Clock genes, hair growth and aging
Abstract
Hair follicles undergo continuous cycles of growth, involution and rest. This process, referred to as the hair growth cycle, has a periodicity of weeks to months. At the same time, skin and hair follicles harbor a functional circadian clock that regulates gene expression with a periodicity of approximately twenty four hours. In our recent study we found that circadian clock genes play a role in regulation of the hair growth cycle during synchronized hair follicle cycling, uncovering an unexpected connection between these two timing systems within skin. This work, therefore, indicates a role for circadian clock genes in a cyclical process of much longer periodicity than twenty four hours.
The hair growth cycle
Beginning after completion of hair morphogenesis
(postnatal day 14 in the mouse), hair growth cycles commence with catagen, an
involution process of the hair follicle during which the majority of its
epithelial compartments undergo apoptosis [1]. This stage is followed by
telogen during which the hair follicle remains in relative quiescence. Telogen
is interrupted by activation of epithelial stem and progenitor cells located in
specialized stem cell compartments in the bulge and secondary hair germ, followed
by rapid proliferation and differentiation of progeny keratinocytes; this
growth phase is referred to as anagen [2]. In mice, the two initial hair
growth cycles are synchronized such that the majority of hair follicles are in
a similar stage of the hair growth cycle at a given time. But as the mouse
ages, the hair growth cycles become progressively less synchronized [3].
Circadian gene expression is hair growth cycle dependent
In order to systematically discover
transcriptional activity associated with the hair growth cycle, we
profiled mRNA expression at a genome-wide level over multiple time points
corresponding to morphogenesis and two synchronized hair growth cycles.
Interestingly, a large fraction of the genome, more than six thousand genes,
exhibits changes in expression that correlates with the progression of the hair
growth cycle, thus underscoring the complexity of this process [4,5]. One
of the surprises that came from this study was the finding that genes regulated
by the core circadian clock mechanism showed expression changes that correlated
with the hair growth cycle, with highest expression during the telogen-anagen
transition.
On a molecular level, the circadian clock consists of
positive and negative feedback loops. At its core are the bHLH-PAS
transcriptional activators CLOCK and BMAL1 (ARNTL), which form a heterodimer
and activate target genes containing E-boxes in their enhancer regions,
including Periods (Per1, 2 and 3) and Cryptochromes (Cry1 and 2). PERs and
CRYs form heterodimeric complexes that translocate into the nucleus where they
inhibit BMAL1-CLOCK transcriptional activity, thus constituting the negative
feedback loop [6]. In other words, the PER/CRY complex inhibits its own
expression, allowing for reactivation of BMAL1/CLOCK leading to rhythmic
expression with a periodicity of 24 hours. Several other proteins, including
kinases, play an important role in generation of rhythmic expression [7]. The
CLOCK-BMAL1 heterodimer activates other genes as well, including Dbp, Tef, Hlf,
and Rev-Erbα, which codes for an orphan nuclear receptor. REV-ERBα
regulates transcription of Bmal1 and other target genes by binding to retinoic
acid-related orphan receptor response elements (ROREs) [8]. The clock genes that
we identified as upregulated in telogen/early anagen were all CLOCK/BMAL1
target genes, including Pers, Dbp and Rev-Erbα. While these genes show a
clear circadian pattern of expression in skin, as was previously demonstrated [9-11],
their amplitude was higher during telogen and early anagen (Figure 1),
indicating that in skin, the expression of clock controlled genes is dependent
both on circadian mechanism and the hair growth cycle.
Since our mRNA expression studies
were performed using whole skin, we asked which compartments of the skin and
hair follicles contribute to the robust rhythmic circadian gene expression in telogen. The hair
follicle contains several functionally and
structurally distinct compartments, including the bulge region, which harbors
slow-cycling hair follicle stem cells [12]; the secondary hair germ, which
contains actively cycling stem and progenitor cells [13-15]; and the dermal
papilla, a source of signals for activating the stem cells at the beginning of
anagen [16] (Figure 2). While in situ hybridization studies revealed that all
cell types of the skin express circadian clock genes, the site of most
prominent rhythmic circadian gene expression during telogen and early anagen
was the secondary hair germ. This compartment, strategically positioned
between the dermal papilla and the bulge, contains proliferative Lrg5-positive
stem cells thought to have migrated from the bulge during late catagen and
early telogen [13,17]. The secondary hair germ cells are the first to be
activated during anagen initiation, giving rise to transient amplifying cells
of the hair matrix and eventually differentiating into the hair shaft [15,17,18]. Additionally, our data shows that as anagen progresses, the circadian
amplitude within the hair follicle proper becomes dampened, while the circadian
amplitude in the dermis and interfollicular epidermis continues to be robust.
Interestingly, suspension of circadian rhythm has been previously noted in
other highly proliferative and differentiating tissues, including testis and
thymus [19-21].
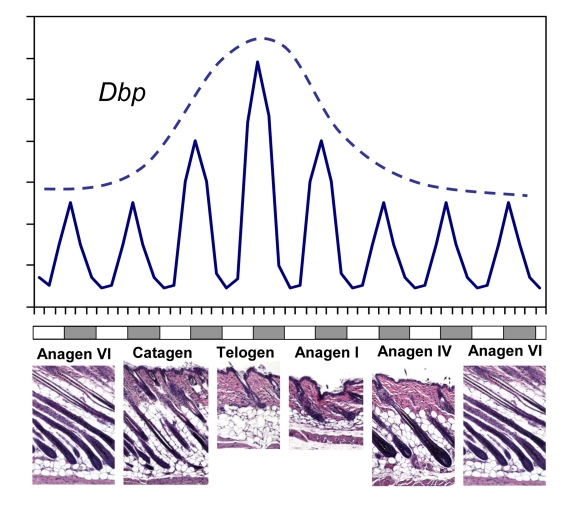
Figure 1. CLOCK-controlled gene expression in skin has a circadian pattern and correlates with synchronized hair growth cycles. A schematic
diagram showing rhythmic circadian expression of clock controlled gene Dbp
over different phases of the hair growth cycle (solid line). The circadian
amplitude of Dbp expression correlates with progression of the hair
follicle cycle with highest expression during telogen (broken line). Skin
histology for representative hair growth cycle stages is shown below. Note
that this schematic does not show the actual length of each phase of the
hair growth cycle.
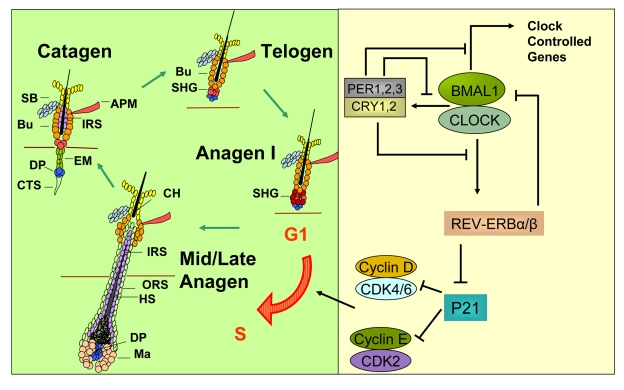
Figure 2. A model for how circadian clock genes participate in regulation of the synchronized hair growth cycle through regulation of cell cycle progression in the secondary hair germ. The hair growth cycle (left panel) is a
continuous process consisting of the quiescent telogen phase followed by
the growth phase (anagen) where signals, presumably originating in the
dermal papilla, activate stem and progenitor cell proliferation leading to
growth and differentiation of the hair shaft. Anagen is followed by catagen
where the lower two-thirds of the follicle undergo apoptosis, sparing the
stem cell compartments and the dermal papilla. The CLOCK/BMAL1 complex is
at the core of the mammalian circadian clock mechanism (right panel). It
activates multiple genes, collectively referred to as clock controlled
genes. Among these genes are Per1,2,3 and Cry1,2 whose protein
products translocate into the nucleus to inhibit the transcriptional
activity of the CLOCK/BMAL1 complex. Rev-erbα is another
clock controlled gene whose protein product negatively regulates expression
of Bmal1. Additionally, REV-ERBα
directly inhibits expression of the G1-S cell cycle inhibitor p21WAF1/CIP.
In the absence of BMAL1, downregulation of Rev-erbα leads to high P21
expression and G1 arrest in the hair germ cells during anagen I of the hair
follicle cycle, thus delaying anagen progression. APM - arrector pili
muscle, Bu - bulge, CH - club hair, CTS - connective tissue sheath, DP -
dermal papilla, EM - epithelial membrane, HS - hair shaft, IRS - inner root
sheath, Ma - matrix, ORS - outer root sheath, SB - sebaceous gland, SHG -
secondary hair germ.
Role for circadian clock genes in hair growth cycling
The robust circadian clock gene
expression within the secondary hair germ led us to test the possibility that
circadian clock genes might play a role in the hair growth cycle. For these
studies, we turned to Clock and Bmal1 mutant mouse models. We found a
significant delay in anagen progression in both mutants and this delay was more
pronounced in Bmal1 deficient mice, possibly due to partial functional
redundancy between Clock and its homologue Npas2. Clock and Bmal1 mutant mice
have no visible defects in hair follicle morphogenesis and enter the first
stage of anagen, characterized by the expansion of the secondary hair germ, at
approximately the same time (post-natal day 22). Yet by day 28, when the
majority of hair follicles in control littermates
have developed hair matrix and hair shaft with the hair bulb growing into the
subcutis, the Bmal1 mutant mice remained in the first anagen phase [5]. After
experiencing a nearly week-long delay, the Bmal1 deficient hair follicles
resumed normal progression of the hair cycle. There were no abnormalities in
the structure of the mature anagen follicles in the Bmal1 or Clock mutant mice,
supporting the idea that circadian clock genes are primarily involved in timing
mechanisms during the telogen-anagen transition.
Further analysis revealed absence of mitotic cells in
the early anagen secondary hair germ in Bmal1 mutant hair follicles, while
wild-type secondary hair germs at the same stage contained mitotic cells.
Importantly, epidermis and dermis of Bmal1 mutant mice contained mitotic cells,
indicating that the proliferation defect was hair follicle specific.
Phosphorylated Retinoblastoma Protein (Rb), a marker of cell cycle progression
through the G1-S cell cycle checkpoint [22], was absent in the secondary hair
germ of Bmal1 mutant hair follicles while it was abundant in control mice.
These results indicate that in Bmal1 mutant hair follicles, progenitor cells of
the early anagen secondary hair germ are arrested at the G1-S cell cycle
checkpoint.
To gain insights into the
molecular mechanisms underlying the G1 arrest of progenitor cells in the
secondary hair germ, we profiled gene expression in the skin of Bmal1 deficient
mice during telogen. As expected, the expression of multiple known CLOCK-BMAL1
target genes was affected, including that of Rev-Erbα, which was
downregulated approximately fifteen fold. Studies in hepatocytes have
demonstrated that REV-ERBα directly represses expression of the gene
encoding the G1 cell cycle inhibitor p21WAF1/CIP [23], and consistently p21 is
upregulated approximately 2.5 fold in Bmal1 mutant skin. These findings led
us to propose that hair growth cycling in Bmal1 mutant mice is delayed due to
upregulation of p21, leading to slowed G1-S cell cycle progression in
progenitor cells of the secondary hair germ (Figure 2). These results are
consistent with the known extensive crosstalk between the circadian clock and
the cell division cycle [24]. We have also considered the possibility that
circadian gene regulation of the hair growth might involve a mechanism that
"counts" the number of circadian peaks to regulate timing in the hair growth
cycle. However, results from our preliminary experiments in mice entrained to
22 and 26 hour days argue against this possibility.
In the mouse, the first two hair growth cycles are
synchronized. After completion of the second telogen, which can last up to
thirty days, the coat begins to grow asynchronously in complex domains created
by waves of anagen moving through the domain until a wave reaches "refractory"
telogen, an area of skin unresponsive to the propagating anagen stimulus. As
the mouse ages, this process creates increasingly complex patterns of hair
growth with each domain consisting of a telogen competent to be activated, a
propagating anagen wave, a catagen, and a refractive telogen [3,25]. In
preliminary experiments, we did not observe differential expression of clock
controlled genes in skin corresponding to different hair growth phases in
asynchronously cycling skin, suggesting the possibility that hair cycle related
regulation of clock gene expression may be particularly important in
synchronized hair follicle cycling.
One plausible role for circadian
mechanisms in the hair growth cycle is in animals with seasonal hair growth, commonly found in mammals living in the wild [26-30].
In these animals, the circadian clock-regulated hormones melatonin and
prolactin are thought to be key regulators of seasonal changes in hair growth [31-34].
Intriguingly, seasonal hair growth has been found to be regulated at the
telogen-anagen transition; in several breeds of sheep, the winter coat is in
telogen and in the spring when duration of daylight increases, hair follicles
enter anagen and the winter hair fibers are shed [35]. Therefore telogen, and
specifically the secondary hair germ, could serve as an important interpreter
of photic hair growth cycle timing in animals bearing seasonal fur.
The aging hair follicle
Hair loss and hair graying are commonly recognized
symptoms of aging in mammals. In addition to the visible location of hair, the
highly regenerative nature of hair follicles may explain why hair loss is a
prominent feature of aging syndromes. Several mouse models with premature aging
phenotypes show progressive hair loss or graying [36-38], and human conditions
with progeria-like symptoms, such as Werner Syndrome and Hutchinson-Gilford
Progeria, present with premature hair loss or hair graying [39]. The common
form of human hair loss, androgenetic alopecia, shows a clear age-related
progression [40]. In addition, some authorities have argued for a distinct
age-related entity, referred to as senescent alopecia [40-43]. This syndrome as
well as androgenetic alopecia are characterized by a reduction in the large
diameter pigmented (terminal) hair and an increased prevalence of thin
(vellus-like) hair [40]. Thus, both syndromes are thought to represent hair
growth cycle defects characterized by increased telogen to anagen hair follicle
ratio due to a shortened anagen phase and persistent telogen follicles [42,44].
Age-associated hair graying has been linked to
ultraviolet light and reactive oxygen species (ROS)-induced cell damage. The
hair follicle bulge harbors melanocyte stem cells that give rise to mature
melanocytes which synthesize and secrete hair pigments during anagen. Graying
human hair follicles have been shown to contain melanocytes with accumulated
oxidative stress [45], and consistent with this finding, genotoxic stress induced
by ionizing radiation in mice leads to premature differentiation of melanocyte
stem cells, followed by stem cell depletion and hair graying [46]. Furthermore,
deficiency in the ATM gene enhanced ectopic differentiation of the melanocyte
stem cells [46]. Additionally, both Werner Syndrome and Hutchinson-Gilford
Progeria are associated with accumulation of DNA damage [47]. Together, this
data suggests that hair graying may be due to melanocyte stem cell depletion
caused by UV radiation and genotoxic ROS.
The mechanisms underlying senescent alopecia have not
been extensively studied. However, a plausible hypothesis is that analogous to
hair graying, alopecia is related to loss of hair follicle epithelial stem
cells either through decreased renewal, premature differentiation, apoptosis or
cellular senescence. While the link between genotoxic stress and hair graying
has been established using mouse models, only correlative data from human
patients are available in regards to life-long UV exposure and hair loss [40,45]. Skin and hair follicles are heavily bombarded by UV radiation and also
contain actively dividing keratinocytes, a likely source of mito-chondrial ROS.
Circadian clock and the aging hair follicle
Recent studies indicate that circadian clock proteins
may be involved in DNA repair and in regulating accumulation of cellular ROS,
thus making them plausible actors in the aging processes [48]. Fu et. al.
demonstrated that a mutation in the Per2 gene leads to an increase in tumor
development as well as hair graying and hair loss after gamma irradiation [49].
PER1 is also known to interact with ATM and CHK2, thus affecting the proper
initiation of double strand break repair [50]. In addition, recent work from
several groups has revealed an important link between the circadian clock and
cellular metabolism [51-54]. This work suggests that cell division and DNA
synthesis are temporally segregated from the oxidative phase of the metabolic
cycle [55], raising the possibility that the circadian clock has evolved to
coordinate cell division with cellular metabolism, thus minimizing DNA damage.
Furthermore, the NADH/NAD+ ratio and heme, indicators of the redox state, have
been shown to directly modulate activity of circadian clock proteins, suggesting
that the circadian clock can read and interpret the cellular metabolic state [56].
In addition, SIRT1, the mammalian orthologue of yeast SIR2, is a conserved
NAD+-dependent protein deacetylase that deacetylates BMAL1 and PER2, and
functions as a histone deacetylase at clock-regulated promoters [51,52,57].
SIR2 and its orthologues are important regulators of longevity in yeast, worms
and flies [58-60], and in mice, several studies demonstrate SIRT1 contribution
to genome stability and DNA repair [61,62]. Together, these data suggest the
possibility that sirtuins could regulate longevity in part through circadian
clock mechanisms [56].
Direct evidence for clock involvement in
the aging process comes from the study of Kontdravov et. al. [38], showing
that the lifespan of Bmal1 mutant mice is decreased by half and that the mice
exhibit a range of premature aging phenotypes. Among the phenotypes reported by
the authors of this study were age-related lens and cornea defects, reduced
subcutaneous fat, and hair regeneration defects, pointing to strong effect of
the Bmal1 deletion in exposed cutaneous tissues, including hair follicles. The
authors demonstrate that by thirty weeks of age, Bmal1 deficient mice
accumulate significantly more ROS than control animals, thus potentially
explaining the progeria-like phenotype. While our work focused on younger
Bmal1 mutated mice [5], prior to the development of aging symptoms, the finding
of circadian clock involvement in cell cycle progression within the secondary
hair germ may provide a partial explanation for the hair regeneration defect.
Alternatively, in the absence of BMAL1, deregulation of DNA damage and
oxidative stress responses could cause depletion of stem populations necessary
for hair regeneration. Furthermore, in several rodent models there is
deregulated suprachiasmatic nucleus electric activity and photic entrainment as
well as abnormal periodicity and amplitude of circadian gene expression during
the normal aging process [63-66]. Thus, a decline in the robustness of
circadian rhythms may contribute to the aging process.
In conclusion, our study demonstrated that
circadian clock genes can regulate the non-circadian cyclical hair growth
cycle, presumably via an effect on the progression of the cell cycle in a
progenitor cell compartment of the hair follicle, the secondary hair germ [5].
We speculate that circadian genes may play a role in aging-related alopecia
which is characterized by aberrations in the hair growth cycle.
Acknowledgments
This work was supported by NIH grant
AR44882 (to B.A.) and California Breast Cancer Research Fellowship 14GB-0163
(to M.G.). We thank Ambica Bhandari and Amelia Soto for reading the
manuscript.
Conflicts of Interest
The authors of this manuscript have no
conflict of interest to declare.
References
-
1.
Paus
R
and Foitzik
K.
In search of the "hair cycle clock": a guided tour.
Differentiation.
2004;
72:
489
-511.
[PubMed]
.
-
2.
Schneider
MR
, Schmidt-Ullrich
R
and Paus
R.
The hair follicle as a dynamic miniorgan.
Curr Biol.
2009;
19:
R132
-142.
[PubMed]
.
-
3.
Plikus
MV
, Mayer
JA
and de la
Cruz D.
Cyclic dermal BMP signalling regulates stem cell activation during hair regeneration.
Nature.
2008;
451:
340
-344.
[PubMed]
.
-
4.
Lin
KK
, Chudova
D
, Hatfield
GW
, Smyth
P
and Andersen
B.
Identification of hair cycle-associated genes from time-course gene expression profile data by using replicate variance.
Proc Natl Acad Sci U S A.
2004;
101:
15955
-15960.
[PubMed]
.
-
5.
Lin
KK
, Kumar
V
and Geyfman
M.
Circadian clock genes contribute to the regulation of hair follicle cycling.
PLoS Genet.
2009;
5:
e1000573
[PubMed]
.
-
6.
Lowrey
PL
and Takahashi
JS.
Mammalian circadian biology: elucidating genome-wide levels of temporal organization.
Annu Rev Genomics Hum Genet.
2004;
5:
407
-441.
[PubMed]
.
-
7.
Ko
CH
and Takahashi
JS.
Molecular components of the mammalian circadian clock.
Hum Mol Genet.
2006;
15 Spec No 2:
R271
-277.
[PubMed]
.
-
8.
Ukai-Tadenuma
M
, Kasukawa
T
and Ueda
HR.
Proof-by-synthesis of the transcriptional logic of mammalian circadian clocks.
Nat Cell Biol.
2008;
10:
1154
-1163.
[PubMed]
.
-
9.
Bjarnason
GA
, Jordan
RC
and Wood
PA.
Circadian expression of clock genes in human oral mucosa and skin: association with specific cell-cycle phases.
Am J Pathol.
2001;
158:
1793
-1801.
[PubMed]
.
-
10.
Zanello
SB
, Jackson
DM
and Holick
MF.
Expression of the circadian clock genes clock and period1 in human skin.
J Invest Dermatol.
2000;
115:
757
-760.
[PubMed]
.
-
11.
Tanioka
M
, Yamada
H
and Doi
M.
Molecular clocks in mouse skin.
J Invest Dermatol.
2009;
129:
1225
-1231.
[PubMed]
.
-
12.
Cotsarelis
G
, Sun
TT
and Lavker
RM.
Label-retaining cells reside in the bulge area of pilosebaceous unit: implications for follicular stem cells, hair cycle, and skin carcinogenesis.
Cell.
1990;
61:
1329
-1337.
[PubMed]
.
-
13.
Ito
M
, Kizawa
K
, Hamada
K
and Cotsarelis
G.
Hair follicle stem cells in the lower bulge form the secondary germ, a biochemically distinct but functionally equivalent progenitor cell population, at the termination of catagen.
Differentiation.
2004;
72:
548
-557.
[PubMed]
.
-
14.
Panteleyev
AA
, Jahoda
CA
and Christiano
AM.
Hair follicle predetermination.
J Cell Sci.
2001;
114:
3419
-3431.
[PubMed]
.
-
15.
Jaks
V
, Barker
N
and Kasper
M.
Lgr5 marks cycling, yet long-lived, hair follicle stem cells.
Nat Genet.
2008;
40:
1291
-1299.
[PubMed]
.
-
16.
Botchkarev
VA
, Botchkareva
NV
, Sharov
AA
, Funa
K
, Huber
O
and Gilchrest
BA.
Modulation of BMP signaling by noggin is required for induction of the secondary (nontylotrich) hair follicles.
J Invest Dermatol.
2002;
118:
3
-10.
[PubMed]
.
-
17.
Zhang
YV
, Cheong
J
, Ciapurin
N
, McDermitt
DJ
and Tumbar
T.
Distinct self-renewal and differentiation phases in the niche of infrequently dividing hair follicle stem cells.
Cell Stem Cell.
2009;
5:
267
-278.
[PubMed]
.
-
18.
Greco
V
, Chen
T
and Rendl
M.
A two-step mechanism for stem cell activation during hair regeneration.
Cell Stem Cell.
2009;
4:
155
-69.
[PubMed]
.
-
19.
Alvarez
JD
and Sehgal
A.
The thymus is similar to the testis in its pattern of circadian clock gene expression.
J Biol Rhythms.
2005;
20:
111
-121.
[PubMed]
.
-
20.
Morse
D
, Cermakian
N
, Brancorsini
S
, Parvinen
M
and Sassone-Corsi
P.
No circadian rhythms in testis: Period1 expression is clock independent and developmentally regulated in the mouse.
Mol Endocrinol.
2003;
17:
141
-151.
[PubMed]
.
-
21.
Okamura
H
Clock genes in cell clocks: roles, actions, and mysteries.
J Biol Rhythms.
2004;
19:
388
-399.
[PubMed]
.
-
22.
Herwig
S
and Strauss
M.
The retinoblastoma protein: a master regulator of cell cycle, differentiation and apoptosis.
Eur J Biochem.
1997;
246:
581
-601.
[PubMed]
.
-
23.
Grechez-Cassiau
A
, Rayet
B
, Guillaumond
F
, Teboul
M
and Delaunay
F.
The circadian clock component BMAL1 is a critical regulator of p21WAF1/CIP1 expression and hepatocyte proliferation.
J Biol Chem.
2008;
283:
4535
-4342.
[PubMed]
.
-
24.
Hunt
T
and Sassone-Corsi
P.
Riding tandem: circadian clocks and the cell cycle.
Cell.
2007;
129:
461
-464.
[PubMed]
.
-
25.
Plikus
MV
, Widelitz
RB
, Maxson
R
and Chuong
CM.
Analyses of regenerative wave patterns in adult hair follicle populations reveal macro-environmental regulation of stem cell activity.
Int J Dev Biol.
2009;
53:
857
-868.
[PubMed]
.
-
26.
Butler
MP
and Zucker
I.
Seasonal pelage changes are synchronized by simulated natural photoperiods in Siberian hamsters (Phodopus sungorus).
J Exp Zool A Ecol Genet Physiol.
2009;
311:
475
-482.
[PubMed]
.
-
27.
Ebling
FJ
, Alexander
IH
, Urbanski
HF
and Hastings
MH.
Effects of N-methyl-D-aspartate (NMDA) on seasonal cycles of reproduction, body weight and pelage colour in the male Siberian hamster.
J Neuroendocrinol.
1995;
7:
555
-566.
[PubMed]
.
-
28.
al-Khateeb
A
and Johnson
E.
Action of hormones on seasonal changes of pelage in the vole.
J Endocrinol.
1969;
43:
43
-44.
[PubMed]
.
-
29.
Rust
CC
, Shackelford
RM
and Meyer
RK.
Hormonal control of pelage cycles in the mink.
J Mammal.
1965;
46:
549
-565.
[PubMed]
.
-
30.
Randall
VA
, Thornton
MJ
, Messenger
AG
, Hibberts
NA
, Loudon
AS
and Brinklow
BR.
Hormones and hair growth: variations in androgen receptor content of dermal papilla cells cultured from human and red deer (Cervus elaphus) hair follicles.
J Invest Dermatol.
1993;
101:
114S
-120S.
[PubMed]
.
-
31.
Kauffman
AS
, Cabrera
A
and Zucker
I.
Energy intake and fur in summer- and winter-acclimated Siberian hamsters (Phodopus sungorus).
Am J Physiol Regul Integr Comp Physiol.
2001;
281:
R519
-527.
[PubMed]
.
-
32.
Rose
J
, Oldfield
J
and Stormshak
F.
Apparent role of melatonin and prolactin in initiating winter fur growth in mink.
Gen Comp Endocrinol.
1987;
65:
212
-215.
[PubMed]
.
-
33.
Rose
J
, Stormshak
F
, Oldfield
J
and Adair
J.
Induction of winter fur growth in mink (Mustela vison) with melatonin.
J Anim Sci.
1984;
58:
57
-61.
[PubMed]
.
-
34.
Sarkar
M
and Prakash
BS.
Circadian variations in plasma concentrations of melatonin and prolactin during breeding and non-breeding seasons in yak (Poephagus grunniens L.).
Anim Reprod Sci.
2005;
90:
149
-162.
[PubMed]
.
-
35.
Rogers
GE
Biology of the wool follicle: an excursion into a unique tissue interaction system waiting to be re-discovered.
Exp Dermatol.
2006;
15:
931
-949.
[PubMed]
.
-
36.
Tyner
SD
, Venkatachalam
S
and Choi
J.
p53 mutant mice that display early ageing-associated phenotypes.
Nature.
2002;
415:
45
-53.
[PubMed]
.
-
37.
Sun
LQ
, Lee
DW
and Zhang
Q.
Growth retardation and premature aging phenotypes in mice with disruption of the SNF2-like gene, PASG.
Genes Dev.
2004;
18:
1035
-1046.
[PubMed]
.
-
38.
Kondratov
RV
, Kondratova
AA
, Gorbacheva
VY
, Vykhovanets
OV
and Antoch
MP.
Early aging and age-related pathologies in mice deficient in BMAL1, the core componentof the circadian clock.
Genes Dev.
2006;
20:
1868
-1873.
[PubMed]
.
-
39.
Kamenisch
Y
and Berneburg
M.
Progeroid syndromes and UV-induced oxidative DNA damage.
J Investig Dermatol Symp Proc.
2009;
14:
8
-14.
.
-
40.
Trueb
RM
Aging of hair.
J Cosmet Dermatol.
2005;
4:
60
-72.
[PubMed]
.
-
41.
Kligman
AM
The comparative histopathology of male-pattern baldness and senescent baldness.
Clin Dermatol.
1988;
6:
108
-118.
[PubMed]
.
-
42.
Courtois
M
, Loussouarn
G
, Hourseau
C
and Grollier
JF.
Ageing and hair cycles.
Br J Dermatol.
1995;
132:
86
-93.
[PubMed]
.
-
43.
Courtois
M
, Loussouarn
G
, Hourseau
C
and Grollier
JF.
Hair cycle and alopecia.
Skin Pharmacol.
1994;
7:
84
-89.
[PubMed]
.
-
44.
Ellis
JA
and Sinclair
RD.
Male pattern baldness: current treatments, future prospects.
Drug Discov Today.
2008;
13:
791
-797.
[PubMed]
.
-
45.
Arck
PC
, Overall
R
and Spatz
K.
Towards a "free radical theory of graying": melanocyte apoptosis in the aging human hair follicle is an indicator of oxidative stress induced tissue damage.
FASEB J.
2006;
20:
1567
-1569.
[PubMed]
.
-
46.
Inomata
K
, Aoto
T
and Binh
NT.
Genotoxic stress abrogates renewal of melanocyte stem cells by triggering their differentiation.
Cell.
2009;
137:
1088
-1099.
[PubMed]
.
-
47.
Cox
LS
and Faragher
RG.
From old organisms to new molecules: integrative biology and therapeutic targets in accelerated human ageing.
Cell Mol Life Sci.
2007;
64:
2620
-2641.
[PubMed]
.
-
48.
Antoch
MP
and Kondratov
RV.
Circadian proteins and genotoxic stress response.
Circ Res.
2010;
106:
68
-78.
[PubMed]
.
-
49.
Fu
L
, Pelicano
H
, Liu
J
, Huang
P
and Lee
C.
The circadian gene Period2 plays an important role in tumor suppression and DNA damage response in vivo.
Cell.
2002;
111:
41
-50.
[PubMed]
.
-
50.
Gery
S
, Komatsu
N
, Baldjyan
L
, Yu
A
, Koo
D
and Koeffler
HP.
The circadian gene per1 plays an important role in cell growth and DNA damage control in human cancer cells.
Mol Cell.
2006;
22:
375
-382.
[PubMed]
.
-
51.
Nakahata
Y
, Sahar
S
, Astarita
G
, Kaluzova
M
and Sassone-Corsi
P.
Circadian control of the NAD+ salvage pathway by CLOCK-SIRT1.
Science.
2009;
324:
654
-657.
[PubMed]
.
-
52.
Nakahata
Y
, Kaluzova
M
and Grimaldi
B.
The NAD+-dependent deacetylase SIRT1 modulates CLOCK-mediated chromatin remodeling and circadian control.
Cell.
2008;
134:
329
-340.
[PubMed]
.
-
53.
Yin
L
, Wu
N
and Curtin
JC.
Rev-erbalpha, a heme sensor that coordinates metabolic and circadian pathways.
Science.
2007;
318:
1786
-1789.
[PubMed]
.
-
54.
Rutter
J
, Reick
M
and McKnight
SL.
Metabolism and the control of circadian rhythms.
Annu Rev Biochem.
2002;
71:
307
-331.
[PubMed]
.
-
55.
Tu
BP
, Kudlicki
A
, Rowicka
M
and McKnight
SL.
Logic of the yeast metabolic cycle: temporal compartmentalization of cellular processes.
Science.
2005;
310:
1152
-1158.
[PubMed]
.
-
56.
Eckel-Mahan
K
and Sassone-Corsi
P.
Metabolism control by the circadian clock and vice versa.
Nat Struct Mol Biol.
2009;
16:
462
-467.
[PubMed]
.
-
57.
Asher
G
, Gatfield
D
and Stratmann
M.
SIRT1 regulates circadian clock gene expression through PER2 deacetylation.
Cell.
2008;
134:
317
-328.
[PubMed]
.
-
58.
Rogina
B
and Helfand
SL.
Sir2 mediates longevity in the fly through a pathway related to calorie restriction.
Proc Natl Acad Sci U S A.
2004;
101:
15998
-6003.
[PubMed]
.
-
59.
Imai
S
, Armstrong
CM
, Kaeberlein
M
and Guarente
L.
Transcriptional silencing and longevity protein Sir2 is an NAD-dependent histone deacetylase.
Nature.
2000;
403:
795
-800.
[PubMed]
.
-
60.
Finkel
T
, Deng
CX
and Mostoslavsky
R.
Recent progress in the biology and physiology of sirtuins.
Nature.
2009;
460:
587
-591.
[PubMed]
.
-
61.
Wang
RH
, Sengupta
K
and Li
C.
Impaired DNA damage response, genome instability, and tumorigenesis in SIRT1 mutant mice.
Cancer Cell.
2008;
14:
312
-323.
[PubMed]
.
-
62.
Oberdoerffer
P
, Michan
S
and McVay
M.
SIRT1 redistribution on chromatin promotes genomic stability but alters gene expression during aging.
Cell.
2008;
135:
907
-918.
[PubMed]
.
-
63.
Yamazaki
S
, Straume
M
, Tei
H
, Sakaki
Y
, Menaker
M
and Block
GD.
Effects of aging on central and peripheral mammalian clocks.
Proc Natl Acad Sci U S A.
2002;
99:
10801
-10806.
[PubMed]
.
-
64.
Sutin
EL
and Kilduff
TS.
Circadian and light-induced expression of immediate early gene mRNAs in the rat suprachiasmatic nucleus.
Brain Res Mol Brain Res.
1992;
15:
281
-290.
[PubMed]
.
-
65.
Van
Reeth O
, Zhang
Y
, Reddy
A
, Zee
P
and Turek
FW.
Aging alters the entraining effects of an activity-inducing stimulus on the circadian clock.
Brain Res.
1993;
607:
286
-292.
[PubMed]
.
-
66.
Kolker
DE
, Fukuyama
H
, Huang
DS
, Takahashi
JS
, Horton
TH
and Turek
FW.
Aging alters circadian and light-induced expression of clock genes in golden hamsters.
J Biol Rhythms.
2003;
18:
159
-169.
[PubMed]
.