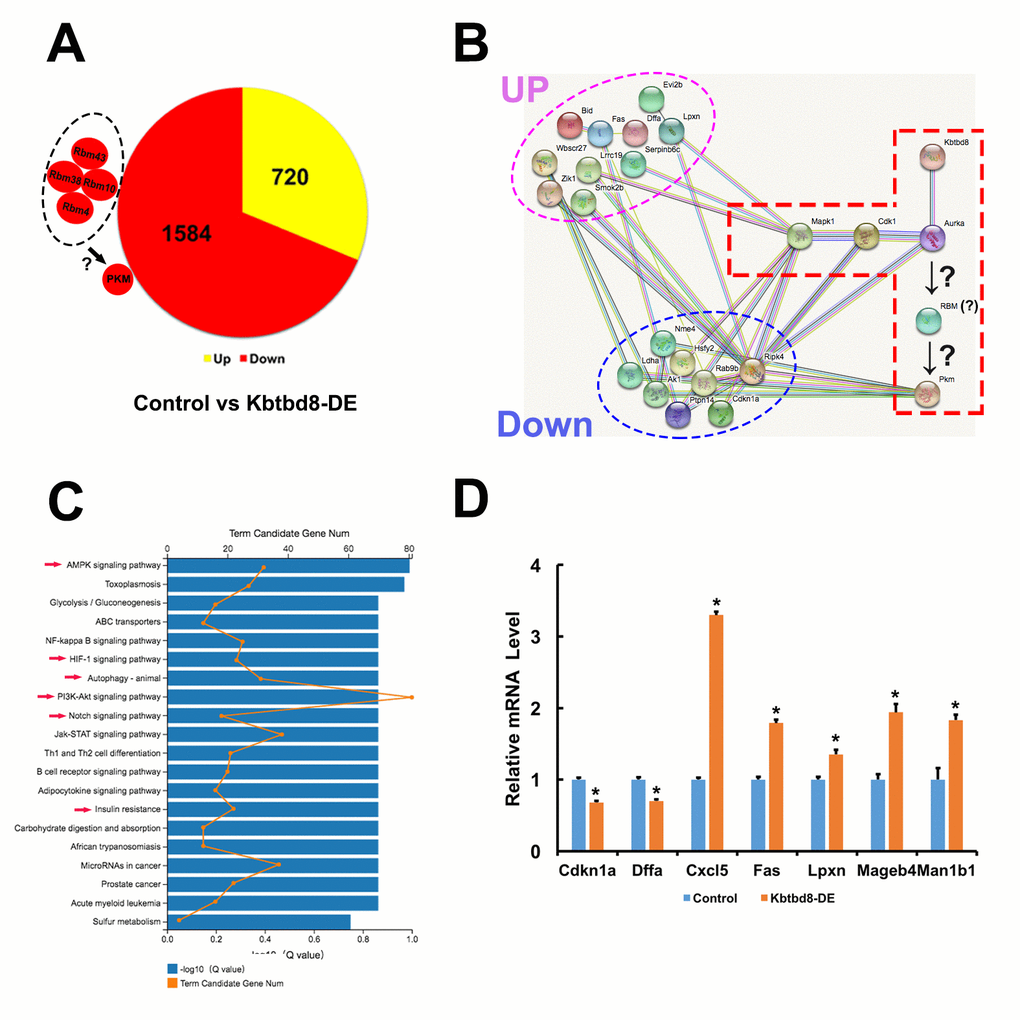
Figure 7.Transcriptome-wide characterization of KBTBD8-regulated pathways. A. Comparative mRNA sequencing in control and KBTBD8-depleted oocytes identified 2304 differentially-expressed genes (KBTBD8/control, log2 > 2). Among these, 1584 (69%) were downregulated while 720 (31%) were upregulated. Although the PKM log2 ratio was lower than 2 (90.83/50.51 = 1.84), its downregulation is consistent with our immunofluorescence and molecular analyses in KBTBD8-depleted oocytes. Also, the log2 ratios of 3 potential splicing enzymes for PKM1, RBM10, 38, and 43, were over or close to 2, so they were also considered as downregulated. Although the log2 ratio of RBM4 was only 0.73, based on its reported splicing activity on PKM1 we included it as another potential PKM1 splicing factor. B. Protein network showing the 18/200 top differentially expressed genes (log2 ratio > 5) that interact with the proteins in our pathway model. Of those, 10 were upregulated and 8 were downregulated. C. Overall pathway classification of all differentially-expressed genes. Multiple pathways, essential for the regulation of oocyte meiosis and quality, are represented; they include AMPK, HIF-1, Notch, PI3K-Akt, insulin resistance, and autophagy (red arrows). D. q-PCR expression analysis of 7 of the 200 differentially expressed genes verified the credibility of our sequencing data.