Introduction
Commensal microbes with symbiotic relationships with their hosts have been actively investigated in several fields of research in attempts to elucidate their interaction with the host. In particular, many studies have been initiated since the development and wide application of metagenomic sequencing analysis. In recent years, studies on the treatment and prevention of diseases through the production of changes in the host’s intestinal environment have been carried out, and many research teams have suggested that the intestinal microbiota can affect the host’s metabolic diseases [1], immune diseases [2], development [3, 4], reproduction [5], biorhythms [6], and even behavior and mood [7].
Studies into the roles of microbiota as they relate to host aging and lifespan have also been conducted. In Caenorhabditis elegans grown axenically, developmental time and lifespan were about twice as long as those in control worms [8]. Furthermore, the use of microorganisms as food items and the secondary metabolites secreted by the commensal microbes are reported to affect the lifespan of C. elegans [9, 10]. The viability of axenic animals suggests that the presence of microbes in the gut is neither obligatory nor essential to the host’s development, under appropriate conditions. However, conflicting results have been reported on the effects of commensal microbe presence on the lifespan of Drosophila melanogaster. Brummel et al. observed that the lifespan of flies reared axenically following egg bleaching or antibiotic treatment was shorter than that of conventionally reared fly, and that effect could be recovered by exposure of the flies to microbes within 2–3 days from eclosion [11]. In contrast, Yamada et al. did not show an alteration of lifespan in axenic condition flies, while Clark et al., Petkau et al., Galenza et al., Tefit et al. and Obata et al. showed that the absence of commensal microbes extends the lifespan in Drosophila [14–19]. Ren et al., Ridley et al., and Iatsenko et al. showed increased lifespan of Drosophila following egg bleaching or antibiotic treatment, but the increases did not reach statistical significance [12, 13, 20]. To clarify the reasons for these inconsistencies, we generated axenic flies using highly elaborate well-controlled methods and observed that elimination of commensal microbes without detrimental side effects increased host lifespan. Moreover, we observed that an age-related increase in microbial load significantly decreased host lifespan, and a change in microbial load had a more critical effect than that from an age-related change in microbial composition.
Results
Lifespan of axenic D. melanogaster from bleached eggs
To clarify the effect of an absence of commensal microbes on host lifespan, we generated axenic (Ax) flies of laboratory wild-type strain w1118 D. melanogaster by using a sodium hypochlorite-based bleaching method as previously reported [15]. The mean lifespan of the first generation (1G) Ax flies hatched from bleached eggs was shorter than that of conventionally reared (Conv) flies (Figure 1A and 1B, Conv fly 59.92 ± 1.52 days; 1G Ax fly 53.83 ± 1.31 days, 10.16% decrease, log-rank test, χ2 = 38.85, p < 0.0001, Wilcoxon test, χ2 = 20.81, p < 0.0001). Because the bleaching process might have a deleterious effect on the health of flies, we measured the lifespan of second- (2G) and third- (3G) generation flies after first-generation bleaching. Interestingly, the 2G and 3G Ax flies had increased lifespans compared to that of Conv flies (Figure 1A and 1B, Supplementary Table 1; 2G Ax fly 68.52 ± 0.94 days, 14.35% increase, log-rank test, χ2 = 0.08, p = 0.77, Wilcoxon test, χ2 = 8.91, p < 0.005; 3G Ax fly 70.73 ± 0.85 days, 18.04% increase, log-rank test, χ2 = 8.20, p < 0.005, Wilcoxon test, χ2 = 18.93, p < 0.0001), suggesting that the bleaching method may have a detrimental effect on D. melanogaster, but the effect diminished over subsequent generations. To confirm whether the lifespan extension by egg bleaching is due to the absence of commensal microbes, we introduced microbes from 10-day-old Conv flies to 3-day-old 3G Ax flies by using fecal transplantation. As expected, the longevity effect of bacterial removal in Ax flies was diminished by fecal microbe transplantation (Figure 1C and 1D, Supplementary Table 1; Ax 73.00 ± 1.40 days; Ax + FecesConv 68.15 ± 1.30 days, 6.64% crease, log-rank test, χ2 = 22.61, p < 0.0001). These results indicate that the bleaching of eggs can have an adverse effect on adult fly lifespan, but the elimination of commensal microbes increases the lifespan of subsequent generations of flies.
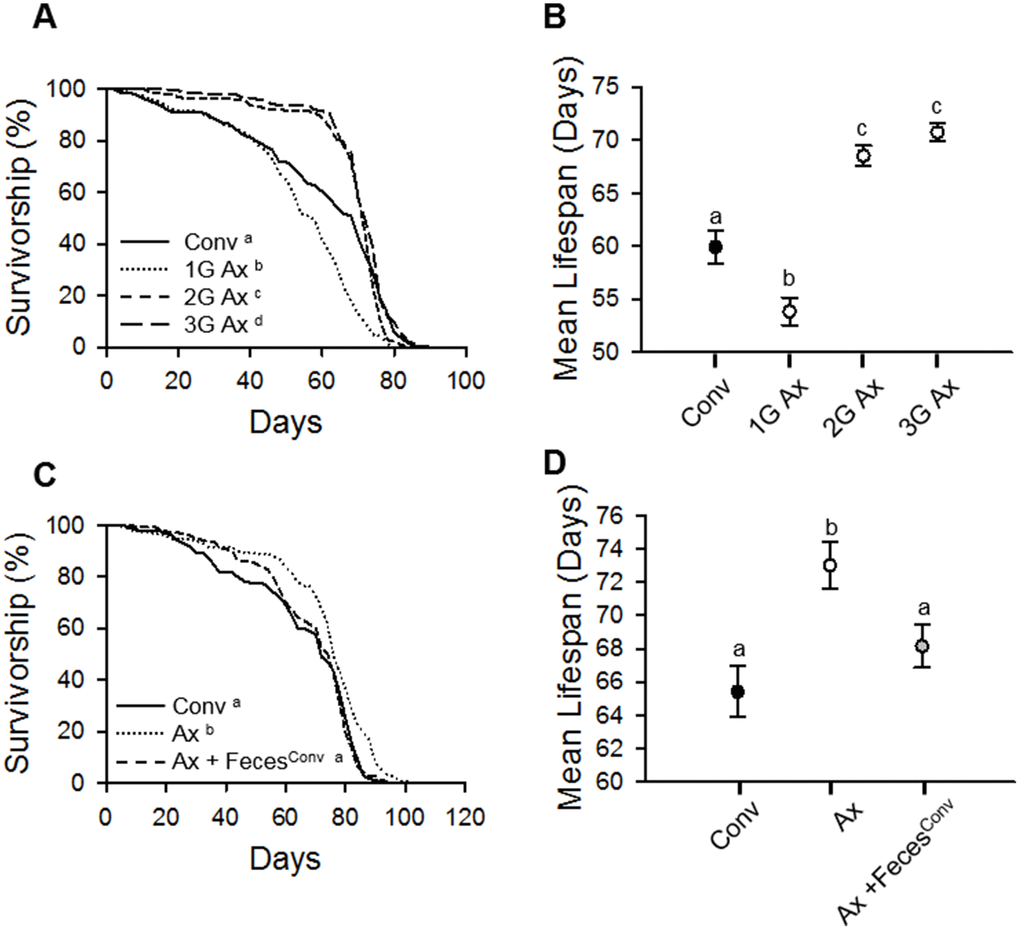
Figure 1. Elimination of commensal microbes by egg bleaching extends host lifespan. (A) Survival of first (1G), second (2G), and third (3G) generations of Ax flies from bleached eggs. (B) Mean lifespan of Conv fly and 1G, 2G, and 3G of Ax flies from bleached eggs. Filled dot, Conv flies; open dots, Ax flies. (C) Survival of Ax flies treated with feces from 10-day-old Conv flies. (D) Mean lifespan of Ax flies treated with feces from 10-day-old Conv flies. Black dot, Conv flies; white dot, Ax flies fed feces from Ax flies; gray dot, Ax flies fed feces from Conv (Ax + FecesConv) flies. Different letters indicate significant differences between groups. Error bars represent the SEM.
Lifespan of axenic D. melanogaster generated by antibiotic treatment
To confirm the effect of microbe removal on host lifespan, we supplied an antibiotic (AB) cocktail to Conv flies as described previously [12]. In our laboratory, bacteria were colonized into the guts of 10-day-old control flies at densities of up to 1.8 × 102 cells per fly; colonization was increased with fly age to densities of up to 2.2 × 105 cells per fly in 50-day-old flies (Figure 2A–2C). Treatment with the AB cocktail reduced the number of bacteria in the gut, and the AB effects were gradually decreased by dilution of the cocktail (Figure 2A–2C). Following AB treatment (×1 AB), the mean lifespans of Conv flies were not significantly different from that of untreated (×0 AB) Conv flies (Figure 2D and 2F, Supplementary Table 1; Conv + ×0 AB 62.73 ± 1.32 days; Conv + ×1.0 AB 62.89 ± 1.21 days, log-rank test, χ2 = 0.53, p = 0.47). However, AB cocktail treatment dramatically reduced the lifespan of Ax flies at the ×1 AB concentration (Figure 2E and 2F, Supplementary Table 1; Ax + ×0 AB 77.17 ± 1.32 days; Ax + ×1.0 AB 65.94 ± 1.34 days, 14.55% decrease, log-rank test, χ2 = 64.01, p < 0.0001). We assumed that the AB cocktail concentration used might have had a toxic effect on fly health. To investigate this possibility, we treated flies with the AB cocktail at 2- and 10-fold concentration dilutions (×0.5 AB and ×0.1 AB, respectively). The ×0.5 AB and ×0.1 AB cocktail treatments did not significantly affect the lifespan of Ax flies (Figure 2E and 2F, Supplementary Table 1; Ax + ×0.1 AB 77.44 ± 1.34 days, log-rank test, χ2 = 1.41, p = 0.23; Ax + ×0.5 AB 77.97 ± 1.06 days, log-rank test, χ2 = 0.64, p = 0.42), whereas the lifespan of Conv flies was increased at these lower AB concentrations (Figure 2D and 2F, Supplementary Table 1; Conv + ×0.1 AB 68.93 ± 1.16 days, 9.88% increase, log-rank test, χ2 = 12.33, p < 0.001; Conv + ×0.5 AB 67.84 ± 0.99 days, log-rank test, χ2 = 2.86, p = 0.09). These results indicate that an AB treatment at a dosage that did not have toxic effects can increase the lifespan of D. melanogaster, which is consistent with the results obtained by using the bleaching method to produce Ax flies.
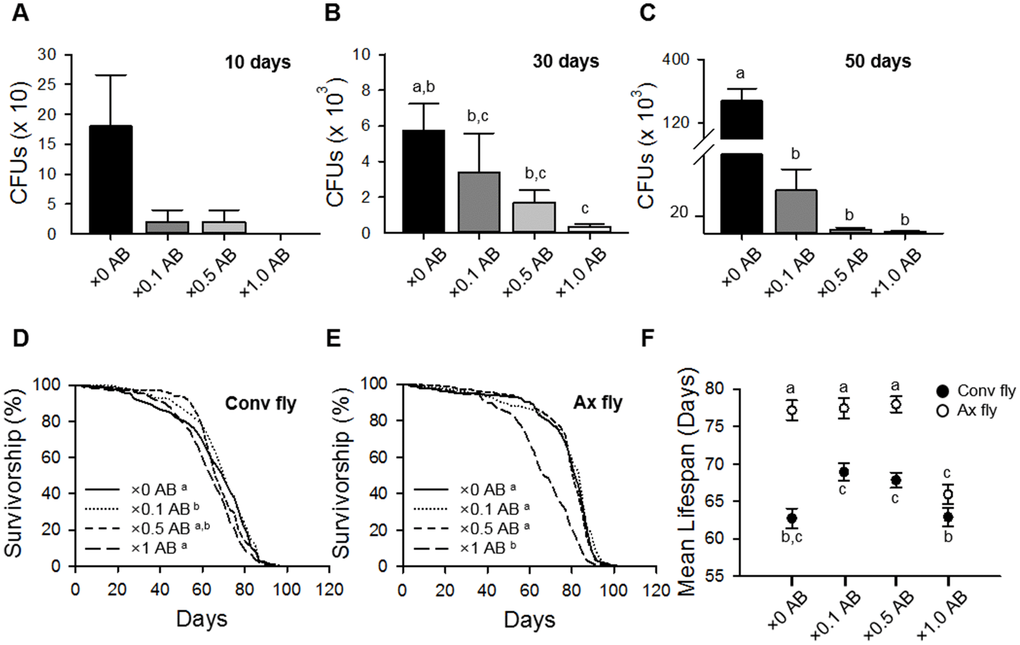
Figure 2. Elimination of commensal microbe by antibiotic treatment extends host lifespan. (A–C) Colony-forming units (CFUs) of Conv flies treated with ×0.1, ×0.5, or ×1 of the antibiotic cocktail AB for 10 days (A), 30 days (B), and 50 days (C). (D–E) Survival of flies treated with ×0.1, ×0.5, or ×1 AB in Conv fly (D) and Ax fly (E). (F) Mean lifespan of flies treated with ×0.1, ×0.5, or ×1 AB. Filled dots, Conv flies; open dots, Ax flies. Different letters indicate significant differences between groups. Error bars represent the SEM.
Strain-specific longevity effect of commensal microbe elimination
Han et al. (2017) reported that Drosophila with different genetic backgrounds had different microbial flora, even though the flies were reared under the same conditions in the same laboratory [21]. Since a different microbial flora may lead to different lifespan results following bacterial removal, we compared the lifespan of Ax flies generated from different strains in our laboratory; wild-type Oregon-R, Canton-S, and w1118 strains. Under the same conditions, the load levels of commensal microbes residing in these strains were different; the number of colony-forming units (CFUs) of commensal microbes was lowest in the Canton-S strain and highest in the w1118 (Figure 3A, ANOVA, p < 0.0001; Tukey’s HSD test, w1118 vs.OR, p < 0.0001; w1118 vs.CS, p < 0.0001; OR vs.CS, p < 0.005). In all three strains, the bleached egg effect on the longevity of 3G Ax flies was observed, but the extent of lifespan extension resulting from bacterial removal was different among the strains. Bacterial removal increased the lifespan of w1118 flies by 32.01%, whereas the lifespan increases were only 16.82% and 3.03% in the Oregon-R and Canton-S strains, respectively (Figure 3B and 3C, Supplementary Table 1, log-rank test, w1118, χ2 = 146.33, p < 0.0001; Oregon-R, χ2 = 37.43, p < 0.0001; Canton-S, χ2 = 6.60, p < 0.05). Moreover, the extent of the lifespan extension was positively correlated with the abundance of commensal microbes residing in each strain (Spearman’s correlation, rho = 0.88, p < 0.0001). To confirm that the longevity effect produced by axenic culture was related to bacteria abundance in flies, we performed an analysis of covariance (ANCOVA) test to adjust the bacterial loads among three strain groups. Following the ANCOVA test and after adjustment of bacterial abundance, we confirmed that the differences in the lifespan extension effect of axenic culture among the three strains disappeared (ANCOVA, p = 0.99). It showed the possibility that the difference in lifespan extension effect by bacteria removal among these three strains was the result of colonized bacterial abundance.
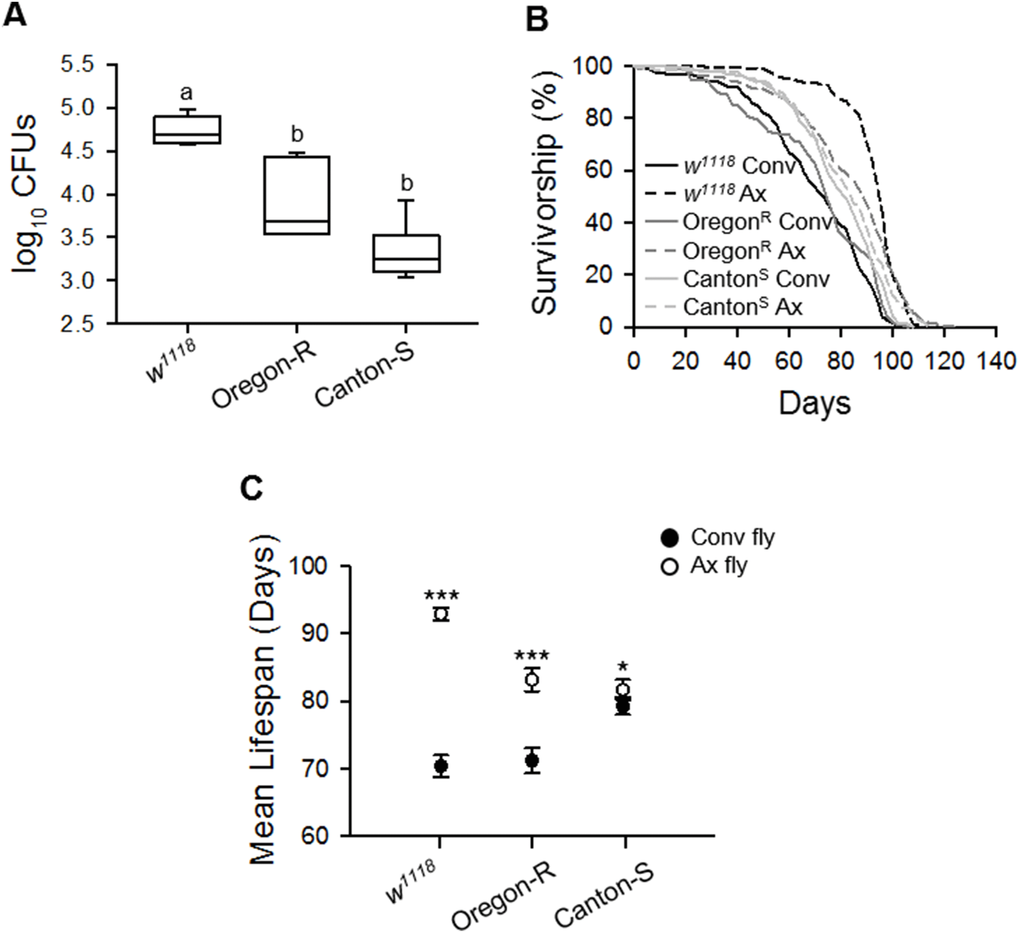
Figure 3. Lifespan extension effect of the elimination of commensal microbes differs among laboratory fly strains. (A) The total number of CFUs from 1-week-old flies in plate count agar (PCA) media. (B) Survival curve of laboratory wild-type w1118, Oregon-R, and Canton-S strains with the elimination of microbes. Solid lines, Conv flies; dashed lines, Ax flies. (C) Mean lifespan of laboratory wild-type w1118, Oregon-R, and Canton-S strains following the elimination of microbes. Filled dots, Conv flies; open dots, Ax flies. Asterisks indicate significant differences between Conv flies and Ax flies (log-rank test, *p < 0.05, ***p < 0.0001). Error bars represent the SEM.
Taken together, the results showed that the elimination of commensal bacteria using a method that is not accompanied by detrimental side effects can increase the lifespan of flies. In addition, strains colonized with a high level of commensal bacteria exhibit a greater increase in lifespan following bacteria elimination.
Discussion
Commensal microbes in a symbiotic relationship are known to affect host health and aging. In this study, we observed that commensal microbes can affect host lifespan in D. melanogaster, and an increase in microbial load in aging flies can be a stronger determinant of lifespan than that from a compositional change in microbiota.
There is a growing body of evidence indicating that commensal microbes, directly and indirectly, affect the lifespan of a host. For example, supplementation with different strains of Escherichia coli can affect the lifespan of C. elegans through direct, metabolic, or species-specific signals [24]. Also, the lifespan of aged killifish can be improved when they are treated with transplanted feces from young fish [25]. Moreover, germ-free mice have lived longer than their germ-bearing counterparts [26]. Similar to those animal model studies, several studies have shown that commensal microbes have effects on lifespan in Drosophila; however, the effects reported have been contradictory.
In this study, we generated axenic flies by using two methods (egg bleaching and antibiotic treatment) and then determined their lifespans. Interestingly, the lifespan of Ax flies was shorter than that of Conv flies after egg bleaching (Figure 1), which indicates that bleaching can have a detrimental effect on lifespan since the effect was diminished as the number of generations of Ax flies increased. In addition, the lifespan of Ax flies was similar to that of Conv flies after antibiotic treatment at a dosage equal to that used by Ren et al. (2007) (Figure 2). However, that antibiotic dose appeared to have a toxic effect on Ax flies since dilution of the antibiotic cocktail increased the lifespan of Ax flies, resulting in a longer lifespan in Ax flies than that in Conv flies. Egg bleaching and antibiotic treatment are commonly used methods to generate Ax flies; regardless, our results suggest that the toxic effects of both of those methods should be considered when generating Ax flies. Most authors have not provided information about which fly generation after egg bleaching they used in their studies [11–13, 15, 27–29], and some tested antibiotics within a very narrow concentration window [11, 12, 17, 27, 28, 30] (see Supplementary Table 12). The inconsistencies in the results reported by different groups on the effect of eliminating microbes on host lifespan might be due to differences in the methods used for microbe elimination.
The above-mentioned inconsistencies among results might also be due to the distinct microbial flora within the flies of each laboratory. Flies reared under the same conditions in the same laboratory can have different microbial flora, depending on their genetic background [21]. In this study, we observed that microbial loads were different among the three tested Drosophila strains, and the strains colonized with a higher level of commensal bacteria exhibited a greater increase in lifespan following bacteria elimination than that in strains with a lower level of commensal bacteria (Figure 3), suggesting that differential bacterial loads within the flies of each laboratory can give rise to differences in lifespan changes after bacterial elimination.
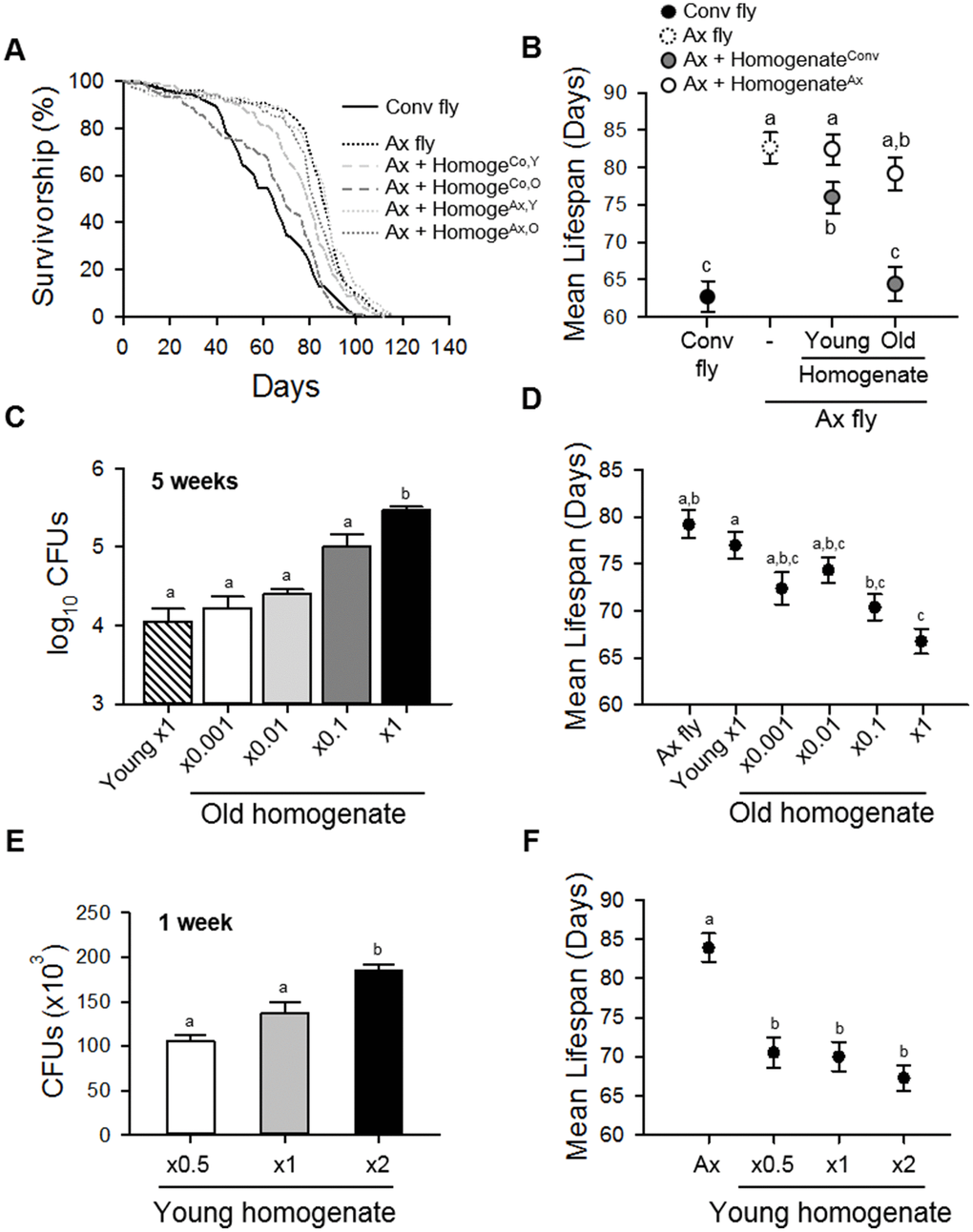
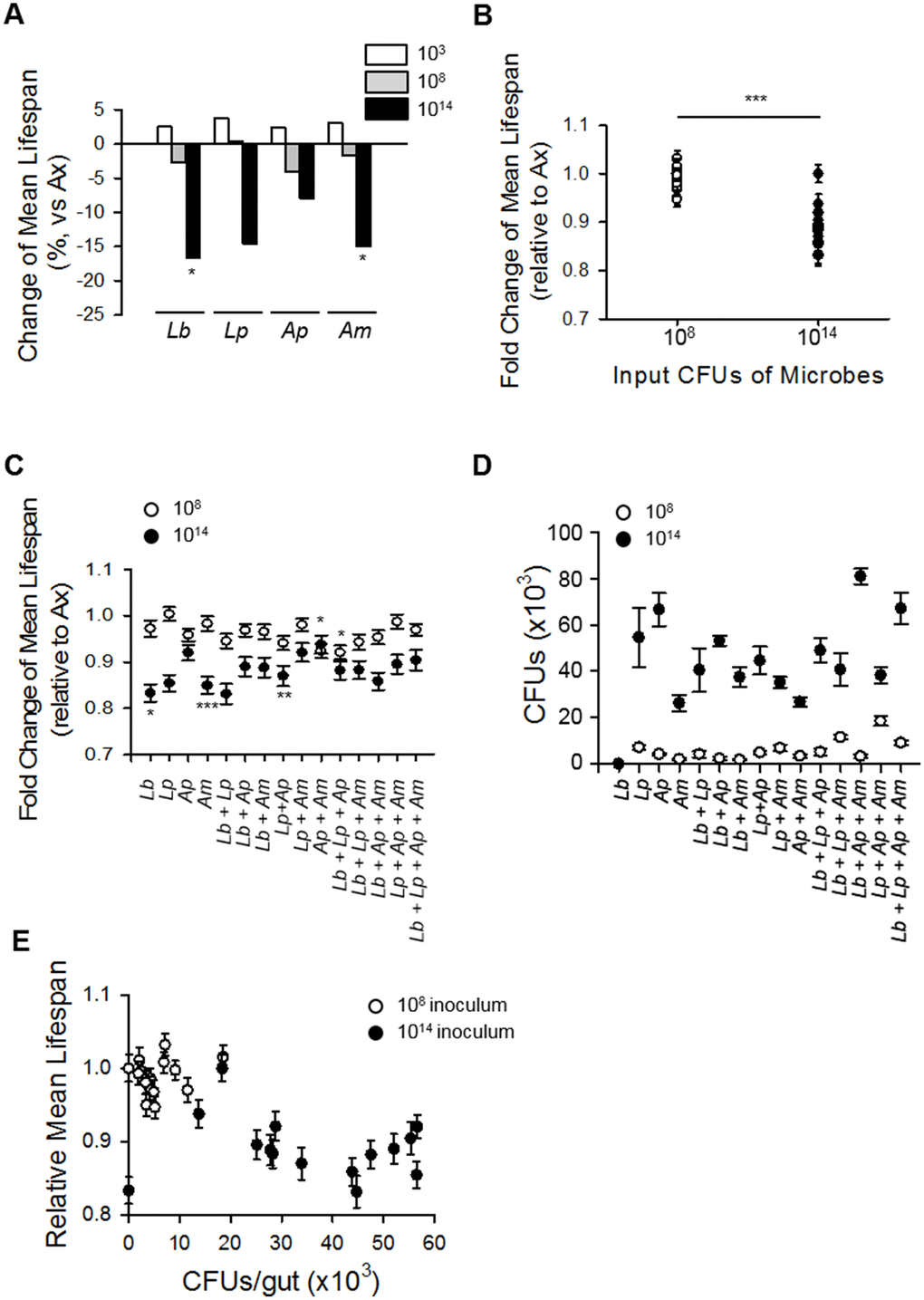
Under our experimental conditions, abundance of commensal microbes was a more important factor than the composition of the microbiota in lifespan determination of Drosophila. Ingestion of diluted old fly homogenate had an effect on lifespan that was similar to that of undiluted young fly homogenate, and the ingestion of young fly homogenate decreased the lifespan of flies in a dose-dependent manner (Figure 4). In addition, gnotobiotic flies colonized with a combination of four dominant bacteria showed that the abundance of commensal microbes exerted a significant influence on fly lifespan, whereas the composition of the microbiota exerted only a moderate effect on lifespan (Figure 5). Consistent with our results, several studies have reported on the effect of commensal microbial load on host lifespan. Flies fed homogenate from old flies have been shown to have a decreased lifespan compared to that in flies fed young fly homogenate [15]. Moreover, an increase in microbiota diversity has been correlated with an increase in bacterial load, resulting in a lifespan decrease [31]. In addition, mutant flies lacking the POU domain transcription factor Nub-PD had a shorter lifespan than control flies with a high diversity and high abundance microbiome [30], while flies treated with rapamycin exhibited delayed microbial expansion with age and had an extended lifespan [32].
Our study was the first to undertake a comparative analysis of the relative effects of microbiota load and microbiota composition on Drosophila lifespan. Our results show that microbiota load has a greater influence than microbiota composition on the lifespan of Drosophila; however, we cannot exclude the possibility that a specific bacterial species proliferates faster than others which could affect the lifespan of axenic flies. In addition, we cannot exclude the possibility that other factors can also affect the host’s lifespan, including microbial diversity and species specificity. There are several reports showing age-related increases in bacterial load and its deleterious effect on health, and there have been trials investigating the modulation of organismal health and lifespan by specific microbes. For example, Bifidobacterium animalis lactis has promoted longevity and reduced tumor incidence in mice [33], and Lactobacillus salivarius isolated from a centenarian's fecal samples extended the lifespan of C. elegans [34]. In addition, many bacteria have been shown to have species- or strain-specific effects on host health. L. plantarum was shown to affect systemic larval growth in Drosophila in a strain-specific manner [3]. Moreover, L. plantarum induces dNox dependent cellular ROS production, but that effect was not observed with other members of the microbiota [35]. In addition, L. brevis and G. morbifer can induce chronic DUOX activation via uracil production, while other commensal microbes cannot [36]. Consistent with those observations, our results showed that colonization of the most dominant microbes combinations at 108 CFUs did not have a significant effect on lifespan, but inoculation of Ap+Am or Lb+Lp+Ap decreased lifespan (Figure 5C, Supplementary Figure 5, Supplementary Table 5–7, ANOVA, p < 0.0005; Tukey's HSD vs. Ax, Ax 85.59 ± 0.97 days; Ap+Am 79.08 ± 1.31 days, 7.61% decrease, p < 0.05; Lb+Lp+Ap 78.80 ± 1.28 days, 7.93% decrease, p < 0.05), even though the final concentrations of the inocula were equal (Figure 5D, Supplementary Table 10, Tukey's HSD vs. each group, p > 0.05). Moreover, although the bacteria decreased lifespan in all species combinations at high concentration (1014 CFUs), the extent of the decrease was different with the different microbe combinations (Figure 5C, Supplementary Table 5, Supplementary Tables 8, 9, ANOVA, p < 0.0005; Ax 76.68 ± 1.58 days; Lb 63.89 ± 1.42 days, 16.80% decrease, Tukey’s HSD test, p < 0.05; Am 65.17 ± 1.42 days, 15.01% decrease, Tukey’s HSD test, p < 0.005; Lp+Ap 66.72 ± 1.70 days, 12.99% decrease, Tukey’s HSD test, p < 0.005), indicating that there may be species-, strain-, or combination-specificity involved in the microbiota effects on lifespan regulation.
In addition, we cannot exclude the role of minor bacteria in lifespan regulation. In this study, we selected the four dominant species (the top two species in each age group) for our microbial combination study. Among the subdominant species, our 10-day-old flies contained K. medellinensis, and there was a high proportion presence of L. pseudomesenteroides in our 50-day-old flies. K. medellinensis (also called Gluconacetobacter xylinus) is reported to reduce triglyceride and glucose levels in Drosophila [37], and a reduced glucose level has been shown in flies with a low insulin-like peptide level, and such flies have been reported to live longer than control flies [38]. Leuconostoc negatively interacts with Lactobacillus indicating that the two genera occupy the same niche in Drosophila [39]; but whether Leuconostoc regulates lifespan along with Lactobacillus has not been reported. Taken collectively, the species-specific or compositional effects of the microbiome on the flies in this study may have been overshadowed by the stronger lifespan effect associated with bacterial abundance.
The mechanism of how commensal bacteria can regulate host lifespan is a fascinating study topic. There are some studies that have described this mechanism in relation to intestinal barrier dysfunction, ROS generation, intestinal stem cell dysplasia, and several lifespan-regulating pathways. Clark et al. showed that deleterious changes in a microbiota can induce intestinal barrier dysfunction and is a primary cause of mortality [15, 40]. In addition, the intracellular ROS level has been considered a critical cause of aging [41], and ROS generation induced by an increase in commensal or pathogenic microbes, as a defensive response, has been suggested to affect host lifespan [36, 42]. Furthermore, microbes are reported to regulate intestinal stem cell (ISC) proliferation by inducing ROS generation [23, 43, 44], and the elimination of the microbiota has induced the quiescent stage in ISCs and reduced the number of progenitor cells [43, 44]. Lastly, lifespan-regulating signaling pathways, including the insulin/IGF-1 signaling pathway and the target of rapamycin pathway, have been reported to be regulated by commensal microbes [3, 4].
Taken together, we suggest that when an axenic fly is generated for use in host-microbe interaction study, it should be noted that both egg bleaching and antibiotic treatment can have an adverse effect on the health of the Drosophila, but these adverse effects can be eliminated by allowing trans-generation of flies hatched from bleached eggs or by using a suitably diluted antibiotics. In this study, we demonstrated that the age-related increase in bacterial load more strongly affects the lifespan of Drosophila than that associated with changes in microbiota composition. Our results present a basic but deterministic connecting point in the relationship between commensal microbes and host lifespan.
Methods
Fly husbandry and generation of axenic D. melanogaster
All experiments, except those related to the strain-specific longevity effect of commensal microbe elimination, were conducted using flies of the D. melanogaster wild-type strain w1118 that were initially provided by the Bloomington Stock Center (Indiana University, USA) and have been adapting to our laboratory environment for 8 years. The enterobacteria Wolbachia, which can affect lifespan, was not present in the strains used in this study as was determined by PCR assay (data not shown). Flies were cultured and reared at 25ºC and 65% humidity on a 12:12 hour light:dark cycle. Sterile standard cornmeal-sugar-yeast (CSY) media (Supplementary Table 13) were used during culture and rearing of the flies. To produce the sterile CSY diet, the above-mentioned CSY medium was autoclaved at 120°C for 20 min, and all vials for food were exposed to UV light for 20 min on a clean bench. For the preparation of antibiotic-treated food, 640 μg/mL doxycycline (Sigma-Aldrich), 640 μg/mL ampicillin (Sigma-Aldrich), and 1 mg/mL kanamycin (Sigma-Aldrich) were added to sterile CSY media [12].
Axenic (Ax) flies were generated by bleaching the embryos. Embryos were collected for 12 h and were then dechorionated for 50 sec in 5% sodium hypochlorite solution (Wako, Japan), rinsed for 50 sec in 70% ethanol, and washed for 1 min in sterile distilled water [15]. Sterile embryos were transferred into sterile CSY food bottles on a clean bench. Eggs in an Ax condition were passed through repeated generations and became third-generation flies. All Ax flies were maintained on a clean bench and were transferred to fresh food every two days. The Ax conditions were confirmed by plating fly homogenate on plate count agar (PCA, Neogen Corporation, MI, USA) (Supplementary Table 13), and by 16S rRNA gene PCR using a bacterial 16S rRNA universal primer (27F and 1492R) provided by Macrogen (Seoul, South Korea).
Bacteria culture
Conventionally, bacteria were cultured on PCA medium. Lactobacillus was grown on 5.5% MRS media (Lactobacilli MRS Broth, BD & Difco, MD, USA) and Acetobacter was grown on Acetobacter-selective media (see Supplementary Table 13). All microbes were incubated at 29°C.
Quantitative analysis of bacteria
For CFU determination, 5 females were rinsed in 70% ethanol for 3 sec for surface decontamination and then homogenized in sterile distilled water. The homogenates were diluted as necessary and plated onto PCA media, MRS media, or Acetobacter-selective media. At least 5 replicates were established for each group. Data are presented as mean ± standard error of the mean (SEM) values. For 16S rRNA PCR, total genomic DNA from 45 surface-sterilized female flies was extracted by using a DNeasy Tissue Kit (Hilden, Germany) in accordance with the manufacturer’s instructions. The PCR assays were performed at a 60°C annealing temperature and for 40–60 cycles using taxon-specific 16S rRNA gene primers for Lactobacillus or Acetobacter designed using Primer3 software, as well as universal PCR primers. Sequences are presented in Supplementary Table 14.
Identification of commensal microbes
For commensal microbe isolation, homogenates from 10- or 50-day-old female flies were plated on a PCA media plate. After incubation of a single colony at 29°C for 3 days, each colony was transferred to Acetobacter-selective or Lactobacillus-selective media broth. After culturing for 24 h, the cell walls of isolated microbes were broken down by bead beating using 0.1 mm diameter glass beads (BioSpec Products, Bartlesville, OK, USA). PCR assays were performed with a 55°C annealing temperature and 45 cycles with universal primers 27F and 1492R. PCR products were sequenced by using 16S sequencing (Macrogen Inc., South Korea) with universal primers 518F and 800R and then analyzed by using EzTaxon BLAST and NCBI BLAST. To determine the dominant commensal bacteria species in the gut of flies, 454 pyrosequencing analysis of the 16S rRNA gene was performed. The 16S rRNA gene amplicons from 100 dissected guts (comprising the Malpighian tubules but excluding the crop) from surface-sterilized females were analyzed by pyrosequencing using the 454 GS FLX Titanium Sequencing System (Roche, Brandford, CT, USA) at Chunlab Inc (South Korea). Phylogenetic relationships were determined by using EzTaxon BLAST and NCBI BLAST.
Introduction of commensal microbes to axenic D. melanogaster
To feed fly homogenate to Ax fly, fly homogenates from surface sterile flies with 70% ethanol were seeded in CSY food vials. The Ax flies were transferred to new homogenate-containing vials three times for a week. To generate adult flies carrying a predetermined composition of bacteria (gnotobiotic flies), 100 μL of commensal bacterial cultures (103, 108, or 1014 CFUs) were added to sterilized food vials containing 2-day-old Ax flies. Every 2 days for 1 week, the flies were transferred to new sterile CSY food vials seeded with experiment-specific compositions of commensal bacteria. For commensal bacteria compositions of more than one species, each species component was added in equal parts to make up the total inoculum. The abundances of colonized microbes were identified via CFU testing 10 days after the initial infection.
Lifespan assay
Newly eclosed adult female flies were collected for 2 days and were provided with a stabilizing time of 1 day with male flies. Mated female flies were randomly assigned to sterile CSY food vials to a final density of 20 flies per vial. Vials were changed every 2 days for new vials containing fresh sterile CSY food; at that time, dead flies were removed and the number was recorded. In the case of gnotobiotic flies, counting of dead flies was performed after bacterial seeding for 1 week. Ten replicate vials were established for each group (n = 200). Once a month, vials were spot checked for contamination by swabbing the food in the spent vials and plating on PCA-bearing culture plates.
Statistical analysis
Log-rank tests were carried out to determine the statistical significance of the results of the survival analysis. The JMP statistical package (SAS, NC, USA) was used for the analyses. The statistical probabilities of the obtained CFU and OTU numbers were determined by using the two-sample t-test. ANOVA, Tukey’s HSD test, and Spearman’s correlation coefficients were derived by using R 3.5.1 software.
Supplementary Materials
Author Contributions
H.Y., S.H., and K.J. designed the research; H.Y., S.H., and J.H. performed the research; H.Y. and S.H. analyzed the data; H.Y., S.H., W.J., and K.J. wrote the paper.
Conflicts of Interest
The authors declare no conflicts of interest.
Funding
This work was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF), funded by the Ministry of Education, Science, and Technology (No. 2015R1A2A2A01005580, 2015R1A3A2033475 and 2016R1D1A1B03930533).
References
- 1. Moran CP, Shanahan F. Gut microbiota and obesity: role in aetiology and potential therapeutic target. Best Pract Res Clin Gastroenterol. 2014; 28:585–97. https://doi.org/10.1016/j.bpg.2014.07.005 [PubMed]
- 2. Kostic AD, Xavier RJ, Gevers D. The microbiome in inflammatory bowel disease: current status and the future ahead. Gastroenterology. 2014; 146:1489–99. https://doi.org/10.1053/j.gastro.2014.02.009 [PubMed]
- 3. Storelli G, Defaye A, Erkosar B, Hols P, Royet J, Leulier F. Lactobacillus plantarum promotes Drosophila systemic growth by modulating hormonal signals through TOR-dependent nutrient sensing. Cell Metab. 2011; 14:403–14. https://doi.org/10.1016/j.cmet.2011.07.012 [PubMed]
- 4. Shin SC, Kim SH, You H, Kim B, Kim AC, Lee KA, Yoon JH, Ryu JH, Lee WJ. Drosophila microbiome modulates host developmental and metabolic homeostasis via insulin signaling. Science. 2011; 334:670–74. https://doi.org/10.1126/science.1212782 [PubMed]
- 5. Diaz SA, Mooring EQ, Rens EG, Restif O. Association with pathogenic bacteria affects life-history traits and population growth in Caenorhabditis elegans.. Ecol Evol. 2015; 5:1653–63. https://doi.org/10.1002/ece3.1461 [PubMed]
- 6. Thaiss CA, Zeevi D, Levy M, Zilberman-Schapira G, Suez J, Tengeler AC, Abramson L, Katz MN, Korem T, Zmora N, Kuperman Y, Biton I, Gilad S, et al. Transkingdom control of microbiota diurnal oscillations promotes metabolic homeostasis. Cell. 2014; 159:514–29. https://doi.org/10.1016/j.cell.2014.09.048 [PubMed]
- 7. Yano JM, Yu K, Donaldson GP, Shastri GG, Ann P, Ma L, Nagler CR, Ismagilov RF, Mazmanian SK, Hsiao EY. Indigenous bacteria from the gut microbiota regulate host serotonin biosynthesis. Cell. 2015; 161:264–76. https://doi.org/10.1016/j.cell.2015.02.047 [PubMed]
- 8. Houthoofd K, Braeckman BP, Lenaerts I, Brys K, De Vreese A, Van Eygen S, Vanfleteren JR. Axenic growth up-regulates mass-specific metabolic rate, stress resistance, and extends life span in Caenorhabditis elegans.. Exp Gerontol. 2002; 37:1371–78. https://doi.org/10.1016/S0531-5565(02)00173-0 [PubMed]
- 9. Garsin DA, Villanueva JM, Begun J, Kim DH, Sifri CD, Calderwood SB, Ruvkun G, Ausubel FM. Long-lived C. elegans daf-2 mutants are resistant to bacterial pathogens. Science. 2003; 300:1921. https://doi.org/10.1126/science.1080147 [PubMed]
- 10. Han B, Sivaramakrishnan P, Lin CJ, Neve IAA, He J, Tay LWR, Sowa JN, Sizovs A, Du G, Wang J, Herman C, Wang MC. Microbial genetic composition tunes host longevity. Cell. 2017; 169:1249–1262.e13. https://doi.org/10.1016/j.cell.2017.05.036 [PubMed]
- 11. Brummel T, Ching A, Seroude L, Simon AF, Benzer S. Drosophila lifespan enhancement by exogenous bacteria. Proc Natl Acad Sci USA. 2004; 101:12974–79. https://doi.org/10.1073/pnas.0405207101 [PubMed]
- 12. Ren C, Webster P, Finkel SE, Tower J. Increased internal and external bacterial load during Drosophila aging without life-span trade-off. Cell Metab. 2007; 6:144–52. https://doi.org/10.1016/j.cmet.2007.06.006 [PubMed]
- 13. Ridley EV, Wong AC, Westmiller S, Douglas AE. Impact of the resident microbiota on the nutritional phenotype of Drosophila melanogaster.. PLoS One. 2012; 7:e36765. https://doi.org/10.1371/journal.pone.0036765 [PubMed]
- 14. Petkau K, Parsons BD, Duggal A, Foley E. A deregulated intestinal cell cycle program disrupts tissue homeostasis without affecting longevity in Drosophila.. J Biol Chem. 2014; 289:28719–29. https://doi.org/10.1074/jbc.M114.578708 [PubMed]
- 15. Clark RI, Salazar A, Yamada R, Fitz-Gibbon S, Morselli M, Alcaraz J, Rana A, Rera M, Pellegrini M, Ja WW, Walker DW. Distinct shifts in microbiota composition during Drosophila aging impair intestinal function and drive mortality. Cell Rep. 2015; 12:1656–67. https://doi.org/10.1016/j.celrep.2015.08.004 [PubMed]
- 16. Yamada R, Deshpande SA, Bruce KD, Mak EM, Ja WW. Microbes promote amino acid harvest to rescue undernutrition in Drosophila.. Cell Rep. 2015; 10:865–72. https://doi.org/10.1016/j.celrep.2015.01.018 [PubMed]
- 17. Galenza A, Hutchinson J, Campbell SD, Hazes B, Foley E. Glucose modulates Drosophila longevity and immunity independent of the microbiota. Biol Open. 2016; 5:165–73. https://doi.org/10.1242/bio.015016 [PubMed]
- 18. Téfit MA, Leulier F. Lactobacillus plantarum favors the early emergence of fit and fertile adult Drosophila upon chronic undernutrition. J Exp Biol. 2017; 220:900–07. https://doi.org/10.1242/jeb.151522 [PubMed]
- 19. Obata F, Fons CO, Gould AP. Early-life exposure to low-dose oxidants can increase longevity via microbiome remodelling in Drosophila.. Nat Commun. 2018; 9:975. https://doi.org/10.1038/s41467-018-03070-w [PubMed]
- 20. Iatsenko I, Boquete JP, Lemaitre B. Microbiota-derived lactate activates production of reactive oxygen species by the intestinal NADPH oxidase nox and shortens Drosophila lifespan. Immunity. 2018; 49:929–942.e5. https://doi.org/10.1016/j.immuni.2018.09.017 [PubMed]
- 21. Han G, Lee HJ, Jeong SE, Jeon CO, Hyun S. Comparative analysis of Drosophila melanogaster gut microbiota with respect to host strain, sex, and age. Microb Ecol. 2017; 74:207–16. https://doi.org/10.1007/s00248-016-0925-3 [PubMed]
- 22. Wong CN, Ng P, Douglas AE. Low-diversity bacterial community in the gut of the fruitfly Drosophila melanogaster.. Environ Microbiol. 2011; 13:1889–900. https://doi.org/10.1111/j.1462-2920.2011.02511.x [PubMed]
- 23. Guo L, Karpac J, Tran SL, Jasper H. PGRP-SC2 promotes gut immune homeostasis to limit commensal dysbiosis and extend lifespan. Cell. 2014; 156:109–22. https://doi.org/10.1016/j.cell.2013.12.018 [PubMed]
- 24. Heintz C, Mair W. You are what you host: microbiome modulation of the aging process. Cell. 2014; 156:408–11. https://doi.org/10.1016/j.cell.2014.01.025 [PubMed]
- 25. Smith P, Willemsen D, Popkes M, Metge F, Gandiwa E, Reichard M, Valenzano DR. Regulation of life span by the gut microbiota in the short-lived African turquoise killifish. eLife. 2017; 6:6. https://doi.org/10.7554/eLife.27014 [PubMed]
- 26. Wostmann BS. Germ-free versus non-germ-free animals in gerontological research. 1968.
- 27. Fast D, Duggal A, Foley E. The symbiont Lactobacillus plantarum causes intestinal pathogenesis in adult Drosophila.. bioRxiv. 2016. https://doi.org/10.1101/049981
- 28. Li H, Qi Y, Jasper H. Preventing age-related decline of gut compartmentalization limits microbiota dysbiosis and extends lifespan. Cell Host Microbe. 2016; 19:240–53. https://doi.org/10.1016/j.chom.2016.01.008 [PubMed]
- 29. Sannino DR, Dobson AJ, Edwards K, Angert ER, Buchon N. The Drosophila melanogaster gut microbiota provisions thiamine to its host. MBio. 2018; 9:e00155–18. https://doi.org/10.1128/mBio.00155-18 [PubMed]
- 30. Dantoft W, Lundin D, Esfahani SS, Engström Y. The POU/Oct transcription factor Pdm1/nub is necessary for a beneficial gut microbiota and normal lifespan of Drosophila.. J Innate Immun. 2016; 8:412–26. https://doi.org/10.1159/000446368 [PubMed]
- 31. Gould A, Zhang V, Lamberti L, Jones E, Obadia B, Gavryushkin A, Carlson J, Beerenwinkel N, Ludington W. High-dimensional microbiome interactions shape host fitness. bioRxiv. 2017. https://doi.org/10.1101/232959
- 32. Fan X, Liang Q, Lian T, Wu Q, Gaur U, Li D, Yang D, Mao X, Jin Z, Li Y, Yang M. Rapamycin preserves gut homeostasis during Drosophila aging. Oncotarget. 2015; 6:35274–83. https://doi.org/10.18632/oncotarget.5895 [PubMed]
- 33. Matsumoto M, Kurihara S, Kibe R, Ashida H, Benno Y. Longevity in mice is promoted by probiotic-induced suppression of colonic senescence dependent on upregulation of gut bacterial polyamine production. PLoS One. 2011; 6:e23652. https://doi.org/10.1371/journal.pone.0023652 [PubMed]
- 34. Zhao Y, Zhao L, Zheng X, Fu T, Guo H, Ren F. Lactobacillus salivarius strain FDB89 induced longevity in Caenorhabditis elegans by dietary restriction. J Microbiol. 2013; 51:183–88. https://doi.org/10.1007/s12275-013-2076-2 [PubMed]
- 35. Jones RM, Luo L, Ardita CS, Richardson AN, Kwon YM, Mercante JW, Alam A, Gates CL, Wu H, Swanson PA, Lambeth JD, Denning PW, Neish AS. Symbiotic lactobacilli stimulate gut epithelial proliferation via Nox-mediated generation of reactive oxygen species. EMBO J. 2013; 32:3017–28. https://doi.org/10.1038/emboj.2013.224 [PubMed]
- 36. Lee KA, Kim SH, Kim EK, Ha EM, You H, Kim B, Kim MJ, Kwon Y, Ryu JH, Lee WJ. Bacterial-derived uracil as a modulator of mucosal immunity and gut-microbe homeostasis in Drosophila. Cell. 2013; 153:797–811. https://doi.org/10.1016/j.cell.2013.04.009 [PubMed]
- 37. Chaston JM, Newell PD, Douglas AE. Metagenome-wide association of microbial determinants of host phenotype in Drosophila melanogaster.. MBio. 2014; 5:e01631–14. https://doi.org/10.1128/mBio.01631-14 [PubMed]
- 38. Broughton SJ, Piper MD, Ikeya T, Bass TM, Jacobson J, Driege Y, Martinez P, Hafen E, Withers DJ, Leevers SJ, Partridge L. Longer lifespan, altered metabolism, and stress resistance in Drosophila from ablation of cells making insulin-like ligands. Proc Natl Acad Sci USA. 2005; 102:3105–10. https://doi.org/10.1073/pnas.0405775102 [PubMed]
- 39. Pais IS, Valente RS, Sporniak M, Teixeira L. Drosophila melanogaster establishes a species-specific mutualistic interaction with stable gut-colonizing bacteria. PLoS Biol. 2018; 16:e2005710. https://doi.org/10.1371/journal.pbio.2005710 [PubMed]
- 40. Salazar AM, Resnik-Docampo M, Ulgherait M, Clark RI, Shirasu-Hiza M, Jones DL, Walker DW. Intestinal snakeskin limits microbial dysbiosis during aging and promotes longevity. iScience. 2018; 9:229–243. https://doi.org/10.1016/j.isci.2018.10.022 [PubMed]
- 41. Barja G. The mitochondrial free radical theory of aging. Prog Mol Biol Transl Sci. 2014; 127:1–27. https://doi.org/10.1016/B978-0-12-394625-6.00001-5 [PubMed]
- 42. Jones RM, Desai C, Darby TM, Luo L, Wolfarth AA, Scharer CD, Ardita CS, Reedy AR, Keebaugh ES, Neish AS. Lactobacilli modulate epithelial cytoprotection through the Nrf2 pathway. Cell Rep. 2015; 12:1217–25. https://doi.org/10.1016/j.celrep.2015.07.042 [PubMed]
- 43. Buchon N, Broderick NA, Chakrabarti S, Lemaitre B. Invasive and indigenous microbiota impact intestinal stem cell activity through multiple pathways in Drosophila.. Genes Dev. 2009; 23:2333–44. https://doi.org/10.1101/gad.1827009 [PubMed]
- 44. Broderick NA, Buchon N, Lemaitre B. Microbiota-induced changes in drosophila melanogaster host gene expression and gut morphology. MBio. 2014; 5:e01117–14. https://doi.org/10.1128/mBio.01117-14 [PubMed]