SOCS1, cancer and senescence
Cytokines are secreted proteins that regulate different cellular processes including survival, proliferation and differentiation. Following binding to their receptors, cytokines activate the Janus kinases (JAK1, JAK2, JAK3 and Tyk2) leading to the phosphorylation of tyrosine residues on the cytoplasmic portion of the receptor creating docking sites for signaling molecules containing a SH2 domain [1,2]. Members of the STAT family of proteins that are recruited to the phosphorylated cytokine receptors themselves become phosphorylation substrates for JAK kinases. Phosphorylated STAT proteins homo- or hetero- dimerize and translocate to the nucleus to activate transcription of target genes by binding to specific response elements in their promoter regions. Among these cytokine-induced proteins, members of the SOCS family constitute important negative regulators of the JAK/STAT signaling pathway.
There are eight members of the SOCS family of proteins (CIS, SOCS1-7), each of which harbor a central SH2 domain and a C-terminal SOCS box region [3] (Figure 1). The suppressor of cytokine signaling SOCS1 was initially identified as a cytokine-inducible inhibitor of STAT signaling [4,5,6]. Through its SH2 domain, SOCS1 can directly bind phosphorylated JAK2 to prevent the phosphorylation of STAT. SOCS1 also possesses a kinase inhibitory region (KIR), a domain composed of less than 30 amino acids, which shares homology with the pseudosubstrate inhibitory region of JAK and leads to inhibition of the catalytic activity of JAK [7,8]. The SOCS box allows recruitment of elongin B/C and Cullin 2 to form an ubiquitin E3 ligase complex [9,10]. This allows the SOCS protein to operate as an adaptor to trigger ubiquitination and degradation of proteins involved in cellular signaling including JAK [11], TEL-JAK2 [12], IRS-1/2 [13], FAK [14], Vav [15] and Mal [16]. It is currently thought that SOCS1 contributes to tumor suppression due to its ability to control and terminate the activation of STATs [17,18,19,20,21,22,23,24,25]. On the other hand, the relationship between SOCS1 and other tumor suppressor pathways and the cellular mechanisms by which SOCS1 might exert its tumor suppression remain largely unexplored.
To prevent the formation of cancer, normal cells possess intrinsic tumor suppressor mechanisms that are triggered upon oncogene activation. Like apoptosis, cellular senescence opposes cellular transformation by limiting the proliferation of cells expressing oncogenes. In normal human diploid cells, oncogene activation causes a permanent growth arrest with features of cellular senescence [26]. We have recently extended the list of oncogenes known to trigger the senescence response to include the JAK/STAT5 pathway. The transcription factor STAT5 is implicated in tumor formation by regulating important cellular processes including cell cycle progression, apoptosis, angiogenesis and metastasis [27]. However, in normal cells, expression of Tel/Jak2 or constitutively activated allele of STAT5A and B initiated a cell cycle arrest in G1 associated with markers of premature cellular senescence and activation of the tumor suppressors Rb and p53 [28,29,30].
SOCS box proteins and the regulation of p53
The activation of the p53 pathway following oncogene activation is crucial to induce senescence in normal cells. In mice, stimulation of p53 is dependent on p19ARF (Alternative Reading Frame), which is induced by several oncogenes [31,32]. However, the role of ARF in oncogene-induced senescence in human cells is still unclear [33]. In order to identify new regulators of p53 activation following constitutively activated STAT5 expression in normal cells, we performed microarray analysis covering the entire human transcriptome. We observed that the expression of SOCS1 was highly increased at both mRNA and protein level during STAT5-induced senescence [34]. Unexpectedly, SOCS1 expression in normal human fibroblasts was sufficient to trigger a p53-dependent cell cycle arrest displaying features of the senescence phenotype. This function of SOCS1 was dependent on the integrity of its SOCS box. In addition, SOCS1, but not a mutant lacking the SOCS box domain, led to the accumulation of phosphorylated p53 on serine 15 and increased transcription of the p53 target gene p21CIP. The knockdown of SOCS1 during STAT5-induced senescence reduced the phosphorylation of p53 on Ser15, diminished the nuclear accumulation of p53 and compromised the development of senescence phenotype [34]. The remaining activated p53 and partial bypass of the senescence response observed following the knockdown of SOCS1 might arise from the ability of STAT5 to engage multiple signaling pathways to ensure p53 activation. For example, STAT5 can directly transactivate the promoter of the PML gene and stimulate its expression in a p53-independent fashion [30]. The PML protein can then inhibit Mdm2 and stimulate p53 [35,36] contributing to the senescence phenotype [37,38].
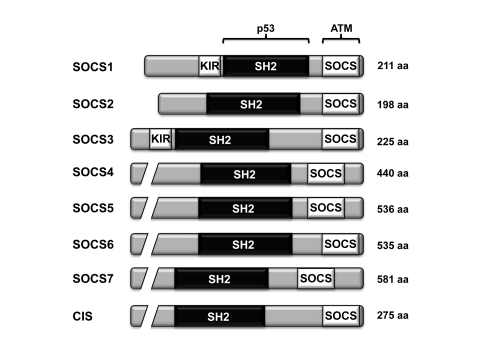
Figure 1. The domain architecture of the different members of the SOCS family of proteins. All eight members of the SOCS family
harbor a central SH2 domain and a C-terminal SOCS box. Both SOCS1 and
SOCS3 also contain a kinase inhibitory region (KIR). The region of
SOCS1 interacting with p53 and ATM are shown [34].
SOCS1 mediated STAT5-induced senescence via an unexpected protein-protein interaction between the SH2 domain of SOCS1 and the transactivation domain of p53 [34]. Because the transactivation domain of p53 harbors no tyrosine residues, the binding should occur independently of tyrosine phosphorylation, as reported before for SOCS1 binding to Vav [15] and for other SH2 domains as well [39,40]. The von Hippel-Lindau protein (VHL), another SOCS box-containing protein, has been recently shown to interact with p53. This interaction does not rely on an SH2 domain but on the SOCS box domain of VHL. However, like SOCS1, VHL facilitates p53 interaction with the DNA damage activated kinase ATM [41]. Hence, SOCS1 links DNA damage signals stimulated by oncogenic activity to p53.
Interestingly, SOCS1 is not the only protein inhibitor of STAT implicated in the regulation of p53 activity. The protein inhibitors of activated STAT, PIAS1 and PIASy both promote the sumoylation and transcriptional activity of p53 [42,43,44]. However, the mechanism of activation of p53 by PIAS is still unclear. While the sumoylation of p53 by PIAS1 has been demonstrated [43], a mutated PIAS1 lacking the RING finger-like domain and defective in promoting p53 sumoylation was sufficient to activate p53 [44]. Furthermore, by controlling the activity of both p53 and Rb, PIASy regulates Ras-induced senescence and apoptosis [42]. These data suggest that the control of STAT signaling is tightly linked to the activation of p53 to possibly control the JAK/STAT oncogenic pathway.
Inhibitors of STATs activity and the DNA damage response
The stimulation of p53 during oncogene-induced senescence is associated with the activation of the DNA damage response [28,45,46]. The DNA damage observed in normal cells expressing activated oncogenes may be due to reactive oxygen species [47] and/or some type of replicative stress [45,46]. SOCS1-induced senescence was accompanied by the activation of the DNA damage-regulated kinases ATM and Chk2. Since the stimulation of p53 reporters by SOCS1 was partially blocked in cells depleted of ATM, ATM might participate in the SOCS1-dependent activation of p53. Using pulldown assays, we demonstrated that SOCS1 interacted with both ATM and ATR through its SOCS box (Figure 1) [34]. ATM is an important mediator of the senescence response by activating the p53 pathway, mainly through phosphorylation of the Ser 15 residue [28,45,46]. Depletion of SOCS1 during STAT5-induced senescence caused a dramatic decrease in Ser15 phosphorylation of p53. In order to form a ternary complex with p53 and ATM, SOCS1 must localize to the nucleus. We confirmed that SOCS1 is able to localize to the nucleus and that endogenous SOCS1 colocalized to DNA damage foci with ATM during STAT5-induced senescence [34], thus reinforcing the notion that SOCS1 is a mediator of the DNA damage response. Not only SOCS1 but also other proteins controlling JAK/STAT signaling are known to localize to DNA damage sites. PIAS1 and PIAS4 were also shown to localize to DNA breaks and contribute to the DNA damage response by sumoylating BRCA1 [48,49]. Together, these findings strongly suggest a close link between cytokine signaling and the DNA damage response.
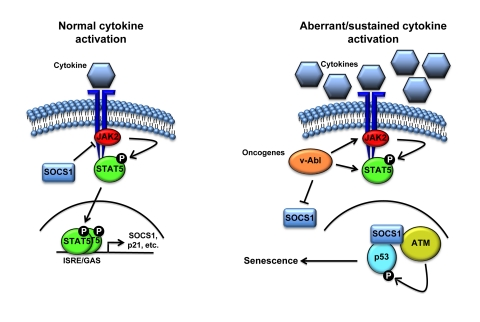
Figure 2. Schematic representation of the cell proliferation control exerted by SOCS1. Following
activation of the receptor by cytokine binding, JAK phosphorylates the
receptor creating a docking site for STATs. JAK then phopshorylates STATs
causing its release from the receptor, allowing dimerization and
translocation to the nucleus to activate the transcription of specific
genes including members of the SOCS family. Subsequently, SOCS terminates
cytokine signaling by blocking JAK activity and STAT recruitment to the
receptor. However, aberrant activation of STAT5 triggered by oncogenic
fusion kinases like TEL-JAK2 might result in sustained levels of SOCS1 that
can activate p53 by forming a complex with ATM and p53.
Cytokines, senescence and SOCS1: an emergency switch to control proliferation
Senescent cells secrete numerous cytokines and other mediators that modify the tissue microenvironment. The sum of these secreted factors constitutes what has been named the senescence-associated secretory phenotype (SASP) [50]. Among the SASP factors, IL-6 is required for the oncogene-induced senescence and induction of the tumor suppressor p15INK4B [51]. Furthermore, persistent, but not transient, DNA damage signaling triggers the ATM-dependent IL-6 secretion, presumably to call attention to the presence of damaged cells [52]. During oncogene-induced senescence, IL-6 also amplifies the secretion of IL-8 [51], which with GROαactivates the CXCR2 receptor to reinforce senescence [53]. Among the factors secreted by senescent cells, IGFBP7 [54] and PAI-1 [55] contribute to the growth arrest response, while p53 regulates expression of chemokines directing the immune system to permit the clearance of senescent cells [56]. Collectively, these reports suggest that cytokine signaling could prevent tumor formation by promoting cellular senescence.
The capacity of SOCS1 to activate the p53 pathway can establish an emergency anti-proliferative program in cells exposed to sustain or aberrant cytokine stimulation (Figure 2). Following normal activation of the JAK/STAT pathway, SOCS1 blocks the phosphorylation of STAT by inhibiting or degrading JAK2. However, aberrant and sustained stimulation of STAT might induce a molecular switch allowing SOCS1 to localize to DNA breaks and stimulate ATM-dependent activation of p53.
A general role for SOCS1 in the DNA damage response
The localization of SOCS1 to DNA breaks during STAT5-induced senescence raises numerous questions. First, does the SOCS1 ubiquitin ligase activity contribute to the DNA damage response? A novel cascade of ubiquitination controlled by the E3 ubiquitin ligases RNF8/RNF168 and HERC2 have recently been reported to control the recruitment of BRCA1 and 53BP1 by ubiquitinating the histones H2A and H2AX [57,58,59,60,61,62]. The presence of SOCS1 at DNA breaks could not only regulate ATM-mediated p53 activation but also control the DNA repair process. Second, what are the mechanisms underlying the nuclear transport of SOCS1 and its presence at DNA damage foci? Since most of its interacting partners were localized to the plasma membrane, SOCS1 was considered to be mostly a cytoplasmic protein, but recent evidences suggest that it can localize to the nucleus under certain conditions including STAT5-induced senescence [34,63]. A bipartite nuclear localization signal (NLS) located between the SH2 domain and the SOCS box allows nuclear localization of SOCS1 [63,64]. However, the mechanism controlling the active transport of SOCS1 remains unclear. A clearer understanding of the mechanisms controlling SOCS1 nuclear localization would be crucial to determine how SOCS1 mediates its tumor suppressor activity. Post-translational modifications like ubiquitination and phosphorylation that have been shown to control the nuclear localization of p53 [65,66,67] and STAT [68] could also control the nucleo-cytoplasmic shuttling of SOCS1. Exclusion of SOCS1 from the nucleus would prevent the formation of the ternary complex with p53 and ATM, preventing the activation of p53. Furthermore, the phosphorylation status of SOCS1 could regulate its activity since aberrant SOCS1 phosphorylation is associated with cellular transformation. Actually, phosphorylation of SOCS1 triggered by the oncogenic v-Abl kinase impedes the SOCS1-Elongin B/C interaction, leading to sustained JAK/STAT signaling [69]. v-Abl signaling induces multiple serine/threonine kinases including members of the Pim kinase family. Pim-1 and Pim-2 are required for efficient cellular transformation mediated by v-Abl [70] and are able to phosphorylate SOCS1 and disrupt its binding to Elongin C [71]. Because SOCS1 requires the SOCS box to form a complex with ATM, v-Abl- or Pim kinase-mediated phosphorylation could potentially interfere with this interaction and block p53 activation. Therefore, it appears that aberrant phosphorylation by oncogenic kinases could interfere with the tumor suppressor activities of SOCS1 by at least two different mechanisms: phosphorylated SOCS1 would not be able to inhibit the JAK/STAT pathway and to interact with ATM and promote p53 activation.
Table I. Identification of SOCS1 interaction partners by mass spectrometry*.
| Protein | Function |
| Elongin C | Interacts with SOCS box [10] |
| Elongin B | Interacts with SOCS box [10] |
| Pericentrin | Cells depleted of pericentrin enter senescence due to p53 activation [72]. |
| SHC (Src homology 2 domain containing) transforming protein 1 (SHC1) | Member of the Shc protein family of molecular adaptors, SHC1 promotes apoptosis by its redox activity. SHC1 is implicated in the control of oxidative stress and life span in mammals [73]. |
| Tripartite motif-containing 28 (TRIM28 or KAP1) | TRIM28 is implicated in transcriptional control through its interaction with the Kruppel-associated box repression domain. TRIM28 contributes to DNA repair mechanisms [74]. |
| 5'-nucleotidase, cytosolic II (NT5C2) | NT5C2 hydrolyzes 5-prime-monophosphate (IMP) and other purine nucleotides. NT5C2 is implicated in the maintenance of a constant composition of intracellular purine/pyrimidine nucleotides [75]. |
| BCL2-associated transcription factor 1 (BCLAF1) | BCLAF1, a transcriptional repressor that interacts with members of the BCL2 family of proteins, promotes apoptosis [76]. |
| Human positive cofactor 4 (PC4) | Suppressor of oxidative mutator phenotype [77]. Accumulates at DNA damage foci [78]. |
Finally, the role of SOCS1 as a mediator facilitating the interactions of ATM and ATR with their targets suggests that other interaction partners of SOCS1 could also become the substrates of ATM/ATR-dependent phosphorylation during the DNA damage response. Proteomic analysis of SOCS1 complexes revealed putative interactions with several proteins that play a role in the DNA damage response, apoptosis or oxidative stress pathways (Table I). Future work will determine which functions of SOCS1 apply to every one of its interaction partners: ubiquitination followed by proteolytic degradation or DNA damage stimulated phosphorylation.
Conclusions
Studies on molecular mechanisms underlying cellular senescence have made significant contributions to the discovery of novel regulators of tumor suppressor pathways. Using microarrays or cDNA / siRNA screens, multiple researchers have identified novel regulators of p53 or Rb in controlling tumor formation. Using this approach to study STAT5-induced senescence, we identified SOCS1 as an important activator of the p53 and the DNA damage response. Surprisingly, the SOCS box represents a binding motif for ATM and ATR [34]. To date, about 40 proteins are known to harbor a SOCS box domain. Clearly further work will determine whether SOCS box-containing proteins also participate in the DNA damage response and control oncogenesis.
Acknowledgments
We thank Gillian Vogel for critical reading of the manuscript and helpful suggestions. F.A.M. and G.F. are supported by the Fonds de Recherche en Santé du Québec and V.C. by the Natural Sciences and Engineering Research Council of Canada (NSERC). This work was funded by grants from the Canadian Institutes of Health Research (CIHR MOP82887 to G.F. and MOP84234 to S.I.).
Conflicts of Interest
The authors of this manuscript have no conflict of interests to declare.
References
- 1. Hanada T and Yoshimura A. Regulation of cytokine signaling and inflammation. Cytokine Growth Factor Rev. 2002; 13: 413 -421. [PubMed] .
- 2. Ward AC , Touw I and Yoshimura A. The Jak-Stat pathway in normal and perturbed hematopoiesis. Blood. 2000; 95: 19 -29. [PubMed] .
- 3. Alexander WS Suppressors of cytokine signalling (SOCS) in the immune system. Nat Rev Immunol. 2002; 2: 410 -416. [PubMed] .
- 4. Endo TA , Masuhara M , Yokouchi M , Suzuki R , Sakamoto H , Mitsui K , Matsumoto A , Tanimura S , Ohtsubo M , Misawa H , Miyazaki T , Leonor N and Taniguchi T. A new protein containing an SH2 domain that inhibits JAK kinases. Nature. 1997; 387: 921 -924. [PubMed] .
- 5. Naka T , Narazaki M , Hirata M , Matsumoto T , Minamoto S , Aono A , Nishimoto N , Kajita T , Taga T , Yoshizaki K , Akira S and Kishimoto T. Structure and function of a new STAT-induced STAT inhibitor. Nature. 1997; 387: 924 -929. [PubMed] .
- 6. Starr R , Willson TA , Viney EM , Murray LJ , Rayner JR , Jenkins BJ , Gonda TJ , Alexander WS , Metcalf D , Nicola NA and Hilton DJ. A family of cytokine-inducible inhibitors of signalling. Nature. 1997; 387: 917 -921. [PubMed] .
- 7. Nicholson SE , Willson TA , Farley A , Starr R , Zhang JG , Baca M , Alexander WS , Metcalf D , Hilton DJ and Nicola NA. Mutational analyses of the SOCS proteins suggest a dual domain requirement but distinct mechanisms for inhibition of LIF and IL-6 signal transduction. EMBO J. 1999; 18: 375 -385. [PubMed] .
- 8. Yasukawa H , Misawa H , Sakamoto H , Masuhara M , Sasaki A , Wakioka T , Ohtsuka S , Imaizumi T , Matsuda T , Ihle JN and Yoshimura A. The JAK-binding protein JAB inhibits Janus tyrosine kinase activity through binding in the activation loop. EMBO J. 1999; 18: 1309 -1320. [PubMed] .
- 9. Kamura T , Maenaka K , Kotoshiba S , Matsumoto M , Kohda D , Conaway RC , Conaway JW and Nakayama KI. VHL-box and SOCS-box domains determine binding specificity for Cul2-Rbx1 and Cul5-Rbx2 modules of ubiquitin ligases. Genes Dev. 2004; 18: 3055 -3065. [PubMed] .
- 10. Kamura T , Sato S , Haque D , Liu L , Kaelin WG Jr , Conaway RC and Conaway JW. The Elongin BC complex interacts with the conserved SOCS-box motif present in members of the SOCS, ras, WD-40 repeat, and ankyrin repeat families. Genes Dev. 1998; 12: 3872 -3881. [PubMed] .
- 11. Ungureanu D , Saharinen P , Junttila I , Hilton DJ and Silvennoinen O. Regulation of Jak2 through the ubiquitin-proteasome pathway involves phosphorylation of Jak2 on Y1007 and interaction with SOCS-1. Mol Cell Biol. 2002; 22: 3316 -3326. [PubMed] .
- 12. Kamizono S , Hanada T , Yasukawa H , Minoguchi S , Kato R , Minoguchi M , Hattori K , Hatakeyama S , Yada M , Morita S , Kitamura T , Kato H and Nakayama K. The SOCS box of SOCS-1 accelerates ubiquitin-dependent proteolysis of TEL-JAK2. J Biol Chem. 2001; 276: 12530 -12538. [PubMed] .
- 13. Rui L , Yuan M , Frantz D , Shoelson S and White MF. SOCS-1 and SOCS-3 block insulin signaling by ubiquitin-mediated degradation of IRS1 and IRS2. J Biol Chem. 2002; 277: 42394 -42398. [PubMed] .
- 14. Liu E , Cote JF and Vuori K. Negative regulation of FAK signaling by SOCS proteins. EMBO J. 2003; 22: 5036 -5046. [PubMed] .
- 15. De Sepulveda P , Ilangumaran S and Rottapel R. Suppressor of cytokine signaling-1 inhibits VAV function through protein degradation. J Biol Chem. 2000; 275: 14005 -14008. [PubMed] .
- 16. Mansell A , Smith R , Doyle SL , Gray P , Fenner JE , Crack PJ , Nicholson SE , Hilton DJ , O'Neill LA and Hertzog PJ. Suppressor of cytokine signaling 1 negatively regulates Toll-like receptor signaling by mediating Mal degradation. Nat Immunol. 2006; 7: 148 -155. [PubMed] .
- 17. Rottapel R , Ilangumaran S , Neale C , La Rose J , Ho JM , Nguyen MH , Barber D , Dubreuil P and de Sepulveda P. The tumor suppressor activity of SOCS-1. Oncogene. 2002; 21: 4351 -4362. [PubMed] .
- 18. Fukushima N , Sato N , Sahin F , Su GH , Hruban RH and Goggins M. Aberrant methylation of suppressor of cytokine signalling-1 (SOCS-1) gene in pancreatic ductal neoplasms. Br J Cancer. 2003; 89: 338 -343. [PubMed] .
- 19. Galm O , Yoshikawa H , Esteller M , Osieka R and Herman JG. SOCS-1, a negative regulator of cytokine signaling, is frequently silenced by methylation in multiple myeloma. Blood. 2003; 101: 2784 -2788. [PubMed] .
- 20. Jiang S , Zhang HW , Lu MH , He XH , Li Y , Gu H , Liu MF and Wang ED. MicroRNA-155 functions as an OncomiR in breast cancer by targeting the suppressor of cytokine signaling 1 gene. Cancer Res. 2010; 70: 3119 -3127. [PubMed] .
- 21. Melzner I , Bucur AJ , Bruderlein S , Dorsch K , Hasel C , Barth TF , Leithauser F and Moller P. Biallelic mutation of SOCS-1 impairs JAK2 degradation and sustains phospho-JAK2 action in the MedB-1 mediastinal lymphoma line. Blood. 2005; 105: 2535 -2542. [PubMed] .
- 22. Pichiorri F , Suh SS , Ladetto M , Kuehl M , Palumbo T , Drandi D , Taccioli C , Zanesi N , Alder H , Hagan JP , Munker R , Volinia S and Boccadoro M. MicroRNAs regulate critical genes associated with multiple myeloma pathogenesis. Proc Natl Acad Sci U S A. 2008; 105: 12885 -12890. [PubMed] .
- 23. Sutherland KD , Lindeman GJ , Choong DY , Wittlin S , Brentzell L , Phillips W , Campbell IG and Visvader JE. Differential hyper-methylation of SOCS genes in ovarian and breast carcinomas. Oncogene. 2004; 23: 7726 -7733. [PubMed] .
- 24. Weniger MA , Melzner I , Menz CK , Wegener S , Bucur AJ , Dorsch K , Mattfeldt T , Barth TF and Moller P. Mutations of the tumor suppressor gene SOCS-1 in classical Hodgkin lymphoma are frequent and associated with nuclear phospho-STAT5 accumulation. Oncogene. 2006; 25: 2679 -2684. [PubMed] .
- 25. Yoshikawa H , Matsubara K , Qian GS , Jackson P , Groopman JD , Manning JE , Harris CC and Herman JG. SOCS-1, a negative regulator of the JAK/STAT pathway, is silenced by methylation in human hepatocellular carcinoma and shows growth-suppression activity. Nat Genet. 2001; 28: 29 -35. [PubMed] .
- 26. Evan GI and d'Adda di Fagagna F. Cellular senescence: hot or what. Curr Opin Genet Dev. 2009; 19: 25 -31. [PubMed] .
- 27. Yu H and Jove R. The STATs of cancer--new molecular targets come of age. Nat Rev Cancer. 2004; 4: 97 -105. [PubMed] .
- 28. Mallette FA , Gaumont-Leclerc MF and Ferbeyre G. The DNA damage signaling pathway is a critical mediator of oncogene-induced senescence. Genes Dev. 2007; 21: 43 -48. [PubMed] .
- 29. Mallette FA , Gaumont-Leclerc MF , Huot G and Ferbeyre G. Myc down-regulation as a mechanism to activate the Rb pathway in STAT5A-induced senescence. J Biol Chem. 2007; 282: 34938 -34944. [PubMed] .
- 30. Mallette FA , Moiseeva O , Calabrese V , Mao B , Gaumont-Leclerc MF and Ferbeyre G. Transcriptome analysis and tumor suppressor requirements of STAT5-induced senescence. Ann N Y Acad Sci. 2010; 1197: 142 -151. [PubMed] .
- 31. de Stanchina E , McCurrach ME , Zindy F , Shieh SY , Ferbeyre G , Samuelson AV , Prives C , Roussel MF , Sherr CJ and Lowe SW. E1A signaling to p53 involves the p19(ARF) tumor suppressor. Genes Dev. 1998; 12: 2434 -2442. [PubMed] .
- 32. Zindy F , Eischen CM , Randle DH , Kamijo T , Cleveland JL , Sherr CJ and Roussel MF. Myc signaling via the ARF tumor suppressor regulates p53-dependent apoptosis and immortalization. Genes Dev. 1998; 12: 2424 -2433. [PubMed] .
- 33. Wei W , Hemmer RM and Sedivy JM. Role of p14(ARF) in replicative and induced senescence of human fibroblasts. Mol Cell Biol. 2001; 21: 6748 -6757. [PubMed] .
- 34. Calabrese V , Mallette FA , Deschenes-Simard X , Ramanathan S , Gagnon J , Moores A , Ilangumaran S and Ferbeyre G. SOCS1 links cytokine signaling to p53 and senescence. Mol Cell. 2009; 36: 754 -767. [PubMed] .
- 35. de Stanchina E , Querido E , Narita M , Davuluri RV , Pandolfi PP , Ferbeyre G and Lowe SW. PML is a direct p53 target that modulates p53 effector functions. Mol Cell. 2004; 13: 523 -535. [PubMed] .
- 36. Bourdeau V , Baudry D and Ferbeyre G. PML links aberrant cytokine signaling and oncogenic stress to cellular senescence. Front Biosci. 2009; 14: 475 -485. [PubMed] .
- 37. Ferbeyre G , de Stanchina E , Querido E , Baptiste N , Prives C and Lowe SW. PML is induced by oncogenic ras and promotes premature senescence. Genes Dev. 2000; 14: 2015 -2027. [PubMed] .
- 38. Pearson M , Carbone R , Sebastiani C , Cioce M , Fagioli M , Saito S , Higashimoto Y , Appella E , Minucci S , Pandolfi PP and Pelicci PG. PML regulates p53 acetylation and premature senescence induced by oncogenic Ras. Nature. 2000; 406: 207 -210. [PubMed] .
- 39. Cleghon V and Morrison DK. Raf-1 interacts with Fyn and Src in a non-phosphotyrosine-dependent manner. J Biol Chem. 1994; 269: 17749 -17755. [PubMed] .
- 40. Park I , Chung J , Walsh CT , Yun Y , Strominger JL and Shin J. Phosphotyrosine-independent binding of a 62-kDa protein to the src homology 2 (SH2) domain of p56lck and its regulation by phosphorylation of Ser-59 in the lck unique N-terminal region. Proc Natl Acad Sci U S A. 1995; 92: 12338 -12342. [PubMed] .
- 41. Roe JS , Kim H , Lee SM , Kim ST , Cho EJ and Youn HD. p53 stabilization and transactivation by a von Hippel-Lindau protein. Mol Cell. 2006; 22: 395 -405. [PubMed] .
- 42. Bischof O , Schwamborn K , Martin N , Werner A , Sustmann C , Grosschedl R and Dejean A. The E3 SUMO ligase PIASy is a regulator of cellular senescence and apoptosis. Mol Cell. 2006; 22: 783 -794. [PubMed] .
- 43. Kahyo T , Nishida T and Yasuda H Involvement of PIAS1 in the sumoylation of tumor suppressor p53. Mol Cell. 2001; 8: 713 -718. [PubMed] .
- 44. Megidish T , Xu JH and Xu CW. Activation of p53 by protein inhibitor of activated Stat1 (PIAS1). J Biol Chem. 2002; 277: 8255 -8259. [PubMed] .
- 45. Bartkova J , Rezaei N , Liontos M , Karakaidos P , Kletsas D , Issaeva N , Vassiliou LV , Kolettas E , Niforou K , Zoumpourlis VC , Takaoka M , Nakagawa H and Tort F. Oncogene-induced senescence is part of the tumorigenesis barrier imposed by DNA damage checkpoints. Nature. 2006; 444: 633 -637. [PubMed] .
- 46. Di Micco R , Fumagalli M , Cicalese A , Piccinin S , Gasparini P , Luise C , Schurra C , Garre M , Nuciforo PG , Bensimon A , Maestro R , Pelicci PG and d'Adda di Fagagna F. Oncogene-induced senescence is a DNA damage response triggered by DNA hyper-replication. Nature. 2006; 444: 638 -642. [PubMed] .
- 47. Mallette FA and Ferbeyre G. The DNA damage signaling pathway connects oncogenic stress to cellular senescence. Cell Cycle. 2007; 6: 1831 -1836. [PubMed] .
- 48. Galanty Y , Belotserkovskaya R , Coates J , Polo S , Miller KM and Jackson SP. Mammalian SUMO E3-ligases PIAS1 and PIAS4 promote responses to DNA double-strand breaks. Nature. 2009; 462: 935 -939. [PubMed] .
- 49. Morris JR , Boutell C , Keppler M , Densham R , Weekes D , Alamshah A , Butler L , Galanty Y , Pangon L , Kiuchi T , Ng T and Solomon E. The SUMO modification pathway is involved in the BRCA1 response to genotoxic stress. Nature. 2009; 462: 886 -890. [PubMed] .
- 50. Coppe JP , Patil CK , Rodier F , Sun Y , Munoz DP , Goldstein J , Nelson PS , Desprez PY and Campisi J. Senescence-associated secretory phenotypes reveal cell-nonautonomous functions of oncogenic RAS and the p53 tumor suppressor. PLoS Biol. 2008; 6: 2853 -2868. [PubMed] .
- 51. Kuilman T , Michaloglou C , Vredeveld LC , Douma S , van Doorn R , Desmet CJ , Aarden LA , Mooi WJ and Peeper DS. Oncogene-induced senescence relayed by an interleukin-dependent inflammatory network. Cell. 2008; 133: 1019 -1031. [PubMed] .
- 52. Rodier F , Coppe JP , Patil CK , Hoeijmakers WA , Munoz DP , Raza SR , Freund A , Campeau E , Davalos AR and Campisi J. Persistent DNA damage signalling triggers senescence-associated inflammatory cytokine secretion. Nat Cell Biol. 2009; 11: 973 -979. [PubMed] .
- 53. Acosta JC , O'Loghlen A , Banito A , Guijarro MV , Augert A , Raguz S , Fumagalli M , Da Costa M , Brown C , Popov N , Takatsu Y , Melamed J and d'Adda di Fagagna F. Chemokine signaling via the CXCR2 receptor reinforces senescence. Cell. 2008; 133: 1006 -1018. [PubMed] .
- 54. Wajapeyee N , Serra RW , Zhu X , Mahalingam M and Green MR. Oncogenic BRAF induces senescence and apoptosis through pathways mediated by the secreted protein IGFBP7. Cell. 2008; 132: 363 -374. [PubMed] .
- 55. Kortlever RM , Higgins PJ and Bernards R. Plasminogen activator inhibitor-1 is a critical downstream target of p53 in the induction of replicative senescence. Nat Cell Biol. 2006; 8: 877 -884. [PubMed] .
- 56. Xue W , Zender L , Miething C , Dickins RA , Hernando E , Krizhanovsky V , Cordon-Cardo C and Lowe SW. Senescence and tumour clearance is triggered by p53 restoration in murine liver carcinomas. Nature. 2007; 445: 656 -660. [PubMed] .
- 57. Bekker-Jensen S , Rendtlew Danielsen J , Fugger K , Gromova I , Nerstedt A , Lukas C , Bartek J , Lukas J and Mailand N. HERC2 coordinates ubiquitin-dependent assembly of DNA repair factors on damaged chromosomes. Nat Cell Biol. 2010; 12: 80 -86; sup pp 81-12. [PubMed] .
- 58. Doil C , Mailand N , Bekker-Jensen S , Menard P , Larsen DH , Pepperkok R , Ellenberg J , Panier S , Durocher D , Bartek J , Lukas J and Lukas C. RNF168 binds and amplifies ubiquitin conjugates on damaged chromosomes to allow accumulation of repair proteins. Cell. 2009; 136: 435 -446. [PubMed] .
- 59. Huen MS , Grant R , Manke I , Minn K , Yu X , Yaffe MB and Chen J. RNF8 transduces the DNA-damage signal via histone ubiquitylation and checkpoint protein assembly. Cell. 2007; 131: 901 -914. [PubMed] .
- 60. Kolas NK , Chapman JR , Nakada S , Ylanko J , Chahwan R , Sweeney FD , Panier S , Mendez M , Wildenhain J , Thomson TM , Pelletier L , Jackson SP and Durocher D Orchestration of the DNA-damage response by the RNF8 ubiquitin ligase. Science. 2007; 318: 1637 -1640. [PubMed] .
- 61. Mailand N , Bekker-Jensen S , Faustrup H , Melander F , Bartek J , Lukas C and Lukas J. RNF8 ubiquitylates histones at DNA double-strand breaks and promotes assembly of repair proteins. Cell. 2007; 131: 887 -900. [PubMed] .
- 62. Stewart GS , Panier S , Townsend K , Al-Hakim AK , Kolas NK , Miller ES , Nakada S , Ylanko J , Olivarius S , Mendez M , Oldreive C , Wildenhain J and Tagliaferro A. The RIDDLE syndrome protein mediates a ubiquitin-dependent signaling cascade at sites of DNA damage. Cell. 2009; 136: 420 -434. [PubMed] .
- 63. Koelsche C , Strebovsky J , Baetz A and Dalpke AH. Structural and functional analysis of a nuclear localization signal in SOCS1. Mol Immunol. 2009; 46: 2474 -2480. [PubMed] .
- 64. Baetz A , Koelsche C , Strebovsky J , Heeg K and Dalpke AH. Identification of a nuclear localization signal in suppressor of cytokine signaling 1. FASEB J. 2008; 22: 4296 -4305. [PubMed] .
- 65. Boyd SD , Tsai KY and Jacks T. An intact HDM2 RING-finger domain is required for nuclear exclusion of p53. Nat Cell Biol. 2000; 2: 563 -568. [PubMed] .
- 66. Geyer RK , Yu ZK and Maki CG. The MDM2 RING-finger domain is required to promote p53 nuclear export. Nat Cell Biol. 2000; 2: 569 -573. [PubMed] .
- 67. Li M , Brooks CL , Wu-Baer F , Chen D , Baer R and Gu W. Mono- versus polyubiquitination: differential control of p53 fate by Mdm2. Science. 2003; 302: 1972 -1975. [PubMed] .
- 68. Sekimoto T , Imamoto N , Nakajima K , Hirano T and Yoneda Y. Extracellular signal-dependent nuclear import of Stat1 is mediated by nuclear pore-targeting complex formation with NPI-1, but not Rch1. EMBO J. 1997; 16: 7067 -7077. [PubMed] .
- 69. Limnander A , Danial NN and Rothman PB. v-Abl signaling disrupts SOCS-1 function in transformed pre-B cells. Mol Cell. 2004; 15: 329 -341. [PubMed] .
- 70. Chen JL , Limnander A and Rothman PB. Pim-1 and Pim-2 kinases are required for efficient pre-B-cell transformation by v-Abl oncogene. Blood. 2008; 111: 1677 -1685. [PubMed] .
- 71. Chen XP , Losman JA , Cowan S , Donahue E , Fay S , Vuong BQ , Nawijn MC , Capece D , Cohan VL and Rothman P. Pim serine/threonine kinases regulate the stability of Socs-1 protein. Proc Natl Acad Sci U S A. 2002; 99: 2175 -2180. [PubMed] .
- 72. Srsen V , Gnadt N , Dammermann A and Merdes A. Inhibition of centrosome protein assembly leads to p53-dependent exit from the cell cycle. J Cell Biol. 2006; 174: 625 -630. [PubMed] .
- 73. Trinei M , Giorgio M , Cicalese A , Barozzi S , Ventura A , Migliaccio E , Milia E , Padura IM , Raker VA , Maccarana M , Petronilli V , Minucci S and Bernardi P. A p53-p66Shc signalling pathway controls intracellular redox status, levels of oxidation-damaged DNA and oxidative stress-induced apoptosis. Oncogene. 2002; 21: 3872 -3878. [PubMed] .
- 74. White DE , Negorev D , Peng H , Ivanov AV , Maul GG and Rauscher FJ. 3rd KAP1, a novel substrate for PIKK family members, colocalizes with numerous damage response factors at DNA lesions. Cancer Res. 2006; 66: 11594 -11599. [PubMed] .
- 75. Yamauchi T , Negoro E , Kishi S , Takagi K , Yoshida A , Urasaki Y , Iwasaki H and Ueda T. Intracellular cytarabine triphosphate production correlates to deoxycytidine kinase/cytosolic 5'-nucleotidase II expression ratio in primary acute myeloid leukemia cells. Biochem Pharmacol. 2009; 77: 1780 -1786. [PubMed] .
- 76. Letsas KP , Frangou-Lazaridis M , Skyrlas A , Tsatsoulis A and Malamou-Mitsi V. Transcription factor-mediated proliferation and apoptosis in benign and malignant thyroid lesions. Pathol Int. 2005; 55: 694 -702. [PubMed] .
- 77. Wang JY , Sarker AH , Cooper PK and Volkert MR. The single-strand DNA binding activity of human PC4 prevents mutagenesis and killing by oxidative DNA damage. Mol Cell Biol. 2004; 24: 6084 -6093. [PubMed] .
- 78. Mortusewicz O , Roth W , Li N , Cardoso MC , Meisterernst M and Leonhardt H. Recruitment of RNA polymerase II cofactor PC4 to DNA damage sites. J Cell Biol. 2008; 183: 769 -776. [PubMed] .


