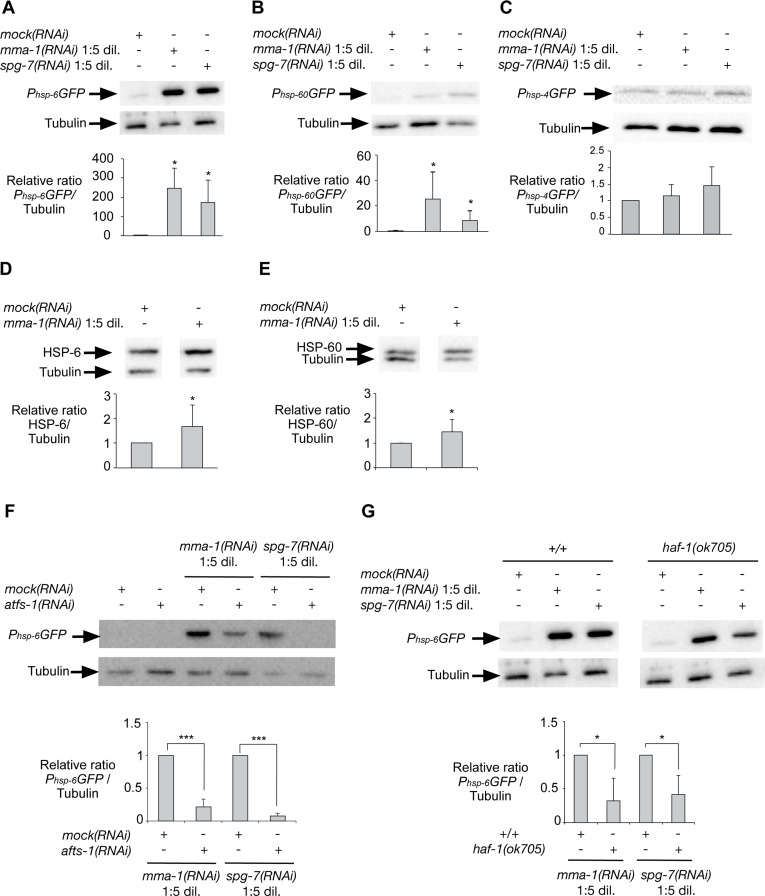
Figure 3.Inactivation of mma-1 by RNAi in C. elegans induces ATFS-1-dependent UPRmtWestern analysis of (A) Phsp-6GFP (mitochondrial Hsp70), (B) Phsp-60GFP (mitochondrial Hsp60) or (C) Phsp-4GFP (ER BiP) reporter strains treated with mock(RNAi), mma-1(RNAi) 1:5 dil. (diluted 1:5 (v/v) with mock(RNAi)) or spg-7(RNAi) 1:5 dil. (diluted 1:5 (v/v) with mock(RNAi)). Ratios of Phsp-6GFP/Tubulin, Phsp-60GFP/Tubulin and Phsp-4GFP/Tubulin relative to the mock(RNAi) treated animals are indicated (n = 5 for Phsp-6GFP + mma-1(RNAi); n = 7 for Phsp-6GFP + spg-7(RNAi); n = 6 for Phsp-60GFP + mma-1(RNAi); n = 9 for Phsp-60GFP + spg-1(RNAi); n = 5 for Phsp-4GFP). Western analysis of the effect of mma-1(RNAi) on endogenous (D) HSP-6 or (E) HSP-60 protein level. Ratios of HSP-6/Tubulin and HSP-60/Tubulin relative to the mock(RNAi) treated animals are indicated (n = 8 for HSP-6 and n = 10 for HSP-60). (F) Western analysis of the effect of ATFS-1 on mma-1(RNAi)-induced UPRmt. mma-1(RNAi) or spg-7(RNAi) were diluted either with mock(RNAi) or atfs-1(RNAi). Relative ratios of Phsp-6GFP/Tubulin are indicated (n = 7 for mma-1(RNAi) and n = 10 for spg-7(RNAi)). (G) Western analysis of the effect of HAF-1 on mma-1(RNAi)-induced UPRmt. Wild-type Phsp-6GFP reporter strain (+/+) or Phsp-6GFP reporter strain carrying the haf-1(ok705) loss-of-function mutation were analyzed. The relative ratios of Phsp-6GFP/Tubulin are indicated (n = 5 for mma-1(RNAi) and n = 6 for spg-7(RNAi)) (For all panels, average values are shown and error bars indicate s.d.; *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001 by one sample t-test).