Introduction
Gliomas are the most common and aggressive primary brain cancers in adult patients [1]. Despite improvements in diagnosis and treatment, the prognosis of glioma remains unfavorable. Especially for glioblastoma (GBM) patients, the most malignant type, the median survival time is only about fifteen months [2, 3]. Epithelial-to-mesenchymal transition (EMT) has been widely reported as a key mechanism in promoting migration, invasion, and tumor progression in glioma [4]. Identification of novel EMT-related markers will facilitate the development of potential molecular targets for glioma patients.
Bradykinin (BK) is a vasoactive peptide produced from kininogen precursor’s cleavage under kallikrein’s action [5]. BK participates in a range of pathophysiological processes, including vasodilation, vascular permeability, pain-sensing, smooth muscle contraction, and inflammation modulation. The BK’s biological function is mediated by the activation of two G protein-coupled receptors: Bradykinin receptor B1 (BDKRB1) and BDKRB2. BDKRB1 is weakly or not expressed under normal physiological conditions, while proinflammatory mediators and oxidative stress can upregulate it, and it usually binds to the active metabolite des-Arg9-BK. However, BDKRB2 is constitutively expressed across different tissues, and it has a high affinity with BK [6]. There is a crosstalk between BDKRB1 and BDKRB2. Upregulation of BDKRB1 can be associated with BDKRB2 downregulation [7].
The BK system plays an important role in cancer occurrence and progression [8]. The BK system stimulates cell proliferation, migration, and angiogenesis, contributing to tumor progression [9]. As a vital receptor of bradykinin, BDKRB2 has been widely reported in a range of malignancies, including cervical cancer [10], triple-negative breast cancer [11], hepatocellular carcinoma (HCC) [12, 13], gastric cancer [14–16], colorectal cancer [17], prostate cancer [18], bladder cancer [19], head and neck squamous cell carcinomas [20], and chondrosarcoma [21]. Across different malignancies, via activating BDKRB2, BK promotes tumor progression through various pathways. For example, the BK-BDKRB2 axis can promote angiogenesis by increasing vascular permeability and by upregulating vascular endothelial growth factor (VEGF) in a sarcoma mouse model [22] and a Walker 256 carcinoma cell-bearing rat model [23]. Yu et al. further demonstrated that the BK-BDKRB2 axis activated Akt-mTOR signaling and downstream NF-κB and activator protein 1 (AP-1), which activated VEGF in human prostate cancer cells [24]. In HCC, BK-BDKRB2 promotes the migration and invasion of tumor cells through transient receptor potential cation channel subfamily M member 7 (TRPM7) and matrix metalloproteinase-2 (MMP2) [13]. In melanoma, BK-BDKRB2 upregulates endothelin-1 and subsequently increases the capacity of migration and invasion [25]. In gastric cancer, the BK-BDKRB2 axis promotes cell proliferation, migration, invasion through ERK signaling pathway [14]. Moreover, several studies sought to investigate the correlation between BDKRB2 expression and clinical characterization and have concluded relatively consistent results across different cancers. A higher expression level of BDKRB2 was reported to be correlated with more malignant features and shorter survival [10, 11, 26, 27].
Heretofore, some researchers have investigated the potential biological functions of BDKRB2 in glioma based on in-vivo and in-vitro experimental studies [7, 28–30]. However, we could not find a systematic report about BDKRB2 expression in pan-glioma from the clinical perspective. In the present study, 998 glioma patients with transcriptome data were enrolled and analyzed, aiming at investigating the clinical significance, characterization of expression profiling, and biological function of BDKRB2 in glioma.
Results
BDKRB2 was associated with EMT
To further illustrate the biological process of BDKRB2 in glioma, GSEA analyses were performed in both CGGA and TCGA datasets. We found that BDKRB2 was most correlated with EMT in CGGA (NES = 2.187, FDR = 0) (Figure 3A, 3E), which was further validated in TCGA (NES = 2.036, FDR = 0.019) (Figure 3B, 3F). Furthermore, GSEA analysis showed a similar pattern of functional enrichment in GBM of both datasets (Figure 3C, 3D, 3G, 3H). These results indicated that BDKRB2 was profoundly associated with EMT phenotype in glioma.
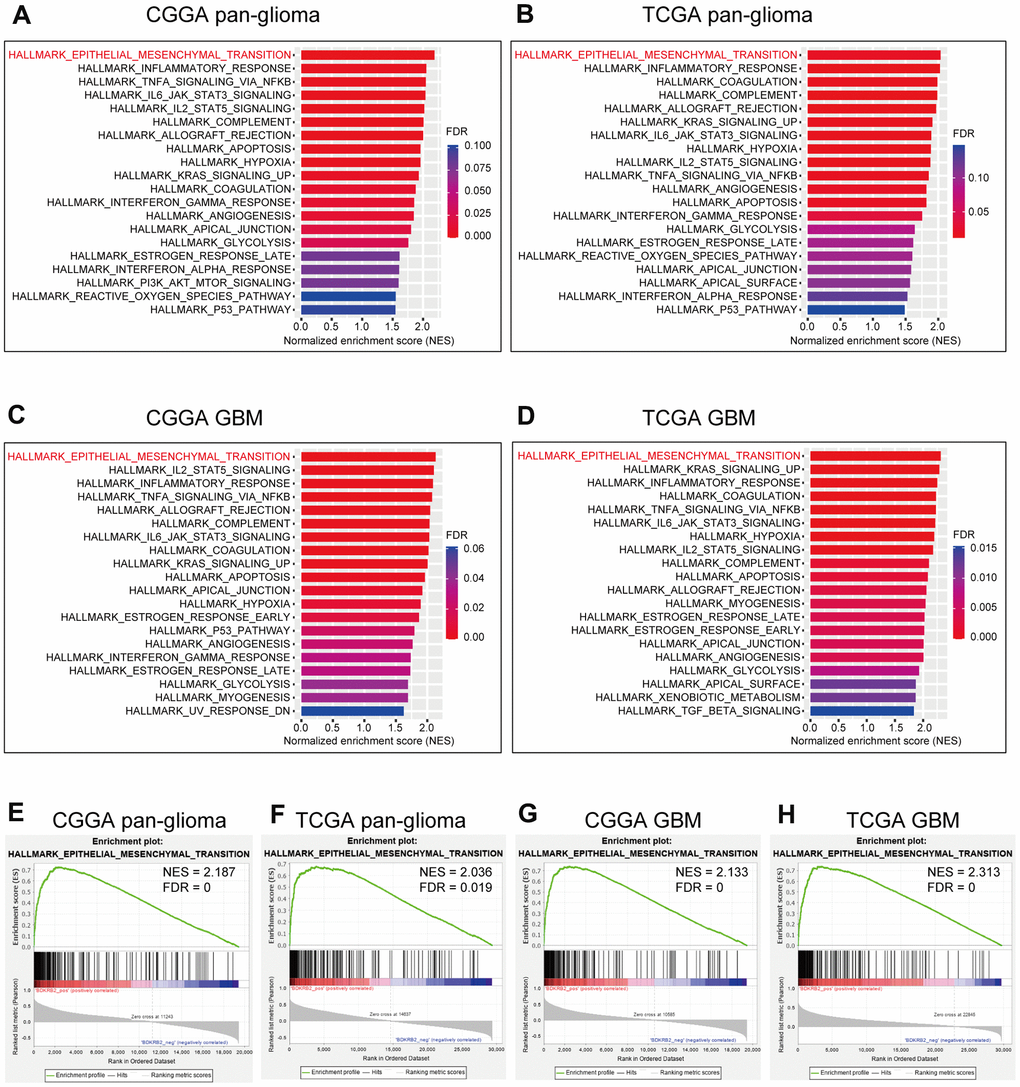
Figure 3. GSEA of BDKRB2 in pan-glioma (A, B) and glioblastoma (C, D), and GSEA plots for EMT enrichment according to BDKRB2 expression in pan-glioma (E, F) and glioblastoma (G, H).
BDKRB2 was associated with EMT biomarkers
To further validate the role of BDKRB2 in the EMT signaling pathway, we examined the correlation between BDKRB2 and EMT biomarkers, including E-cadherin, N-cadherin, snail, and slug. Circos plots revealed that BDKRB2 expression was significantly associated with N-cadherin, snail, and slug (Figure 5A, 5B). Pearson correlation tests were additionally performed in GBM. As shown in Figure 5C, 5D, the correlation between BDKRB2 and these markers in GBM was also very robust in both datasets. While the correlation between BDKRB2 and E-cadherin was very weak, which might be deemed as a noise.
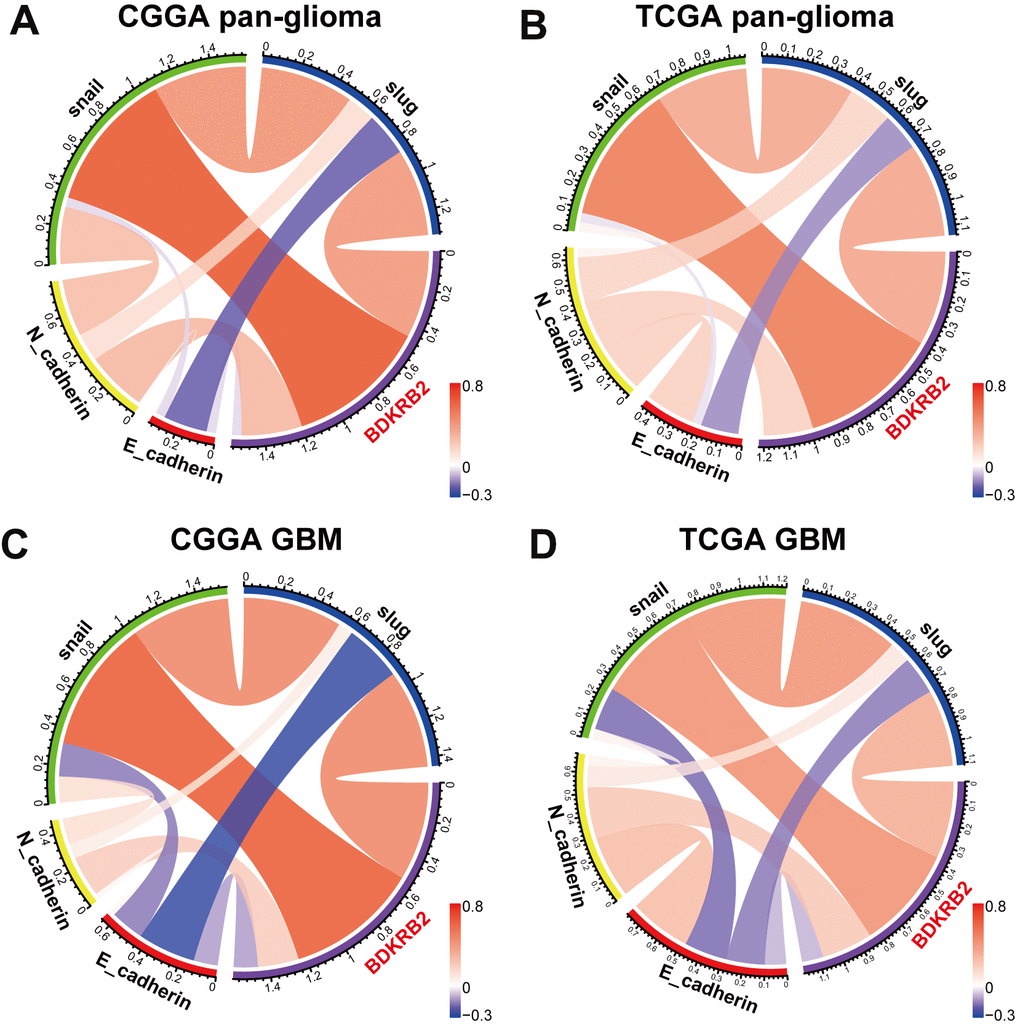
Figure 5. Correlation of BDKRB2 and key EMT biomarkers in pan-glioma (A, B) and glioblastoma (C, D).
Many other biomarkers have been identified as EMT-related targets in EMT [38]. We additionally enrolled EMT-related markers, including β-catenin, vimentin, TWIST1, and TWIST2, and put them into analysis together with BDKRB2. Subsequent Circos plots revealed that BDKRB2 expression was especially correlated with vimentin, TWIST1, and TWIST2 (Supplementary Figure 5).
Higher BDKRB2 predicted shorter survival for glioma
Kaplan-Meier (KM) survival analyses were performed to examine the prognostic role of BDKRB2 in glioma. According to BDKRB2 expression, pan-glioma samples were divided into two groups in each dataset. As shown in Figure 6A, 6D, a higher level of BDKRB2 expression predicted a significantly shorter survival. Moreover, a similar KM survival curve pattern was observed among patients with LGG (Figures 6B, 6E) and GBM (Figure 6C, 6F). To identify the independent effect of BDKRB2 on glioma prognosis, Cox regression analyses were performed with covariates, including BDKRB2 expression, age, and WHO grade. Multivariate analyses revealed that BDKRB2 expression was a significant prognosticator independent of age and WHO grade in both CGGA and TCGA (Table 1).
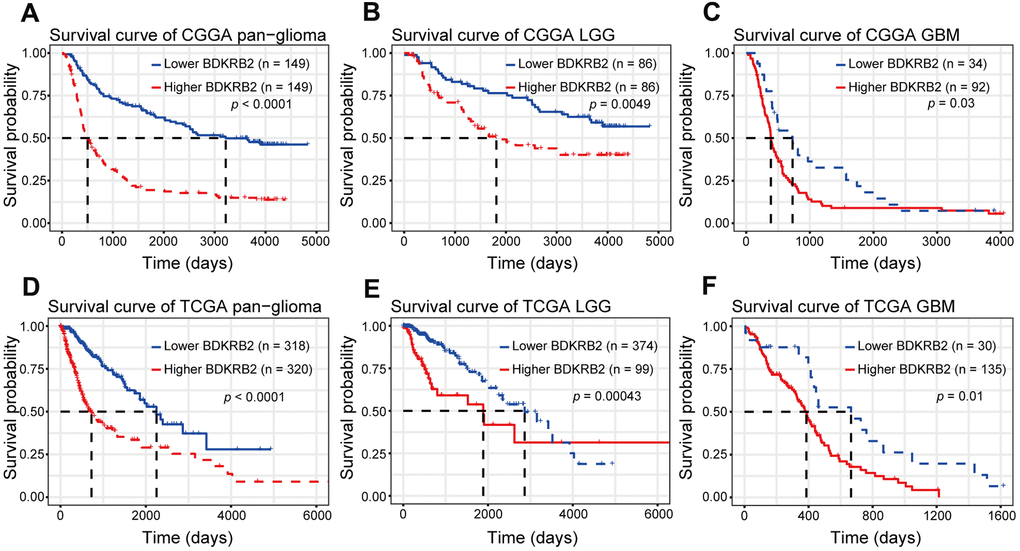
Figure 6. Survival analysis for BDKRB2 in pan-glioma (A, D), LGG (B, E) and GBM (C, F).
Table 1. Cox regression analysis of overall survival in glioma.
| Covariates | CGGA_301 | TCGA | |||||||||
| Univariate | Multivariate | Univariate | Multivariate | ||||||||
| HR(95% CI) | P | HR(95% CI) | P | HR(95% CI) | P | HR(95% CI) | P | ||||
| Age | 1.041 (1.027-1.055) | 0.000 | 1.018 (1.005-1.1.031) | 0.008 | 1.075 (1.062-1.087) | 0.000 | 1.048 (1.034-1.062) | 0.000 | |||
| Grade | 2.670 (2.221-3.210) | 0.000 | 2.329 (1.905-2.847) | 0.000 | 5.057 (3.915-6.532) | 0.000 | 3.011 (2.255-4.019) | 0.000 | |||
| BDKRB2 | 1.420 (1.282-1.573) | 0.000 | 1.126 (1.012-1.253) | 0.030 | 1.443 (1.335-1.559) | 0.000 | 1.155 (1.055-1.265) | 0.002 | |||
Discussion
Emerging evidence indicates BDKRB2 as a pivotal target in tumorigenesis. BDKRB2 as a frequently amplified molecule has been observed in a range of cancers, including cervical cancer [10], triple-negative breast cancer [11], hepatocellular carcinoma (HCC) [12, 13], gastric cancer [14–16], colorectal cancer [17], prostate cancer [18], bladder cancer [19], head and neck squamous cell carcinomas [20], and chondrosarcoma [21]. In glioma, only a few studies have reported that BDKRB2 was dysregulated in GBM cell lines [7, 28–30]. However, the expression profile and prognostic value of BDKRB2 in glioma are still largely unknown.
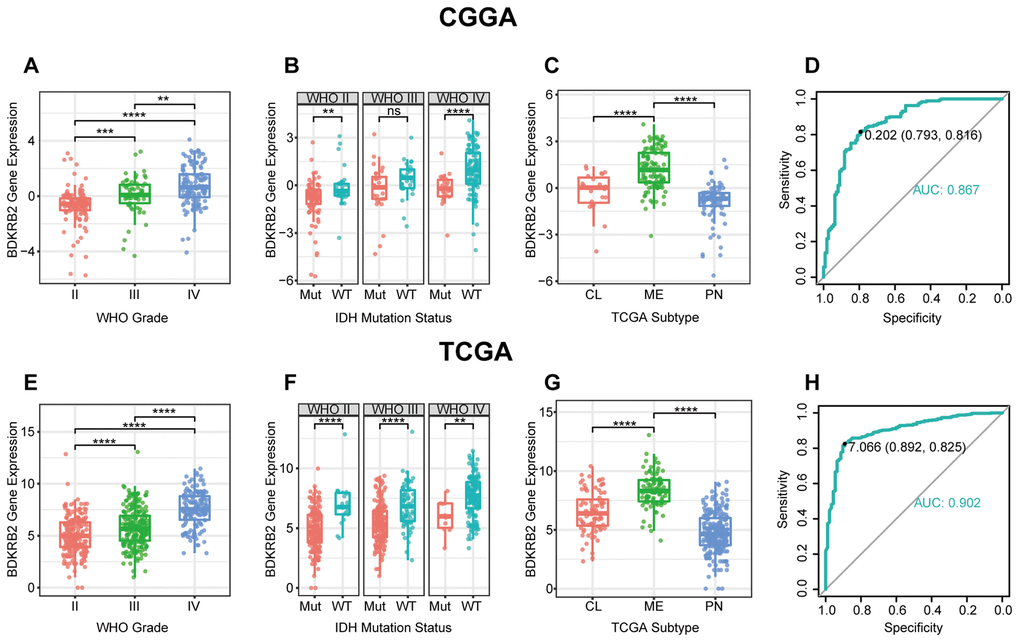
In the present study, we investigated the transcriptional expression profiles of BDKRB2 in 998 glioma patients and revealed that BDKRB2 expression showed a significantly positive correlation with the WHO grade of glioma. Furthermore, higher BDKRB2 expression was usually accompanied by a more aggressive and malignant phenotype in glioma, including GBM, IDH wildtype, and mesenchymal subtype. Moreover, higher BDKRB2 expression indicated a significantly shorter survival for patients with glioma across different WHO grades. These findings suggested that BDKRB2 played a vital role in the malignant progression of glioma, in line with other malignancies reported previously. Understanding the molecular mechanism of BDKRB2 in glioma may provide a novel therapeutic target to overcome this fatal disease.
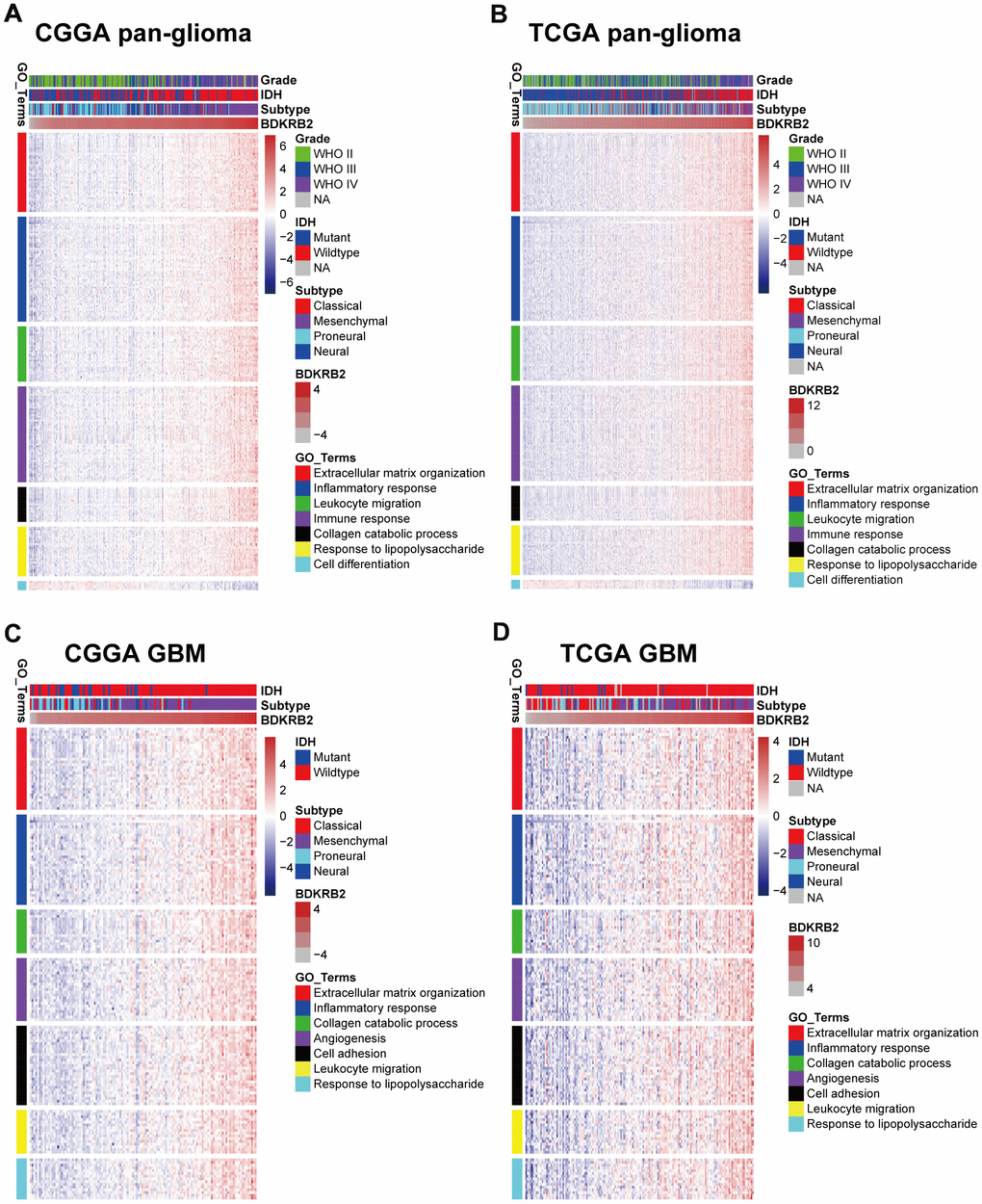
To elucidate the biological function of BDKRB2 in glioma, we further performed GO and GSEA analysis. GO analysis revealed that BDKRB2 was highly associated with extracellular matrix organization and collagen catabolic process in both pan-glioma and GBM, suggesting that glioma cells through their interactions with BDKRB2 might acquire functions that enhance matrix remodeling, cell migration, invasion, and tumor progression, consistent with the results presented by Montana et al. [28]. They concluded that the activation of BDKRB2 in glioma cells caused intracellular Ca2+ oscillations and subsequently enhanced glioma cell migration/invasion. In other types of malignancies, BDKRB2 also exhibited the biological function of promoting cell migration, invasion, and metastasis in hepatocellular carcinoma [12, 13], gastric cancer [14, 16], colorectal cancer [17], prostate cancer [18], head and neck squamous cell carcinoma [20] and chondrosarcoma [21]. In addition, GSEA analyses revealed remarkable evidence that BDKRB2 expression was particularly correlated with EMT, which had been extensively confirmed to play a key role not only in glioma migration/invasion but also in glioma recurrence and therapeutic resistance [39–41]. These results enlightened us that BDKRB2 might promote tumorigenesis and glioma progression mainly through modulating the EMT signaling pathway. Besides, GO and GSEA also revealed that BDKRB2 played a vital role in the tumor-induced inflammatory response in both pan-glioma and GBM subgroup, which might be another mechanism for the oncogenic role of BDKRB2 in glioma.
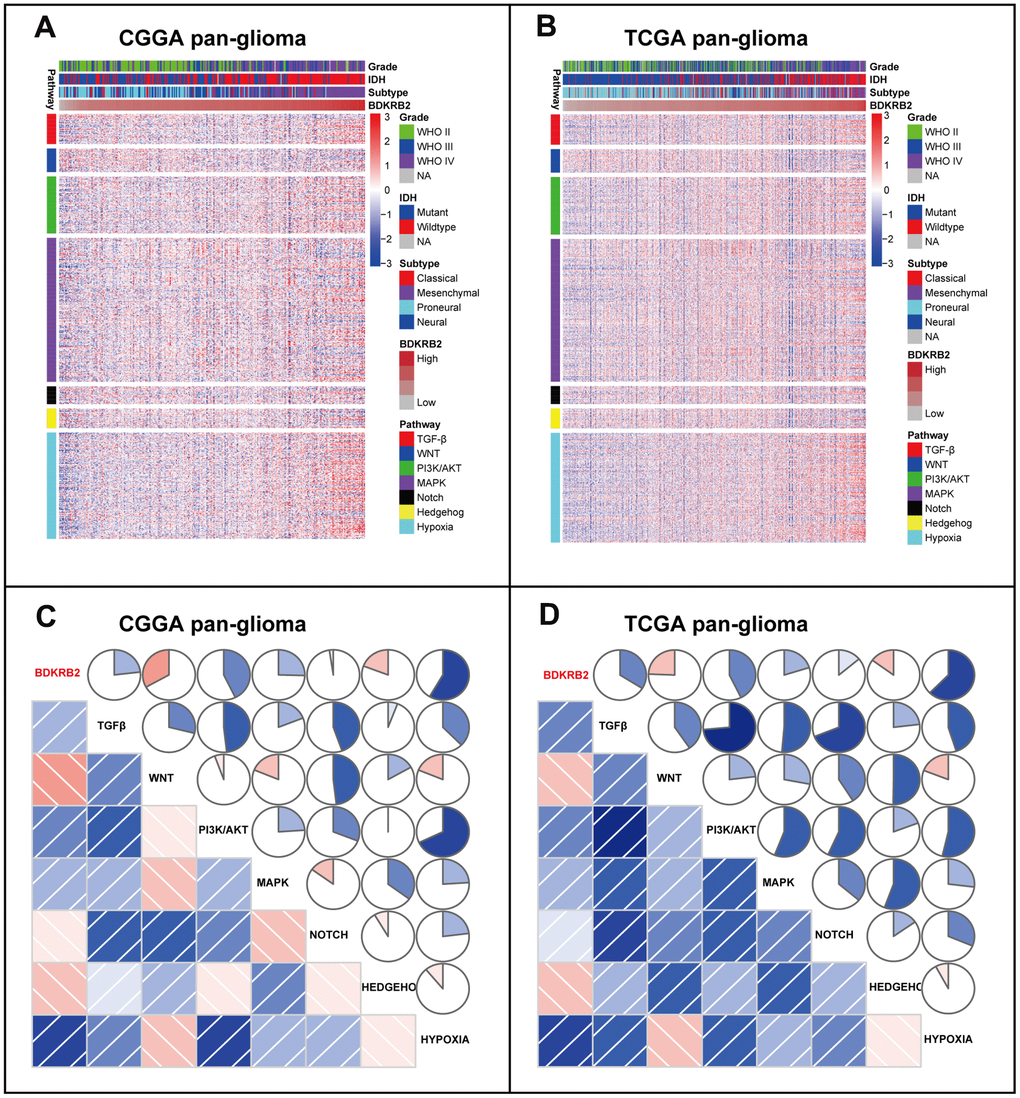
To further validate the role of BDKRB2 in the glioma EMT process, we selected a series of EMT-related signaling pathways and biomarker, which were then analyzed to determine their interaction with BDKRB2. We found that BDKRB2 expression showed a robust correlation with PI3K/AKT, hypoxia, and TGF-β signaling pathway, suggesting that BDKRB2 might promote the EMT process through these pathways. Moreover, most EMT biomarkers, including N-cadherin, snail, slug, vimentin, TWIST1, and TWIST2, were significantly correlated with BDKRB2, which suggested that BDKRB2 might profoundly interact with these key molecules of EMT, further confirming the involvement of BDKRB2 in glioma EMT. Thus, our findings might bring a novel EMT target for potential glioma treatment.
In conclusion, the present study demonstrated that BDKRB2 expression was associated with more malignant glioma phenotypes and predicted much worse survival for patients. Moreover, BDKRB2 was significantly associated with the EMT process. However, a limitation of the current study was that no experimental validation was performed. Further in-vitro and in-vivo studies are needed to validate its role in glioma.
Materials and Methods
Sample and data collection
Transcriptome and clinical data of glioma patients were available on Chinese Glioma Genome Atlas (CGGA) website (http://www.cgga.org.cn/) [42, 43] and TCGA website (http://cancergenome.nih.gov/) [44]. A total of 998 glioma patients, including 301 CGGA microarray data (GeneSpring GX 11.0 normalization) and 697 TCGA RNAseq data (RSEM normalization, level 3), were enrolled. The baseline characteristics of patients in both cohorts were described in Supplementary Table 5. This study was based on two large public databases, with no use of personally identifiable information. The ethics approval was waived by the Ethics Committee of Shenzhen People’s Hospital.
Statistical analysis
For TCGA cohort, RSEM RNAseq data were log2 transformed. For CGGA cohort, microarray data (already normalized and centered by data provider) were directly analyzed. Statistical analysis was performed with R language. Multiple R packages, including ggplot2, pROC, pheatmap, corrgram, circlize, and survival, were used to generate figures. Cox proportional hazard regression analyses were performed with coxph function of survival package. The biological processes of BDKRB2-related genes were annotated using Gene Ontology (GO) (DAVID, https://david.ncifcrf.gov/) enrichment analysis. The hallmarks.all.v7.1.symbols.gmt gene set was selected for Gene Set Enrichment Analysis (GSEA, http://software.broadinstitute.org/). All 301 samples in CGGA and 697 in TCGA were included in pan-glioma GSEA analysis, and 128 GBM samples in CGGA and 167 in TCGA were included in GBM GSEA analysis. The number of permutations was 1000. The enrichment statistic was set as weighted, and the metric for ranking genes was set as Pearson. All statistical tests were two-sided, and a p-value of < 0.05 indicated a statistical significance.
Supplementary Materials
Author Contributions
Ying Yang, Jin Wang, Aijun Shan, and Fei Shi made substantial contributions to the study conception and design. Ying Yang, Jin Wang, Shihai Xu, and Wen Lv performed data acquisition, data analysis, drafted the manuscript, and revised it critically. All authors have read and approved the final manuscript.
Acknowledgments
We appreciate the generosity of CGGA project and TCGA project for sharing data.
Conflicts of Interest
The authors declare no conflicts of interest.
Funding
This work was supported by Medical Scientific Research Foundation of Shenzhen Health Commission (szfz2018022), Shenzhen Science and Technology Innovation Foundation (JCYJ20190806150005453), and Futian Public Welfare Scientific Research Project (FTWS2020099).
References
- 1. Meng X, Zhao Y, Han B, Zha C, Zhang Y, Li Z, Wu P, Qi T, Jiang C, Liu Y, Cai J. Dual functionalized brain-targeting nanoinhibitors restrain temozolomide-resistant glioma via attenuating EGFR and MET signaling pathways. Nat Commun. 2020; 11:594. https://doi.org/10.1038/s41467-019-14036-x [PubMed]
- 2. Yang W, Wu PF, Ma JX, Liao MJ, Wang XH, Xu LS, Xu MH, Yi L. Sortilin promotes glioblastoma invasion and mesenchymal transition through GSK-3β/β-catenin/twist pathway. Cell Death Dis. 2019; 10:208. https://doi.org/10.1038/s41419-019-1449-9 [PubMed]
- 3. Wei J, Ouyang X, Tang Y, Li H, Wang B, Ye Y, Jin M, Al Azab M, Li W, Li X. ER-stressed MSC displayed more effective immunomodulation in RA CD4+CXCR5+ICOS+ follicular helper-like T cells through higher PGE2 binding with EP2/EP4. Mod Rheumatol. 2020; 30:509–16. https://doi.org/10.1080/14397595.2019.1651446 [PubMed]
- 4. Tome-Garcia J, Erfani P, Nudelman G, Tsankov AM, Katsyv I, Tejero R, Zhang B, Walsh M, Friedel RH, Zaslavsky E, Tsankova NM. Analysis of chromatin accessibility uncovers TEAD1 as a regulator of migration in human glioblastoma. Nat Commun. 2018; 9:4020. https://doi.org/10.1038/s41467-018-06258-2 [PubMed]
- 5. Leeb-Lundberg LM, Marceau F, Müller-Esterl W, Pettibone DJ, Zuraw BL. International union of pharmacology. Xlv. Classification of the kinin receptor family: from molecular mechanisms to pathophysiological consequences. Pharmacol Rev. 2005; 57:27–77. https://doi.org/10.1124/pr.57.1.2 [PubMed]
- 6. Couture R, Blaes N, Girolami JP. Kinin receptors in vascular biology and pathology. Curr Vasc Pharmacol. 2014; 12:223–48. https://doi.org/10.2174/1570161112666140226121627 [PubMed]
- 7. Nicoletti NF, Sénécal J, da Silva VD, Roxo MR, Ferreira NP, de Morais RL, Pesquero JB, Campos MM, Couture R, Morrone FB. Primary role for kinin B1 and B2 receptors in glioma proliferation. Mol Neurobiol. 2017; 54:7869–82. https://doi.org/10.1007/s12035-016-0265-9 [PubMed]
- 8. Regoli D, Plante GE, Gobeil F
Jr . Impact of kinins in the treatment of cardiovascular diseases. Pharmacol Ther. 2012; 135:94–111. https://doi.org/10.1016/j.pharmthera.2012.04.002 [PubMed] - 9. da Costa PL, Sirois P, Tannock IF, Chammas R. The role of kinin receptors in cancer and therapeutic opportunities. Cancer Lett. 2014; 345:27–38. https://doi.org/10.1016/j.canlet.2013.12.009 [PubMed]
- 10. Zhou Y, Wang W, Wei R, Jiang G, Li F, Chen X, Wang X, Long S, Ma D, Xi L. Serum bradykinin levels as a diagnostic marker in cervical cancer with a potential mechanism to promote VEGF expression via BDKRB2. Int J Oncol. 2019; 55:131–41. https://doi.org/10.3892/ijo.2019.4792 [PubMed]
- 11. Dubuc C, Savard M, Bovenzi V, Lessard A, Fortier A, Côté J, Neugebauer W, Rizzolio F, Geha S, Giordano A, Chemtob S, Gobeil F. Targeting intracellular B2 receptors using novel cell-penetrating antagonists to arrest growth and induce apoptosis in human triple-negative breast cancer. Oncotarget. 2018; 9:9885–906. https://doi.org/10.18632/oncotarget.24009 [PubMed]
- 12. Zhao J, Dong QZ, Zhong F, Cai LL, Qin ZY, Liu Y, Lin CZ, Qin LX, He FC. NMI promotes hepatocellular carcinoma progression via BDKRB2 and MAPK/ERK pathway. Oncotarget. 2017; 8:12174–85. https://doi.org/10.18632/oncotarget.14556 [PubMed]
- 13. Chen Y, Yu Y, Sun S, Wang Z, Liu P, Liu S, Jiang J. Bradykinin promotes migration and invasion of hepatocellular carcinoma cells through TRPM7 and MMP2. Exp Cell Res. 2016; 349:68–76. https://doi.org/10.1016/j.yexcr.2016.09.022 [PubMed]
- 14. Wang G, Sun J, Liu G, Fu Y, Zhang X. Bradykinin promotes cell proliferation, migration, invasion, and tumor growth of gastric cancer through ERK signaling pathway. J Cell Biochem. 2017; 118:4444–53. https://doi.org/10.1002/jcb.26100 [PubMed]
- 15. Wang D, Luo L, Guo J. miR-129-1-3p inhibits cell migration by targeting BDKRB2 in gastric cancer. Med Oncol. 2014; 31:98. https://doi.org/10.1007/s12032-014-0098-1 [PubMed]
- 16. Jiang J, Liu W, Guo X, Zhang R, Zhi Q, Ji J, Zhang J, Chen X, Li J, Zhang J, Gu Q, Liu B, Zhu Z, Yu Y. IRX1 influences peritoneal spreading and metastasis via inhibiting BDKRB2-dependent neovascularization on gastric cancer. Oncogene. 2011; 30:4498–508. https://doi.org/10.1038/onc.2011.154 [PubMed]
- 17. Wang G, Ye Y, Zhang X, Song J. Bradykinin stimulates IL-6 production and cell invasion in colorectal cancer cells. Oncol Rep. 2014; 32:1709–14. https://doi.org/10.3892/or.2014.3366 [PubMed]
- 18. Yu HS, Lin TH, Tang CH. Bradykinin enhances cell migration in human prostate cancer cells through B2 receptor/PKCδ/c-Src dependent signaling pathway. Prostate. 2013; 73:89–100. https://doi.org/10.1002/pros.22544 [PubMed]
- 19. Sgnaolin V, Pereira TC, Bogo MR, Zanin R, Battastini AM, Morrone FB, Campos MM. Functional and molecular characterization of kinin B1 and B2 receptors in human bladder cancer: implication of the PI3Kγ pathway. Invest New Drugs. 2013; 31:812–22. https://doi.org/10.1007/s10637-012-9907-6 [PubMed]
- 20. Zhang W, Bhola N, Kalyankrishna S, Gooding W, Hunt J, Seethala R, Grandis JR, Siegfried JM. Kinin b2 receptor mediates induction of cyclooxygenase-2 and is overexpressed in head and neck squamous cell carcinomas. Mol Cancer Res. 2008; 6:1946–56. https://doi.org/10.1158/1541-7786.MCR-07-2197 [PubMed]
- 21. Yang WH, Chang JT, Hsu SF, Li TM, Cho DY, Huang CY, Fong YC, Tang CH. Bradykinin enhances cell migration in human chondrosarcoma cells through BK receptor signaling pathways. J Cell Biochem. 2010; 109:82–92. https://doi.org/10.1002/jcb.22383 [PubMed]
- 22. Ishihara K, Kamata M, Hayashi I, Yamashina S, Majima M. Roles of bradykinin in vascular permeability and angiogenesis in solid tumor. Int Immunopharmacol. 2002; 2:499–509. https://doi.org/10.1016/s1567-5769(01)00193-x [PubMed]
- 23. Ikeda Y, Hayashi I, Kamoshita E, Yamazaki A, Endo H, Ishihara K, Yamashina S, Tsutsumi Y, Matsubara H, Majima M. Host stromal bradykinin B2 receptor signaling facilitates tumor-associated angiogenesis and tumor growth. Cancer Res. 2004; 64:5178–85. https://doi.org/10.1158/0008-5472.CAN-03-3589 [PubMed]
- 24. Yu HS, Wang SW, Chang AC, Tai HC, Yeh HI, Lin YM, Tang CH. Bradykinin promotes vascular endothelial growth factor expression and increases angiogenesis in human prostate cancer cells. Biochem Pharmacol. 2014; 87:243–53. https://doi.org/10.1016/j.bcp.2013.10.016 [PubMed]
- 25. Andoh T, Akira A, Saiki I, Kuraishi Y. Bradykinin increases the secretion and expression of endothelin-1 through kinin B2 receptors in melanoma cells. Peptides. 2010; 31:238–41. https://doi.org/10.1016/j.peptides.2009.12.003 [PubMed]
- 26. Desposito D, Chollet C, Taveau C, Descamps V, Alhenc-Gelas F, Roussel R, Bouby N, Waeckel L. Improvement of skin wound healing in diabetic mice by kinin B2 receptor blockade. Clin Sci (Lond). 2016; 130:45–56. https://doi.org/10.1042/CS20150295 [PubMed]
- 27. Chen S, Zhang L, Xu R, Ti Y, Zhao Y, Zhou L, Zhao J. TheBDKRB2 +9/-9 polymorphisms influence pro-inflammatory cytokine levels in knee osteoarthritis by altering TLR-2 expression: clinical and in vitro studies. Cell Physiol Biochem. 2016; 38:1245–56. https://doi.org/10.1159/000443072 [PubMed]
- 28. Montana V, Sontheimer H. Bradykinin promotes the chemotactic invasion of primary brain tumors. J Neurosci. 2011; 31:4858–67. https://doi.org/10.1523/JNEUROSCI.3825-10.2011 [PubMed]
- 29. Pillat MM, Oliveira MN, Motaln H, Breznik B, Glaser T, Lah TT, Ulrich H. Glioblastoma-mesenchymal stem cell communication modulates expression patterns of kinin receptors: possible involvement of bradykinin in information flow. Cytometry A. 2016; 89:365–75. https://doi.org/10.1002/cyto.a.22800 [PubMed]
- 30. Nicoletti NF, Erig TC, Zanin RF, Pereira TC, Bogo MR, Campos MM, Morrone FB. Mechanisms involved in kinin-induced glioma cells proliferation: the role of ERK1/2 and PI3K/Akt pathways. J Neurooncol. 2014; 120:235–44. https://doi.org/10.1007/s11060-014-1549-4 [PubMed]
- 31. Liu S, Wang Z, Wang Y, Fan X, Zhang C, Ma W, Qiu X, Jiang T. PD-1 related transcriptome profile and clinical outcome in diffuse gliomas. Oncoimmunology. 2017; 7:e1382792. https://doi.org/10.1080/2162402X.2017.1382792 [PubMed]
- 32. Wang Z, Zhang C, Liu X, Wang Z, Sun L, Li G, Liang J, Hu H, Liu Y, Zhang W, Jiang T. Molecular and clinical characterization of PD-L1 expression at transcriptional level via 976 samples of brain glioma. Oncoimmunology. 2016; 5:e1196310. https://doi.org/10.1080/2162402X.2016.1196310 [PubMed]
- 33. Li G, Wang Z, Zhang C, Liu X, Cai J, Wang Z, Hu H, Wu F, Bao Z, Liu Y, Zhao L, Liang T, Yang F, et al. Molecular and clinical characterization of TIM-3 in glioma through 1,024 samples. Oncoimmunology. 2017; 6:e1328339. https://doi.org/10.1080/2162402X.2017.1328339 [PubMed]
- 34. Liu S, Zhang C, Maimela NR, Yang L, Zhang Z, Ping Y, Huang L, Zhang Y. Molecular and clinical characterization of CD163 expression via large-scale analysis in glioma. Oncoimmunology. 2019; 8:1601478. https://doi.org/10.1080/2162402X.2019.1601478 [PubMed]
- 35. Jiang Y, Zhou J, Hou D, Luo P, Gao H, Ma Y, Chen YS, Li L, Zou D, Zhang H, Zhang Y, Jing Z. Prosaposin is a biomarker of mesenchymal glioblastoma and regulates mesenchymal transition through the TGF-β1/smad signaling pathway. J Pathol. 2019; 249:26–38. https://doi.org/10.1002/path.5278 [PubMed]
- 36. Wang J, Xu S, Lv W, Shi F, Mei S, Shan A, Xu J, Yang Y. Uridine phosphorylase 1 is a novel immune-related target and predicts worse survival in brain glioma. Cancer Med. 2020; 9:5940–47. https://doi.org/10.1002/cam4.3251 [PubMed]
- 37. Gonzalez DM, Medici D. Signaling mechanisms of the epithelial-mesenchymal transition. Sci Signal. 2014; 7:re8. https://doi.org/10.1126/scisignal.2005189 [PubMed]
- 38. Xu J, Zhang Z, Qian M, Wang S, Qiu W, Chen Z, Sun Z, Xiong Y, Wang C, Sun X, Zhao R, Xue H, Li G. Cullin-7 (CUL7) is overexpressed in glioma cells and promotes tumorigenesis via NF-κB activation. J Exp Clin Cancer Res. 2020; 39:59. https://doi.org/10.1186/s13046-020-01553-7 [PubMed]
- 39. Li C, Zheng H, Hou W, Bao H, Xiong J, Che W, Gu Y, Sun H, Liang P. Long non-coding RNA linc00645 promotes TGF-β-induced epithelial-mesenchymal transition by regulating miR-205-3p-ZEB1 axis in glioma. Cell Death Dis. 2019; 10:717. https://doi.org/10.1038/s41419-019-1948-8 [PubMed]
- 40. Li H, Li J, Chen L, Qi S, Yu S, Weng Z, Hu Z, Zhou Q, Xin Z, Shi L, Ma L, Huang A, Lu Y. HERC3-mediated SMAD7 ubiquitination degradation promotes autophagy-induced EMT and chemoresistance in glioblastoma. Clin Cancer Res. 2019; 25:3602–16. https://doi.org/10.1158/1078-0432.CCR-18-3791 [PubMed]
- 41. Li H, Li J, Zhang G, Da Q, Chen L, Yu S, Zhou Q, Weng Z, Xin Z, Shi L, Ma L, Huang A, Qi S, Lu Y. HMGB1-induced p62 overexpression promotes snail-mediated epithelial-mesenchymal transition in glioblastoma cells via the degradation of GSK-3β. Theranostics. 2019; 9:1909–22. https://doi.org/10.7150/thno.30578 [PubMed]
- 42. Wang Y, Wang Y, Fan X, Li S, Liu X, Wang J, Jiang T. Putamen involvement and survival outcomes in patients with insular low-grade gliomas. J Neurosurg. 2017; 126:1788–94. https://doi.org/10.3171/2016.5.JNS1685 [PubMed]
- 43. Fang S, Liang J, Qian T, Wang Y, Liu X, Fan X, Li S, Wang Y, Jiang T. Anatomic location of tumor predicts the accuracy of motor function localization in diffuse lower-grade gliomas involving the hand knob area. AJNR Am J Neuroradiol. 2017; 38:1990–97. https://doi.org/10.3174/ajnr.A5342 [PubMed]
- 44. Weinstein JN, Collisson EA, Mills GB, Shaw KR, Ozenberger BA, Ellrott K, Shmulevich I, Sander C, Stuart JM, and Cancer Genome Atlas Research Network. The cancer genome atlas pan-cancer analysis project. Nat Genet. 2013; 45:1113–20. https://doi.org/10.1038/ng.2764 [PubMed]