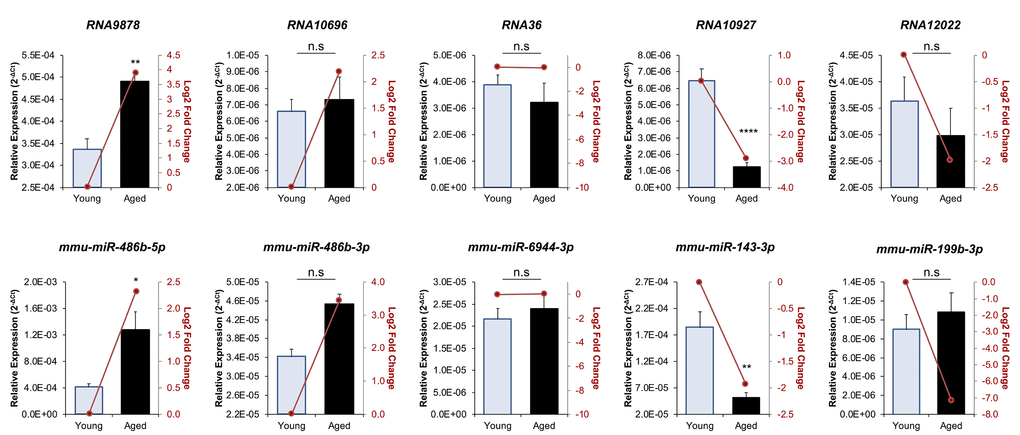
Figure 2.RT-qPCR validation of miRNA and endo-siRNA abundance in young and aged oocytes. To validate RNA-Seq data, five endo-siRNAs and five miRNAs were selected for quantification using RT-qPCR. Candidate sRNA included four representatives with increased expression in aged oocytes (RNA9878, RNA10696, mmu-miR-486b-5p, and mmu-miR-486b-3p), four with decreased expression in aged oocytes (RNA10927, RNA12022, mmu-miR-143-3p, and mmu-miR-199b-3p), and two that remained at equivalent levels (RNA36 and mmu-miR-6944-3p). cDNA generation and RT-qPCR experiments were performed in technical and biological triplicate, with each biological replicate comprising 10 oocytes randomly sampled from a pool of oocytes isolated from three animals. The U6 small nuclear RNA was employed as an endogenous control to normalize the expression levels of target sRNAs. Values are shown as a mean of all replicates ± SEM. Statistical analyses were performed using Student’s t-test, * p < 0.05, ** p < 0.01, **** p < 0.0001. Log2 fold changes based on RNA-Seq are represented as dark red line graphs while the relative abundance (2-ΔCt) of each sRNA determined by RT-qPCR is represented by columns.