, Downing
JR
, Ashmun
RA
, Grosveld
G
and Sherr
CJ.
Tumor suppression at the mouse INK4a locus mediated by the alternative reading frame product p19ARF.
Cell.
1997;
91:
649
-659.
[PubMed]
.
-
11.
Harvey
DM
and Levine
AJ.
p53 alteration is a common event in the spontaneous immortalization of primary BALB/c murine embryo fibroblasts.
Genes Dev.
1991;
5:
2375
-2385.
[PubMed]
.
-
12.
Serrano
M
, Lin
AW
, McCurrach
ME
, Beach
D
and Lowe
SW.
Oncogenic ras provokes premature cell senescence associated with accumulation of p53 and p16INK4a.
Cell.
1997;
88:
593
-602.
[PubMed]
.
-
13.
Kamijo
T
, Weber
JD
, Zambetti
G
, Zindy
F
, Roussel
MF
and Sherr
CJ.
Functional and physical interactions of the ARF tumor suppressor with p53 and Mdm2.
Proc Natl Acad Sci U S A.
1998;
95:
8292
-8297.
[PubMed]
.
-
14.
Roussel
MF
p53: regular or super.
Cancer Cell.
2002;
2:
434
-435.
[PubMed]
.
-
15.
Luo
JL
, Yang
Q
, Tong
WM
, Hergenhahn
M
, Wang
ZQ
and Hollstein
M.
Knock-in mice with a chimeric human/murine p53 gene develop normally and show wild-type p53 responses to DNA damaging agents: a new biomedical research tool.
Oncogene.
2001;
20:
320
-328.
[PubMed]
.
-
16.
Reinbold
M
, Luo
JL
, Nedelko
T
, Jerchow
B
, Murphy
ME
, Whibley
C
, Wei
Q
and Hollstein
M.
Common tumour p53 mutations in immortalized cells from Hupki mice heterozygous at codon 72.
Oncogene.
2008;
27:
2788
-2794.
[PubMed]
.
-
17.
Song
H
, Hollstein
M
and Xu
Y.
p53 gain-of-function cancer mutants induce genetic instability by inactivating ATM.
Nat Cell Biol.
2007;
9:
573
-580.
[PubMed]
.
-
18.
Dudgeon
C
, Kek
C
, Demidov
ON
, Saito
S
, Fernandes
K
, Diot
A
, Bourdon
JC
, Lane
DP
, Appella
E
, Fornace
AJ Jr
and Bulavin
DV.
Tumor susceptibility and apoptosis defect in a mouse strain expressing a human p53 transgene.
Cancer Res.
2006;
66:
2928
-2936.
[PubMed]
.
-
19.
Liu
Z
, Muehlbauer
KR
, Schmeiser
HH
, Hergenhahn
M
, Belharazem
D
and Hollstein
MC.
p53 mutations in benzo(a)pyrene-exposed human p53 knock-in murine fibroblasts correlate with p53 mutations in human lung tumors.
Cancer Res.
2005;
65:
2583
-2587.
[PubMed]
.
-
20.
Whibley
C
, Odell
AF
, Nedelko
T
, Balaburski
G
, Murphy
M
, Liu
Z
, Stevens
L
, Walker
JH
, Routledge
M
and Hollstein
M.
Wild-type and HUPKI (human P53 knock-in) murine embryonic fibroblasts: P53/ARF pathway disruption in spontaneous escape from senescence.
J Biol Chem.
2010;
Epub Feb 4
.
-
21.
Quon
KC
and Berns
A.
Haplo-insufficiency? Let me count the ways.
Genes Dev.
2001;
15:
2917
-2921.
[PubMed]
.
-
22.
Inoue
K
, Zindy
F
, Randle
DH
, Rehg
JE
and Sherr
CJ.
Dmp1 is haplo-insufficient for tumor suppression and modifies the frequencies of Arf and p53 mutations in Myc-induced lymphomas.
Genes Dev.
2001;
15:
2934
-2939.
[PubMed]
.
-
23.
Hannon
GJ
and Rossi
JJ.
Unlocking the potential of the human genome with RNA interference.
Nature.
2004;
431:
371
-378.
[PubMed]
.
-
24.
Berns
K
, Hijmans
EM
, Mullenders
J
, Brummelkamp
TR
, Velds
A
, Heimerikx
M
, Kerkhoven
RM
, Madiredjo
M
, Nijkamp
W
, Weigelt
B
, Agami
R
, Ge
W
and Cavet
G.
A large-scale RNAi screen in human cells identifies new components of the p53 pathway.
Nature.
2004;
428:
431
-437.
[PubMed]
.
-
25.
Wajapeyee
N
, Serra
RW
, Zhu
X
, Mahalingam
M
and Green
MR.
Oncogenic BRAF induces senescence and apoptosis through pathways mediated by the secreted protein IGFBP7.
Cell.
2008;
132:
363
-374.
[PubMed]
.
-
26.
Humbert
N
, Martien
S
, Augert
A
, Da
Costa M
, Mauen
S
, Abbadie
C
, de Launoit
Y
, Gil
J
and Bernard
D.
A genetic screen identifies topoisomerase 1 as a regulator of senescence.
Cancer Res.
2009;
69:
4101
-4106.
[PubMed]
.
-
27.
Dirac
AM
and Bernards
R.
Reversal of senescence in mouse fibroblasts through lentiviral suppression of p53.
J Biol Chem.
2003;
278:
11731
-11734.
[PubMed]
.
-
28.
Leal
JF
, Fominaya
J
, Cascon
A
, Guijarro
MV
, Blanco-Aparicio
C
, Lleonart
M
, Castro
ME
, Ramon
YCS
, Robledo
M
, Beach
DH
and Carnero
A.
Cellular senescence bypass screen identifies new putative tumor suppressor genes.
Oncogene.
2008;
27:
1961
-1970.
[PubMed]
.
-
29.
Shvarts
A
, Brummelkamp
TR
, Scheeren
F
, Koh
E
, Daley
GQ
, Spits
H
and Bernards
R.
A senescence rescue screen identifies BCL6 as an inhibitor of anti-proliferative p19(ARF)-p53 signaling.
Genes Dev.
2002;
16:
681
-686.
[PubMed]
.
-
30.
Brummelkamp
TR
, Kortlever
RM
, Lingbeek
M
, Trettel
F
, MacDonald
ME
, van
Lohuizen M
and Bernards
R.
TBX-3, the gene mutated in Ulnar-Mammary Syndrome, is a negative regulator of p19ARF and inhibits senescence.
J Biol Chem.
2002;
277:
6567
-6572.
[PubMed]
.
-
31.
Jacobs
JJ
, Keblusek
P
, Robanus-Maandag
E
, Kristel
P
, Lingbeek
M
, Nederlof
PM
, van
Welsem T
, van
de Vijver MJ
, Koh
EY
, Daley
GQ
and van
Lohuizen M.
Senescence bypass screen identifies TBX2, which represses Cdkn2a (p19(ARF)) and is amplified in a subset of human breast cancers.
Nat Genet.
2000;
26:
291
-299.
[PubMed]
.
-
32.
Peeper
DS
, Shvarts
A
, Brummelkamp
T
, Douma
S
, Koh
EY
, Daley
GQ
and Bernards
R.
A functional screen identifies hDRIL1 as an oncogene that rescues RAS-induced senescence.
Nat Cell Biol.
2002;
4:
148
-153.
[PubMed]
.
-
33.
Dasari
A
, Bartholomew
JN
, Volonte
D
and Galbiati
F.
Oxidative stress induces premature senescence by stimulating caveolin-1 gene transcription through p38 mitogen-activated protein kinase/Sp1-mediated activation of two GC-rich promoter elements.
Cancer Res.
2006;
66:
10805
-10814.
[PubMed]
.
-
34.
Volonte
D
, Zhang
K
, Lisanti
MP
and Galbiati
F.
Expression of caveolin-1 induces premature cellular senescence in primary cultures of murine fibroblasts.
Mol Biol Cell.
2002;
13:
2502
-2517.
[PubMed]
.
-
35.
Binet
R
, Ythier
D
, Robles
AI
, Collado
M
, Larrieu
D
, Fonti
C
, Brambilla
E
, Brambilla
C
, Serrano
M
, Harris
CC
and Pedeux
R.
WNT16B is a new marker of cellular senescence that regulates p53 activity and the phosphoinositide 3-kinase/AKT pathway.
Cancer Res.
2009;
69:
9183
-9191.
[PubMed]
.
-
36.
Kortlever
RM
, Higgins
PJ
and Bernards
R.
Plasminogen activator inhibitor-1 is a critical downstream target of p53 in the induction of replicative senescence.
Nat Cell Biol.
2006;
8:
877
-884.
[PubMed]
.
-
37.
Nogueira
V
, Park
Y
, Chen
CC
, Xu
PZ
, Chen
ML
, Tonic
I
, Unterman
T
and Hay
N.
Akt determines replicative senescence and oxidative or oncogenic premature senescence and sensitizes cells to oxidative apoptosis.
Cancer Cell.
2008;
14:
458
-470.
[PubMed]
.
-
38.
Das
M
, Jiang
F
, Sluss
HK
, Zhang
C
, Shokat
KM
, Flavell
RA
and Davis
RJ.
Suppression of p53-dependent senescence by the JNK signal transduction pathway.
Proc Natl Acad Sci U S A.
2007;
104:
15759
-15764.
[PubMed]
.
-
39.
MacLaren
A
, Black
EJ
, Clark
W
and Gillespie
DA.
c-Jun-deficient cells undergo premature senescence as a result of spontaneous DNA damage accumulation.
Mol Cell Biol.
2004;
24:
9006
-9018.
[PubMed]
.
-
40.
Utikal
J
, Polo
JM
, Stadtfeld
M
, Maherali
N
, Kulalert
W
, Walsh
RM
, Khalil
A
, Rheinwald
JG
and Hochedlinger
K.
Immortalization eliminates a roadblock during cellular reprogramming into iPS cells.
Nature.
2009;
460:
1145
-1148.
[PubMed]
.
-
41.
Banito
A
, Rashid
ST
, Acosta
JC
, Li
S
, Pereira
CF
, Geti
I
, Pinho
S
, Silva
JC
, Azuara
V
, Walsh
M
, Vallier
L
and Gil
J.
Senescence impairs successful reprogramming to pluripotent stem cells.
Genes Dev.
2009;
23:
2134
-2139.
[PubMed]
.
-
42.
Gjoerup
OV
, Wu
J
, Chandler-Militello
D
, Williams
GL
, Zhao
J
, Schaffhausen
B
, Jat
PS
and Roberts
TM.
Surveillance mechanism linking Bub1 loss to the p53 pathway.
Proc Natl Acad Sci U S A.
2007;
104:
8334
-8339.
[PubMed]
.
-
43.
Tarunina
M
, Alger
L
, Chu
G
, Munger
K
, Gudkov
A
and Jat
PS.
Functional genetic screen for genes involved in senescence: role of Tid1, a homologue of the Drosophila tumor suppressor l(2)tid, in senescence and cell survival.
Mol Cell Biol.
2004;
24:
10792
-10801.
[PubMed]
.
-
44.
Gil
J
, Bernard
D
, Martinez
D
and Beach
D.
Polycomb CBX7 has a unifying role in cellular lifespan.
Nat Cell Biol.
2004;
6:
67
-72.
[PubMed]
.
-
45.
Kortlever
RM
, Brummelkamp
TR
, van
Meeteren LA
, Moolenaar
WH
and Bernards
R.
Suppression of the p53-dependent replicative senescence response by lysophosphatidic acid signaling.
Mol Cancer Res.
2008;
6:
1452
-1460.
[PubMed]
.
-
46.
Michaloglou
C
, Vredeveld
LC
, Soengas
MS
, Denoyelle
C
, Kuilman
T
, van der Horst
CM
, Majoor
DM
, Shay
JW
, Mooi
WJ
and Peeper
DS.
BRAFE600-associated senescence-like cell cycle arrest of human naevi.
Nature.
2005;
436:
720
-724.
[PubMed]
.
-
47.
Rowland
BD
, Bernards
R
and Peeper
DS.
The KLF4 tumour suppressor is a transcriptional repressor of p53 that acts as a context-dependent oncogene.
Nat Cell Biol.
2005;
7:
1074
-1082.
[PubMed]
.
-
48.
Leal
JF
, Ferrer
I
, Blanco-Aparicio
C
, Hernandez-Losa
J
, Ramon
YCS
, Carnero
A
and Lleonart
ME.
S-adenosylhomocysteine hydrolase downregulation contributes to tumorigenesis.
Carcinogenesis.
2008;
29:
2089
-2095.
[PubMed]
.
-
49.
Acosta
JC
, O'Loghlen
A
, Banito
A
, Guijarro
MV
, Augert
A
, Raguz
S
, Fumagalli
M
, Da
Costa M
, Brown
C
, Popov
N
, Takatsu
Y
, Melamed
J
and d'Adda
di Fagagna F.
Chemokine signaling via the CXCR2 receptor reinforces senescence.
Cell.
2008;
133:
1006
-1018.
[PubMed]
.