More than a decade ago, our group discovered that dying yeasts do not succumb in an unregulated manner but rather follow a highly orchestrated molecular choreography, which resembles mammalian apoptosis in both morphological and molecular terms. On the one hand, morphological apoptotic markers, such as phosphatidylserine externalization, DNA degradation, chromatin condensation, and the generation of reactive oxygen species (ROS), accompany yeast apoptosis [1,2]. On the other hand, crucial molecular regulators of apoptosis in yeast, including core executors like a caspase [3], the apoptosis-inducing factor [4], endonuclease G [5] or the serine protease OMI [6], as well as pivotal inhibitors like the AAA-ATPase CDC48 (p97/VCP) [7] or the IAP (inhibitor of apoptosis protein) BIR1 [8], have been identified by different groups. Moreover, yeast apoptosis has been causally linked to complex metabolic scenarios such as the Warburg effect [9] or lipotoxicity, a form of cellular demise resulting from lipid overload [10]. Other „classical” apoptosis features connected to dying yeasts are deregulated mitochondrial fission and fusion, cytochrome c release, perturbations of the actin or tubulin cytoskeleton, and epigenetic modifications of the chromatin [11-15].
Research in this area has also provided a teleological explanation for regulated yeast cell death, which a priori should be counterproductive for a unicellular organism, by proving its fundamental role in several physiological scenarios, among others viral infection, meiosis, mating and aging [16-18]. In these scenarios, the death of damaged individual cells yields a selective advantage for the yeast population as a whole [17-19], facilitating the spreading of the clone. This is also the case during chronological aging of yeast cells, a model invented and developed by V. D. Longo in 1996 [20] and defined by the decline of surviving cells in the postmitotic stationary phase, thus simulating the aging of the mostly postmitotic cells of higher organisms. Here, programmed death of old, damaged yeast cells (both by apoptosis and necrosis [17,18,21]) favors the long-term survival of the population. For instance, a strain devoid of the apoptotic machinery or overexpressing superoxide dismutase (and therefore with diminished levels of superoxide) shows an initial advantage in a direct over-time competition assay with a wild type strain; however, it gets finally outcompeted by the wild type strain because it accumulates damaged or unfit cells [17,18]. Programmed cell death seems to clean the population over time, suggesting that aging in yeast (and possibly in higher organisms) may be programmed, since single cells sacrifice themselves for the benefit of the group. In fact, these data may be regarded as the first experimental proof for the so called „group selection theory" as proposed by A. Wallace, in which it is suggested that alleles can become selected because of the benefits they might render to the group, not to the individual [17,22].
Besides such philosophical considerations, the yeast chronological aging system (Figure 1) has led to the discovery of aging mechanisms and anti-aging drugs that have subsequently been confirmed in higher organisms [23]. Examples include branched chain amino acids (BCAA), first found to extend yeast chronological lifespan (CLS) and then confirmed as regulators in mice [24,25] or spermidine, first detected in yeast as an antiaging compound upon external administration and afterwards shown to also prolong life of flies, worms, human immune cells, and, possibly mice [21,26]. CLS extension by rapamycin, was first shown in budding yeast and meanwhile shown to promote longevity in higher eukaryotes (i.e. flies and mice) as well [27-29]. Furthermore, FCCP a mitochondrial uncoupler extended CLS of yeast as well as lifespan of worms [30,31].
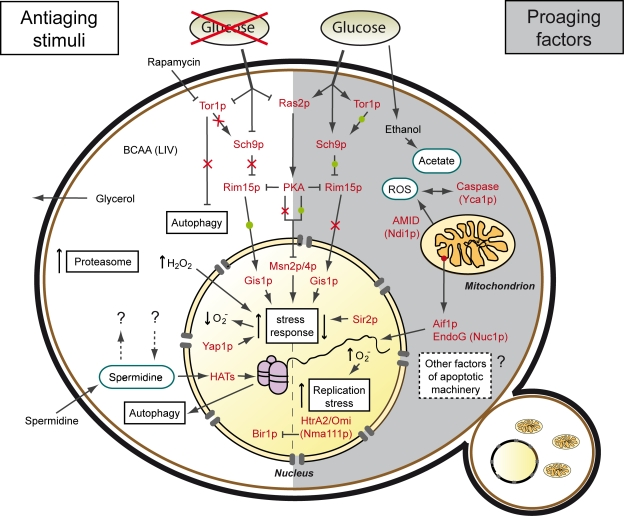
Figure 1. Stimuli and factors involved in yeast chronological aging. The process of chronological aging ultimately relies on a cell's decision to stall or promote its growth in a given scenario. If growth is inhibited, for instance due to low nutrient availability (caloric restriction), the cell enters a state of low metabolic activity (quiescence), thus arresting the aging process (antiaging). If nutrients are available the cell activates growth progression, elevates metabolic rates, promotes its reproduction and progressively ages (non-quiescence or senescence), eventually culminating in its demise (proaging).
Consequently, glucose availability has a major impact on longevity, specifically via the Tor-Sch9p and Ras2-PKA pathways, which require the serine-threonine kinase Rim15p and the transcription factors Gis1p, Msn2p and Msn4p to regulate expression of stress response genes. While longevity is promoted upon low glucose availability (↓glucose → ↓Tor/Sch9p/Ras2 → ↑stress response), it is shortened when glucose availability is high (↑glucose → ↑Tor/Sch9p/Ras2 → ↓stress response). Stress response is also regulated via other factors like the transcription factor Yap1p or the histone deacetylase Sir2.
The Tor1p kinase is also involved in the regulation of autophagy, a self-recycling pathway under nutrient starvation. Upregulation of autophagy may occur via inhibition of the Tor1p kinase (for instance with rapamycin) or elevation of intracellular spermidine levels (by external supplementation or internal regulation mechanisms still to be discerned). Spermidine, which induces autophagy via inhibition of histone acetyl transferases (HATs) and resulting histone hypoacetylation, could also regulate longevity via autophagy-independent alternative pathways. In addition, chronological lifespan is extended by increased availability of branched side chain amino acids (BCAA; leucine, isoleucine and valine) and a rise in synthesis and release of glycerol, which in turn is inhibited by sustained growth signalling (see text).
On the other hand, under conditions where chronological aging is promoted ethanol is metabolized to acetate, which acts as a proaging trigger, in part by influencing internal and external pH as well as TOR signaling. Additionally, growth signaling leads to cell cycle progression and replication stress. Chronological aging-induced cell death has been shown to be regulated by a number of mitochondrial, nuclear and cytosolic lethal effectors and might also involve further factors associated with the yeast apoptotic machinery.
The causative role for ROS as a driving force of the aging process serves as the unifying feature of the CLS model. Fabrizio et al. presented evidence, that the superoxide-induced demise of yeast cells during chronological aging provides the population with nutrients and could favor either spontaneous or specific mutations leading to the so-called adaptive regrowth [17]. Adaptive regrowth occurs in late phases of chronological aging experiments where already 90-99% of the population is dead and describes the growth of better adapted mutants, which is facilitated by nutrients released from dead cells. Interestingly, long-lived mutants (like sch9 or ras2 deletion strains) do not show adaptive regrowth, possibly due to the diminished superoxide production of these strains [17].
In this issue of AGING Weinberger et al. present further evidence supporting the idea of superoxide anions acting as signal molecules that determine yeast CLS [32]. While activation of the major growth signaling pathways (TOR, AKT/Sch9p in Saccharomyces cerevisiae and RAS) by several triggers (e.g. high glucose levels) increases superoxide levels, inactivation of these signaling pathways by caloric restriction (CR) or gene deletions (SCH9 or TOR1) diminishes superoxide levels. CR-effects are in part dependent on Rim15p [33]; in addition, Weinberger et al. could show that Rim15p-independent effects are mediated by H2O2-induced expression of superoxide dismutases, which results in reduced superoxide levels [32,34]. Furthermore, high superoxide levels correlate with the inability to cleanly arrest in G0/G1 growth phase shifting the arrest rather to the S phase, which in turn induces replication stress and hinders quiescence [32]. Quiescence is a prerequisite for postmitotic cell survival [19] and replication stress a known inducer of apoptosis in S. cerevisiae [35].
One of the key triggers of growth signaling used by Weinberger et al. is elevated glucose. Interestingly, exposure to pure glucose in aqueous solution has been shown to induce cell death in stationary yeast cultures [36]. The occurring cell death, which is of apoptotic nature, can be prevented by inhibiting ROS production through addition of ascorbic acid [37]. Intriguingly, Granot and colleagues could also demonstrate that glucose induced early, pre-budding growth events (culminating in faster budding upon transfer to rich media) [36,38], suggesting that apoptotic death can even occur when budding is in preparation, but not visible yet. Given that these conditions (stationary yeast in aqueous glucose solution) possibly resemble later phases of chronological aging experiments with glucose excess, the superoxide-based mechanism proposed by Weinberger et al. [32] might underlie the effects observed by Granot et al. [37].
The understanding of the proposed superoxide signal cascade will require clarifying the molecular connection between growth signaling pathways and the generation of superoxide. Notably, superoxide is a byproduct naturally occurring during respiration and other enzymatic reactions [39]. Mitochondria, well-established executive organelles in both apoptosis and aging [40-42], represent the main sites for superoxide leakage, in particular complex I (in yeast consisting only of NADH-ubiquinone oxidoreductases) and complex III of the respiratory chain [43]. The increase in superoxide levels accompanying stable growth signaling could, thus, be an unavoidable byproduct of enzymatic reactions (i.e. during limited respiration). However, it is also conceivable that under such circumstances superoxide might be produced specifically as a signaling molecule by appropriate enzymes (e.g. NAD(P)H oxidases).
In budding yeast administration of glucose leads to drastically diminished respiratory activity due to a very effective glucose repression circuit [44,45]. Albeit, NADH (that is generated during the conversion of sugar into biomass) and possibly other upstream metabolites for reductive potential and use in respiration will still lead to basic respiratory activity and could derive in a saturation of the electron transport capacity under conditions, where low expression of respiratory chain components limit respiration. While overall ROS production (via respiration) should remain rather low, specific superoxide generation could occur. Interestingly, the group around G. S. Schadel has previously published that TOR signaling and Sch9p kinase are downstream effectors of respiration and ROS production [46]. Limiting TOR signaling during glucose availability results in enhanced O2 consumption (without rising ATP levels) and elevated expression of respiratory complexes but diminished ROS production during early time points [46]. These observations might support the above mentioned concept of specific superoxide generation under conditions of glucose repression during early growth phases in an aerobic environment.
Under anaerobic conditions glycolytically generated NADH cannot be deployed to respiration and thus is re-oxidized by generating glycerol, a reaction that consumes reductive potential by NADH oxidation [47,48]. Intriguingly, Wei et al. could show that the sch9 long-lived deletion mutant, which generates less superoxide, produces and exports glycerol under aerobic growth conditions when glucose or ethanol are available, thus generating a CR-like environment [49]. In this case - taking together the aforementioned considerations - NADH would be redirected from being used in respiration and could account, in part, for the lowered ROS accumulation found in this mutant strain. This is in line with the observed altered respiration phenotype of sch9 mutants, whose respiration is higher during early [46] and lower during later growth phases [49] as compared to the wild type: while during early growth (high levels of glucose) an overload of electrons in the respiratory chain would be circumvented, thus making respiration more efficient in comparison to the wild type, overall NADH availability for respiration would be reduced in later growth phases. It is tempting to speculate that glycerol production and release by long-lived mutant yeast strains reflects a general concept also important for higher organisms. In that sense it is remarkable that worms fed on glucose present reduced longevity likely due to a down-regulation of a specific aquaporin glycerol-tansport channel [50].
Of note, in a different yeast model with continuous high growth rates (grown as colonies on agar plates) cells can be protected from ROS-induced apoptosis in early growth phases by eliminating the onset of respiration either via higher glucose levels or via deletions resulting in loss of respiration competence [9]. Additionally, abrupt induction of respiration in highly proliferative cells (i.e. by shifting exponentially growing yeast cells on respiratory media) results in ROS-mediated apoptosis that can be avoided by addition of glutathione [9]. These results emphasize the importance of different growth models and hint to different roles for ROS during different growth phases and scenarios.
Besides its actual production, both the cellular localization of superoxide and the mechanism signaling an S phase rather than an G0/G1 phase arrest ultimately driving cells into replication stress are of great interest. ROS have been known for a long time to induce cell cycle alterations; however, focus was directed towards secondary effects mediating cellular damage [51]. Intriguingly, recent publications ascribe ROS cell cycle-associated signaling attributes: for instance, Hole et al. [52] recently described that Ras-induced ROS generation by an NADPH oxidase promotes growth factor-independent proliferation in human CD34+ hematopoietic progenitor cells and Cova et al. [53] evidenced that alteration of cell cycle in Amyotrophic Lateral Sclerosis disease is linked to mutated Superoxide dismutase 1.
The results by Weinberger et al. suggest that different ROS trigger specific effects, making it particularly important to accurately distinguish them and their downstream activity (either as damaging agent or signal molecule). At the same time, Weinberger et al. also provide evidence that acetic acid production (and/or resulting pH alterations), which, in some settings is able to restrict yeast CLS [54-56], can lead to sustained growth signaling. This observation could have theoretical and practical implications for a variety of human diseases [55]. For example, blood pH, which declines in humans with ongoing aging due to renal insufficiencies, can be influenced by our diet. In fact, modern diets could lead to a low-grade systemic metabolic acidosis [57,58].
Metabolic homeostasis, which can be regulated by autophagy during chronological aging, is a key component of longevity. Intriguingly, main signaling pathways involved in autophagy (e.g. TOR) were shown by Weinberger et al. to modulate superoxide levels [32]. Consistently, lifespan extension by diminished TOR/Sch9p signaling leads to decreased superoxide levels [32,59] and elevated autophagy [24]. Our group could demonstrate that addition of spermidine (a natural occurring polyamine) drastically extends CLS dependent on induction of autophagy and accompanied by elevated fractions of quiescent cells. The longevity-promoting effects of spermidine addition were absent in autophagy deficient cells in a pH-independent manner [21]. Furthermore, cultivation of yeast in the presence of rapamycin elongates lifespan in an autophagy-dependent manner as shown by Alvers et al. [60]. In these experiments, rapamycin was added right after inoculation of yeast cultures emphasizing that early onset of autophagic events might already pave the impact on longevity.
It will be interesting to see if autophagy and superoxide effects are strictly parallel, but cooperative events in promoting longevity, or if superoxides have a direct effect on the regulation and/or occurrence of autophagy and vice versa. Indeed, a number of works have shown that ROS stress is an important inducer and regulator of autophagy [61]. For instance, Atg4p (an essential protease of the autophagic machinery) is a direct target for oxidation by H2O2 [62].
After the pioneering work of Longo and colleagues [17,20,59] on the role of superoxide in yeast CLS, Weinberger et al. now reinforces the importance of superoxide and provides exciting new aspects of its role in determining CLS [32]. It now needs to be clarified how it is interconnected to other known parts of the regulatory picture determining (yeast) lifespan. Given that high glucose uptake also negatively influences lifespan of higher organisms like Caenorhabditis elegans [31,50] accompanied by elevated superoxide levels [31], these results may well open doors to gain regulatory insights into how our diet influences our lifespan and our health.
Acknowledgments
We are grateful to the Austrian Science Fund FWF (Austria) for grant SFB LIPOTOX, and to the European Commission for project APOSYS.
Conflicts of Interest
The authors of this manuscript have no conflict of interests to declare.
References
- 1. Carmona-Gutierrez D, Eisenberg T, Buttner S, Meisinger C, Kroemer G, Madeo F. Apoptosis in yeast: triggers, pathways, subroutines. Cell Death Differ. 2010; 17: 763 -773. [PubMed] .
- 2. Madeo F, Frohlich E, Ligr M, Grey M, Sigrist SJ, Wolf DH, Frohlich K. U. Oxygen stress: a regulator of apoptosis in yeast. J Cell Biol. 1999; 145: 757 -67. [PubMed] .
- 3. Madeo F, Herker E, Maldener C, Wissing S, Lachelt S, Herlan M, Fehr M, Lauber K, Sigrist SJ, Wesselborg S, Frohlich KU. A caspase-related protease regulates apoptosis in yeast. Mol Cell. 2002; 9: 911 -917. [PubMed] .
- 4. Wissing S, Ludovico P, Herker E, Buttner S, Engelhardt SM, Decker T, Link A, Proksch A, Rodrigues F, Corte-Real M, Frohlich KU, Manns J, Cande C, Sigrist SJ, Kroemer G, Madeo F. An AIF orthologue regulates apoptosis in yeast. J Cell Biol. 2004; 166: 969 -974. [PubMed] .
- 5. Buttner S, Eisenberg T, Carmona-Gutierrez D, Ruli D, Knauer H, Ruckenstuhl C, Sigrist C, Wissing S, Kollroser M, Frohlich KU, Sigrist S, Madeo F. Endonuclease G regulates budding yeast life and death. Mol Cell. 2007; 25: 233 -246. [PubMed] .
- 6. Fahrenkrog B, Sauder U, Aebi U. The S. cerevisiae HtrA-like protein Nma111p is a nuclear serine protease that mediates yeast apoptosis. J Cell Sci. 2004; 117: 115 -26. [PubMed] .
- 7. Madeo F, Frohlich E, Frohlich KU. A yeast mutant showing diagnostic markers of early and late apoptosis. J Cell Biol. 1997; 139: 729 -734. [PubMed] .
- 8. Walter D, Wissing S, Madeo F, Fahrenkrog B. The inhibitor-of-apoptosis protein Bir1p protects against apoptosis in S. cerevisiae and is a substrate for the yeast homologue of Omi/HtrA2. J Cell Sci. 2006; 119: 1843 -1851. [PubMed] .
- 9. Ruckenstuhl C, Buttner S, Carmona-Gutierrez D, Eisenberg T, Kroemer G, Sigrist SJ, Frohlich KU, Madeo F. The Warburg effect suppresses oxidative stress induced apoptosis in a yeast model for cancer. PLoS One. 2009; 4: e4592 [PubMed] .
- 10. Rockenfeller P, Ring J, Muschett V, Beranek A, Buettner S, Carmona-Gutierrez D, Eisenberg T, Khoury C, Rechberger G, Kohlwein SD, Kroemer G, Madeo F. Fatty acids trigger mitochondrion-dependent necrosis. Cell Cycle. 2010; 9: 2836 -2842. [PubMed] .
- 11. Ahn SH, Cheung WL, Hsu JY, Diaz RL, Smith MM, Allis CD. Sterile 20 kinase phosphorylates histone H2B at serine 10 during hydrogen peroxide-induced apoptosis in S. cerevisiae. Cell. 2005; 120: 25 -36. [PubMed] .
- 12. Ahn SH, Diaz RL, Grunstein M, Allis CD. Histone H2B deacetylation at lysine 11 is required for yeast apoptosis induced by phosphorylation of H2B at serine 10. Mol Cell. 2006; 24: 211 -220. [PubMed] .
- 13. Fannjiang Y, Cheng WC, Lee SJ, Qi B, Pevsner J, McCaffery JM, Hil RB, Basanez G, Hardwick JM. Mitochondrial fission proteins regulate programmed cell death in yeast. Genes Dev. 2004; 18: 2785 -2797. [PubMed] .
- 14. Gourlay CW, Carpp LN, Timpson P, Winder SJ, Ayscough KR. A role for the actin cytoskeleton in cell death and aging in yeast. J Cell Biol. 2004; 164: 803 -809. [PubMed] .
- 15. Ludovico P, Rodrigues F, Almeida A, Silva MT, Barrientos A, Corte-Real M. Cytochrome c release and mitochondria involvement in programmed cell death induced by acetic acid in Saccharomyces cerevisiae. Mol Biol Cell. 2002; 13: 2598 -2606. [PubMed] .
- 16. Buttner S, Eisenberg T, Herker E, Carmona-Gutierrez D, Kroemer G, Madeo F. Why yeast cells can undergo apoptosis: death in times of peace, love, and war. J Cell Biol. 2006; 175: 521 -525. [PubMed] .
- 17. Fabrizio P, Battistella L, Vardavas R, Gattazzo C, Liou LL, Diaspro A, Dossen JW, Gralla EB, Longo VD. Superoxide is a mediator of an altruistic aging program in Saccharomyces cerevisiae. J Cell Biol. 2004; 166: 1055 -67. [PubMed] .
- 18. Herker E, Jungwirth H, Lehmann KA, Maldener C, Frohlich KU, Wissing S, Buttner S, Fehr M, Sigrist S, Madeo F. Chronological aging leads to apoptosis in yeast. J Cell Biol. 2004; 164: 501 -507. [PubMed] .
- 19. Allen C, Buttner S, Aragon AD, Thomas JA, Meirelles O, Jaetao JE, Benn D, Ruby SW, Veenhuis M, Madeo F, Werner-Washburne M. Isolation of quiescent and nonquiescent cells from yeast stationary-phase cultures. J Cell Biol. 2006; 174: 89 -100. [PubMed] .
- 20. Longo VD, Gralla EB, Valentine JS. Superoxide dismutase activity is essential for stationary phase survival in Saccharomyces cerevisiae. Mitochondrial production of toxic oxygen species in vivo. J Biol Chem. 1996; 271: 12275 -12280. [PubMed] .
- 21. Eisenberg T, Knauer H, Schauer A, Buttner S, Ruckenstuhl C, Carmona-Gutierrez D, Ring J, Schroeder S, Magnes C, Antonacci L, Fussi H, Deszcz L, et al. Induction of autophagy by spermidine promotes longevity. Nat Cell Biol. 2009; 11: 1305 -1314. [PubMed] .
- 22. Rose MR. 1991; Evolutionary Biology of Aging. New York Oxford University Press .
- 23. Fontana L, Partridge L, Longo VD. Extending healthy life span--from yeast to humans. Science. 2010; 328: 321 -326. [PubMed] .
- 24. Alvers AL, Fishwick LK, Wood MS, Hu D, Chung HS, Dunn WA Jr, Aris JP. Autophagy and amino acid homeostasis are required for chronological longevity in Saccharomyces cerevisiae. Aging Cell. 2009; 8: 353 -369. [PubMed] .
- 25. D'Antona G, Ragni M, Cardile A, Tedesco L, Dossena M, Bruttini F, Caliaro F, Corsetti G, Bottinelli R, Carruba MO, Valerio A, Nisoli E. Branched-chain amino acid supplementation promotes survival and supports cardiac and skeletal muscle mitochondrial biogenesis in middle-aged mice. Cell Metab. 2010; 12: 362 -372. [PubMed] .
- 26. Soda K, Dobashi Y, Kano Y, Tsujinaka S, Konishi F. Polyamine-rich food decreases age-associated pathology and mortality in aged mice. Exp Gerontol. 2009; 44: 727 -732. [PubMed] .
- 27. Harrison DE, Strong R, Sharp ZD, Nelson JF, Astle CM, Flurkey K, Nadon NL, Wilkinson JE, Frenkel K, Carter CS, Pahor M, Javors MA, Fernandez E, Miller RA. Rapamycin fed late in life extends lifespan in genetically heterogeneous mice. Nature. 2009; 460: 392 -395. [PubMed] .
- 28. Bjedov I, Toivonen JM, Kerr F, Slack C, Jacobson J, Foley A, Partridge L. Mechanisms of life span extension by rapamycin in the fruit fly Drosophila melanogaster. Cell Metab. 2010; 11: 35 -46. [PubMed] .
- 29. Powers RW 3rd, Kaeberlein M, Caldwell SD, Kennedy BK, Fields S. Extension of chronological life span in yeast by decreased TOR pathway signaling. Genes Dev. 2006; 20: 174 -184. [PubMed] .
- 30. Longo VD, Liou LL, Valentine JS, Gralla EB. Mitochondrial superoxide decreases yeast survival in stationary phase. Arch Biochem Biophys. 1999; 365: 131 -142. [PubMed] .
- 31. Schlotterer A, Kukudov G, Bozorgmehr F, Hutter H, Du X, Oikonomou D, Ibrahim Y, Pfisterer F, Rabbani N, Thornalley P, Sayed A, Fleming T, et al. C. elegans as model for the study of high glucose- mediated life span reduction. Diabetes. 2009; 58: 2450 -2406. [PubMed] .
- 32. Weinberger M, Mesquita A, Carroll T, Marks L, Yang H, Zhang Z, Ludovico P, Burhans WC. Growth signaling promotes chronological aging in budding yeast by inducing superoxide anions that inhibit quiescence. Aging. 2010; 2: this issue .
- 33. Wei M, Fabrizio P, Hu J, Ge H, Cheng C, Li L, Longo VD. Life span extension by calorie restriction depends on Rim15 and transcription factors downstream of Ras/PKA, Tor, and Sch9. PLoS Genet. 2008; 4: e13 [PubMed] .
- 34. Mesquita A, Weinberger M, Silva A, Sampaio-Marques B, Almeida B, Leao C, Costa V, Rodrigues F, Burhans WC, Ludovico P. Caloric restriction or catalase inactivation extends yeast chronological lifespan by inducing H2O2 and superoxide dismutase activity. Proc Natl Acad Sci U S A. 2010; 107: 15123 -15128. [PubMed] .
- 35. Weinberger M, Ramachandran L, Feng L, Sharma K, Sun X, Marchetti M, Huberman JA, Burhans WC. Apoptosis in budding yeast caused by defects in initiation of DNA replication. J Cell Sci. 2005; 118: 3543 -3553. [PubMed] .
- 36. Granot D and Snyder M. Glucose induces cAMP-independent growth-related changes in stationary-phase cells of Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 1991; 88: 5724 -5728. [PubMed] .
- 37. Granot D, Levine A, Dor-Hefetz E. Sugar-induced apoptosis in yeast cells. FEMS Yeast Res. 2003; 4: 7 -13. [PubMed] .
- 38. Granot D and Snyder M. Carbon source induces growth of stationary phase yeast cells, independent of carbon source metabolism. Yeast. 1993; 9: 465 -479. [PubMed] .
- 39. Brand MD. The sites and topology of mitochondrial superoxide production. Exp Gerontol. 2010; 45: 466 -472. [PubMed] .
- 40. Zamzami N, Marchetti P, Castedo M, Decaudin D, Macho A, Hirsch T, Susin SA, Petit PX, Mignotte B, Kroemer G. Sequential reduction of mitochondrial transmembrane potential and generation of reactive oxygen species in early programmed cell death. J Exp Med. 1995; 182: 367 -377. [PubMed] .
- 41. Zamzami N, Susin SA, Marchetti P, Hirsch T, Gomez-Monterrey I, Castedo M, Kroemer G. Mitochondrial control of nuclear apoptosis. J Exp Med. 1996; 183: 1533 -1544. [PubMed] .
- 42. Eisenberg T, Buttner S, Kroemer G, Madeo F. The mitochondrial pathway in yeast apoptosis. Apoptosis. 2007; 12: 1011 -1023. [PubMed] .
- 43. Murphy MP. How mitochondria produce reactive oxygen species. Biochem J. 2009; 417: 1 -13. [PubMed] .
- 44. De Deken RH. The Crabtree effect: a regulatory system in yeast. J Gen Microbiol. 1966; 44: 149 -156. [PubMed] .
- 45. Piskur J, Rozpedowska E, Polakova S, Merico A, Compagno C. How did Saccharomyces evolve to become a good brewer? Trends Genet. 2006; 22: 183 -186. [PubMed] .
- 46. Pan Y and Shadel GS. Extension of chronological life span by reduced TOR signaling requires down-regulation of Sch9p and involves increased mitochondrial OXPHOS complex density. Aging. 2009; 1: 131 -145. [PubMed] .
- 47. Pronk JT, Yde Steensma H, Van Dijken JP. Pyruvate metabolism in Saccharomyces cerevisiae. Yeast. 1996; 12: 1607 -1633. [PubMed] .
- 48. Weusthuis RA, Visser W, Pronk JT, Scheffers WA, van Dijken JP. Effects of oxygen limitation on sugar metabolism in yeasts: a continuous-culture study of the Kluyver effect. Microbiology. 1994; 140: (Pt 4) 703 -715. [PubMed] .
- 49. Wei M, Fabrizio P, Madia F, Hu J, Ge H, Li LM, Longo VD. Tor1/Sch9-regulated carbon source substitution is as effective as calorie restriction in life span extension. PLoS Genet. 2009; 5: e1000467 [PubMed] .
- 50. Lee SJ, Murphy CT, Kenyon C. Glucose shortens the life span of C. elegans by downregulating DAF-16/FOXO activity and aquaporin gene expression. Cell Metab. 2009; 10: 379 -391. [PubMed] .
- 51. Shackelford RE, Kaufmann WK, Paules RS. Oxidative stress and cell cycle checkpoint function. Free Radic Biol Med. 2000; 28: 1387 -1404. [PubMed] .
- 52. Hole PS, Pearn L, Tonks AJ, James PE, Burnett AK, Darley RL, Tonks A. Ras-induced reactive oxygen species promote growth factor-independent proliferation in human CD34+ hematopoietic progenitor cells. Blood. 2010; 115: 1238 -1246. [PubMed] .
- 53. Cova E, Ghiroldi A, Guareschi S, Mazzini G, Gagliardi S, Davin A, Bianchi M, Ceroni M, Cereda C. G93A SOD1 alters cell cycle in a cellular model of Amyotrophic Lateral Sclerosis. Cell Signal. 2010; 22: 1477 -1484. [PubMed] .
- 54. Almeida B, Ohlmeier S, Almeida AJ, Madeo F, Leao C, Rodrigues F, Ludovico P. Yeast protein expression profile during acetic acid-induced apoptosis indicates causal involvement of the TOR pathway. Proteomics. 2009; 9: 720 -732. [PubMed] .
- 55. Burhans WC and Weinberger M. Acetic acid effects on aging in budding yeast: are they relevant to aging in higher eukaryotes? Cell Cycle. 2009; 8: 2300 -2302. [PubMed] .
- 56. Burtner CR, Murakami CJ, Kennedy BK, Kaeberlein M. A molecular mechanism of chronological aging in yeast. Cell Cycle. 2009; 8: 1256 -1270. [PubMed] .
- 57. Frassetto L, Morris RC Jr, Sellmeyer DE, Todd K, Sebastian A. Diet, evolution and aging--the pathophysiologic effects of the post-agricultural inversion of the potassium-to-sodium and base-to-chloride ratios in the human diet. Eur J Nutr. 2001; 40: 200 -213. [PubMed] .
- 58. Frassetto LA, Morris RC Jr, Sebastian A. Effect of age on blood acid-base composition in adult humans: role of age-related renal functional decline. Am J Physiol. 1996; 271: F1114 -1122. [PubMed] .
- 59. Fabrizio P, Pozza F, Pletcher SD, Gendron CM, Longo VD. Regulation of longevity and stress resistance by Sch9 in yeast. Science. 2001; 292: 288 -290. [PubMed] .
- 60. Alvers AL, Wood MS, Hu D, Kaywell AC, Dunn WA Jr, Aris JP. Autophagy is required for extension of yeast chronological life span by rapamycin. Autophagy. 2009; 5: 847 -849. [PubMed] .
- 61. Scherz-Shouval R and Elazar Z. ROS, mitochondria and the regulation of autophagy. Trends Cell Biol. 2007; 17: 422 -427. [PubMed] .
- 62. Scherz-Shouval R, Shvets E, Fass E, Shorer H, Gil L, Elazar Z. Reactive oxygen species are essential for autophagy and specifically regulate the activity of Atg4. EMBO J. 2007; 26: 1749 -1760. [PubMed] .


