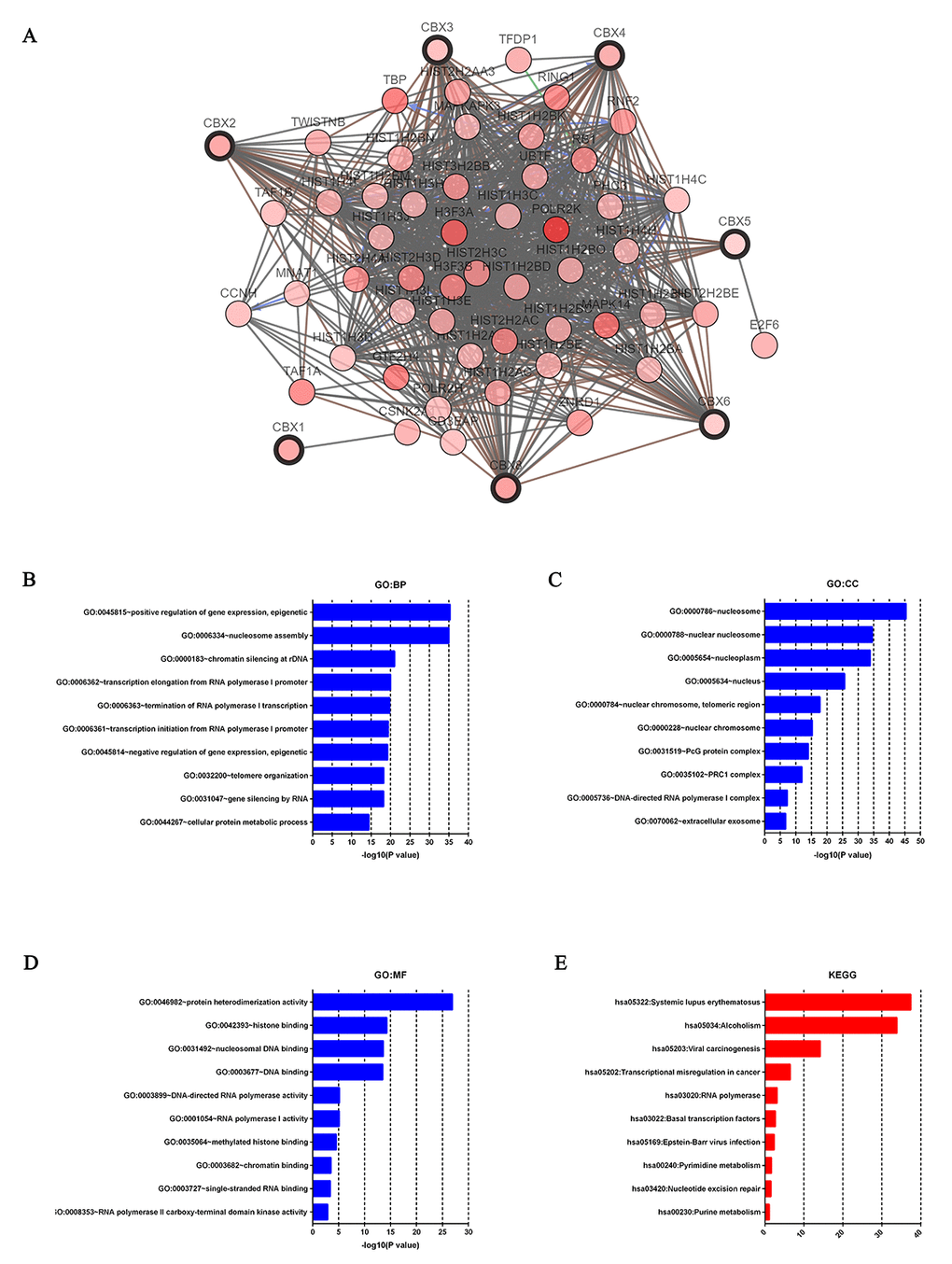
Figure 8.Predicted functions and pathways of the mutations in CBXs and their 50 frequently altered neighbor genes in HCC patients (c-BioPortal and DAVID). Network of CBXs mutations and their 50 frequently altered neighbor genes was constructed. DNA repair and DNA replication related genes including HIST1H3A, HIST1H2AB, H3F3A, HIST2H2AA3, HIST2H3C and HIST3H2BB were significantly related to CBXs mutations (A). GO functional enrichment analysis predicted three main functions of CBXs mutations and their 50 frequently altered neighbor genes, including biological process, cellular components and molecular functions (B-D). KEGG pathway analysis on CBXs and their 50 most frequently altered neighbor genes was shown at figure E.