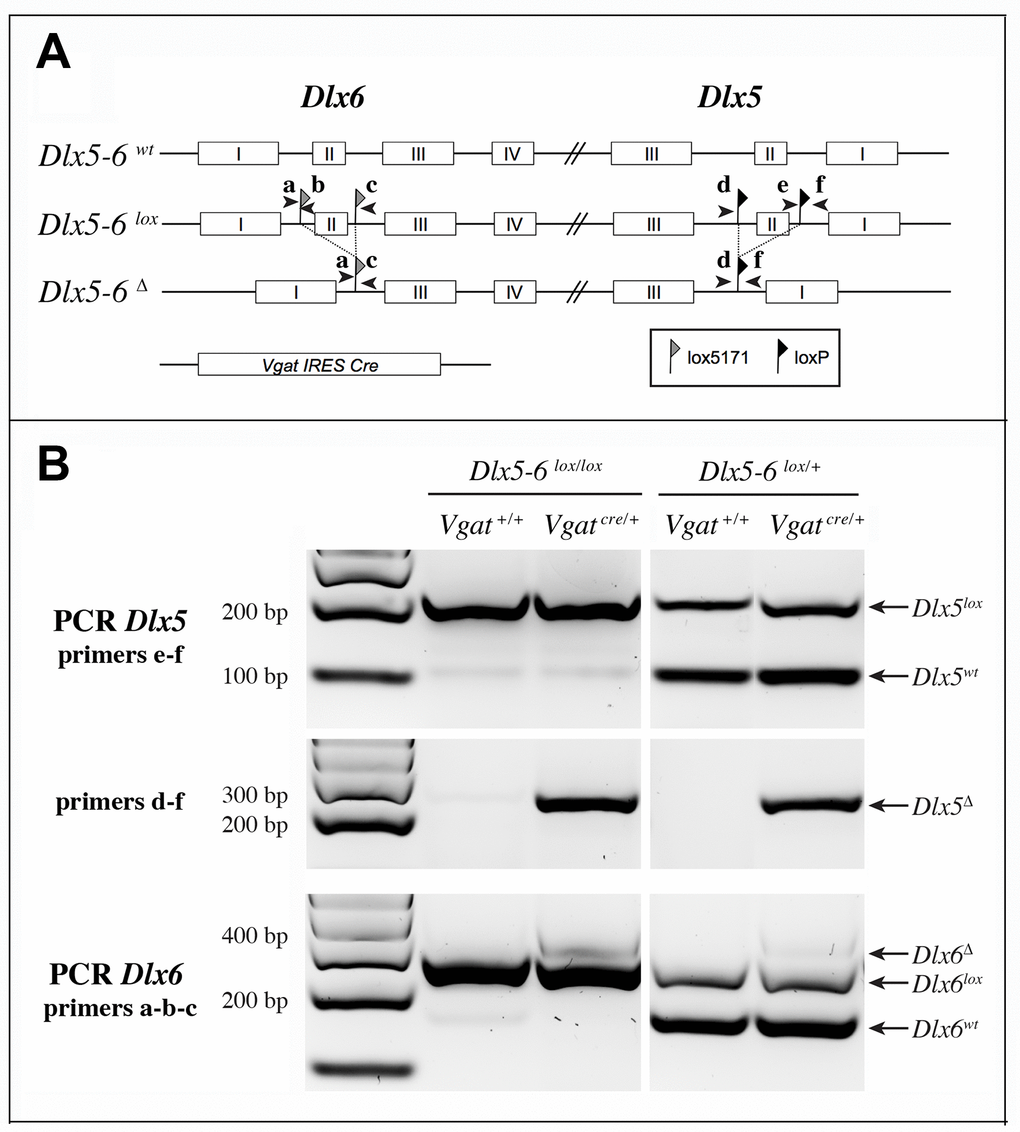
Figure 2.Strategy of Dlx5 and Dlx6 simultaneous invalidation and mouse genotyping. (A) Exons 2 of Dlx5 and Dlx6 were respectively framed with loxP and lox5171 sequences as described in [35]. In the presence of a Slc32a1-IRES-Cre(Vgat-Cre), exons II of Dlx5 and Dlx6 are deleted in GABAergic interneurons generating a Dlx5-6Δ allele. Arrowheads indicate the position and name (a to f) of the primers used for genotyping. (B) PCRs on cortical DNA extracts. Primers a, b, c and d, e, f were respectively utilized to reveal Dlx6 and Dlx5 recombination. The floxed and wild type Dlx5 alleles (primers e-f) were revealed in a separate PCR than that used to reveal the recombinant (Dlx5Δ) allele (primers d-f). Wild type, floxed and recombinant Dlx6 alleles were identified with a single PCR with primers a, b, c. In the presence of Cre recombinase, a band corresponding to the recombinant allele (Δ) can be detected for both Dlx5 (primers d-f) and Dlx6 (primers a, b, c) in VgatΔDlx5-6. In VgatΔDlx5-6/+ mice wild type and recombinant alleles are detected.