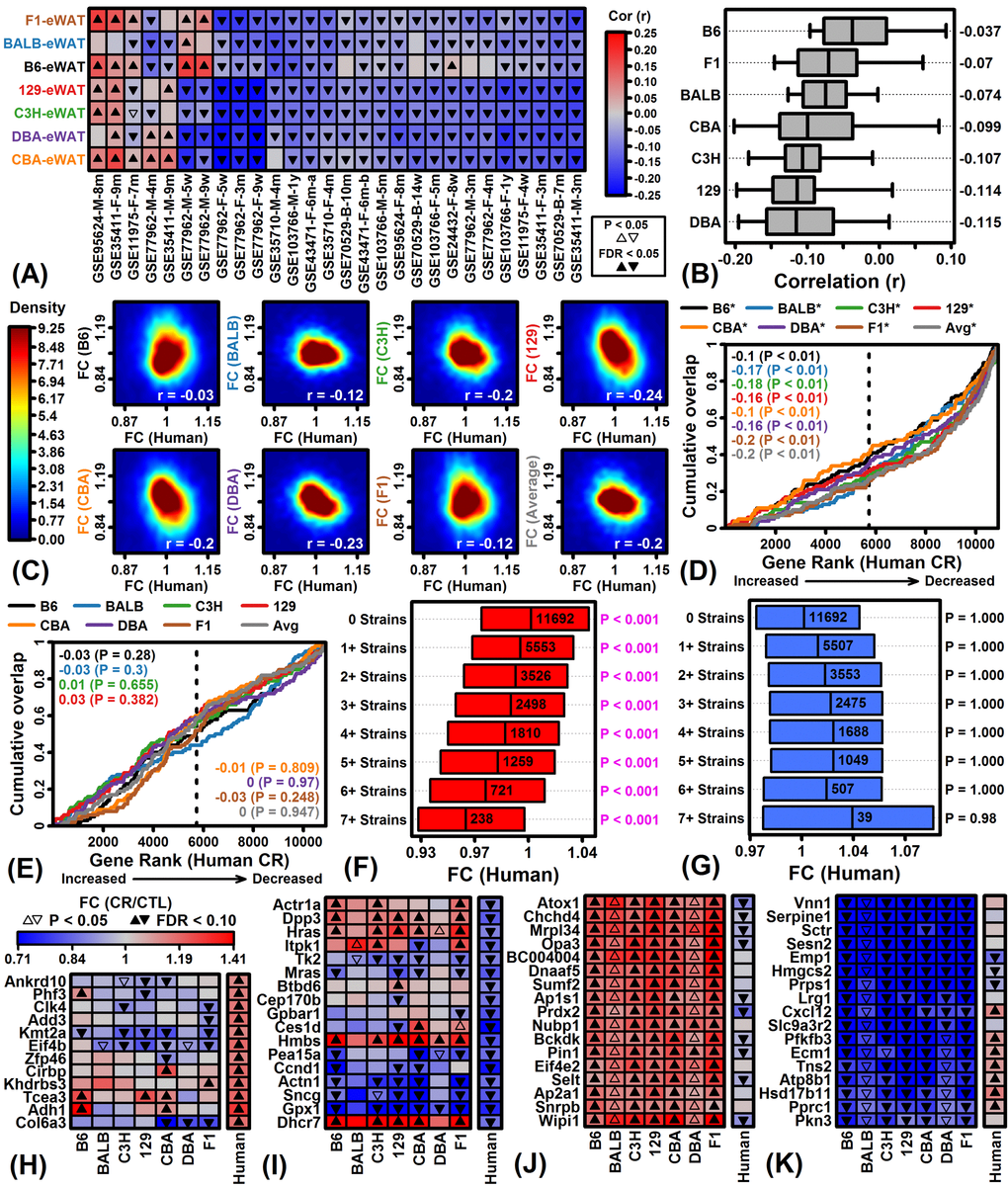
Figure 6.Comparison of CR responses in mouse eWAT and human scWAT. (A) Spearman rank correlations. FC estimates (CR/CTL) in 7 mouse strains (left margin) were compared to those in 28 human experiments (bottom margin). Bottom labels indicate GEO accession identifier, sex, and length of dietary intervention. (B) Correlation estimates by strain. Bars span the middle 50% of correlations for each strain (whiskers: middle 80%; right margin: median correlation). (C) FC scatterplots. FC estimates in each strain are compared to average FC estimates across the 28 human experiments. Colors denote gene density (see scale; lower right: Spearman rank correlation). (D) Gene set enrichment analysis (GSEA) of top 100 CR-increased genes in each mouse strain. (E) GSEA of top 100 CR-decreased genes in each mouse strain. In (D) and (E), genes were ranked according to their expression change with CR in humans (horizontal axis) and cumulative overlap was examined with respect to 100 CR-increased/decreased genes from each strain (vertical axis) (*P < 0.05, upper margin labels; enrichment statistics with p-values listed in each figure). Positive enrichment statistics indicate significant overlap with respect to genes increased by CR in human scWAT, while negative statistics indicate significant overlap with respect to genes decreased by CR in human scWAT (dashed vertical line: number of CR-increased genes in human, FC > 1.00). (F) Human FC estimates of genes increased by CR in multiple mouse strains. (G) Human FC estimates of genes decreased by CR in multiple mouse strains. In (F) and (G), bars span the middle 50% of human FC estimates. Significant p-values indicate that the median human FC estimate is significantly different from 1.00 (Wilcoxon rank sum test). (H, I) Genes most strongly altered by CR in human scWAT. (J, K) Genes most consistently altered by CR across 7 mouse strains (eWAT).