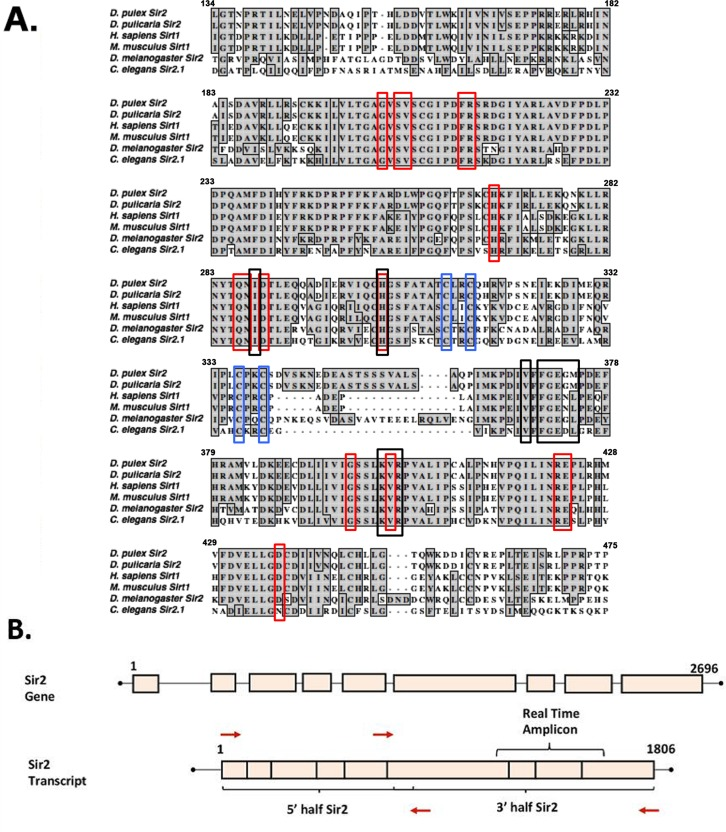
Figure 1.Sequence alignment of the Sirt1 homologs from invertebrates and vertebrates(A) Sir2 related proteins from 5 model organisms are aligned with Daphnia pulex Sir2 on the top. Red boxes depict residues involved in NAD+ binding, black boxes depict substrate-binding residues, and blue boxes depict zinc-binding residues. Alignments made using ClustalW in MacVector. The accession numbers of the sequences used are: Daphnia pulex Sir2: DappuDraft_290541 (wFleaBase), Daphnia pulicaria Sir2: KT960973, H. sapiens SIRT1: NP_036370.2, C. elegans SIR2.1: NP_001255484.1, D. melanogaster Sir2: AAC79684.1, and M. musculus SIRT1: NP_062786.1. (B) Schematic representation of Daphnia Sir2 gene and Sir2 transcript. PCR primers (red arrows) were designed such that the open reading frame (ORF) was PCR amplified in two parts, a 5′ half and a 3′ half as indicated. Real-time amplicon is the region of the transcript amplified via real time PCR for expression analysis.