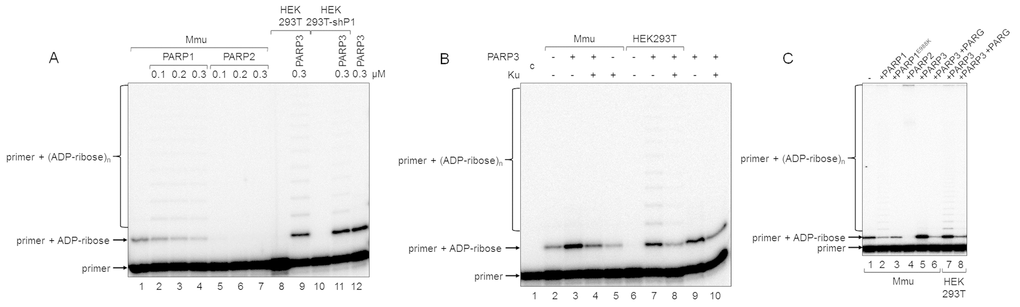
Figure 4.Influence of exogenous proteins on (ADP-ribosyl)ation of primer in WCEs. (A) WCE proteins in the absence or presence of extra recombinant proteins were incubated for 5 min with 100 nM DNA duplex dRP, 0.5 mM NAD+, and 5 mM spermine as described in the section ‘DNA (ADP-ribosyl)ation assay’. Recombinant PARP1, PARP2, or PARP3 were added to the extracts at the indicated concentrations prior to initiation of the (ADP-ribosyl)ation reaction. Lane 1, no extra recombinant proteins were added; lane 12, no extract proteins were added. (B) Mmu or HEK293T WCE proteins (0.5 mg/mL) were incubated for 5 min with 100 nM DNA duplex containing dRP moiety in the presence of 0.5 mM NAD+ and 5 mM spermine as described in the section ‘DNA (ADP-ribosyl)ation assay’. PARP3 and/or Ku (each at the final concentration of 300 nM) were added to the extracts prior to initiation of the (ADP-ribosyl)ation reaction. Lane 1 corresponds to the initial primer (control). (C) Mmu or HEK293T WCE proteins (0.5 mg/mL) were incubated at 37 °C for 5 min with 100 nM DNA duplex containing the dRP moiety in the presence of 0.5 mM NAD+ as described in the section ‘DNA (ADP-ribosyl)ation assay’. PARP1, PARP1E988K, PARP2, or PARP3 (the final concentrations of 300 nM) were initially added to the reaction mixtures and indicated. 50 nM PARG was added to some mixtures (lanes 6 and 8) after the reactions were stopped by the addition of EDTA, and the mixtures were incubated at 37 °C for another 10 min.