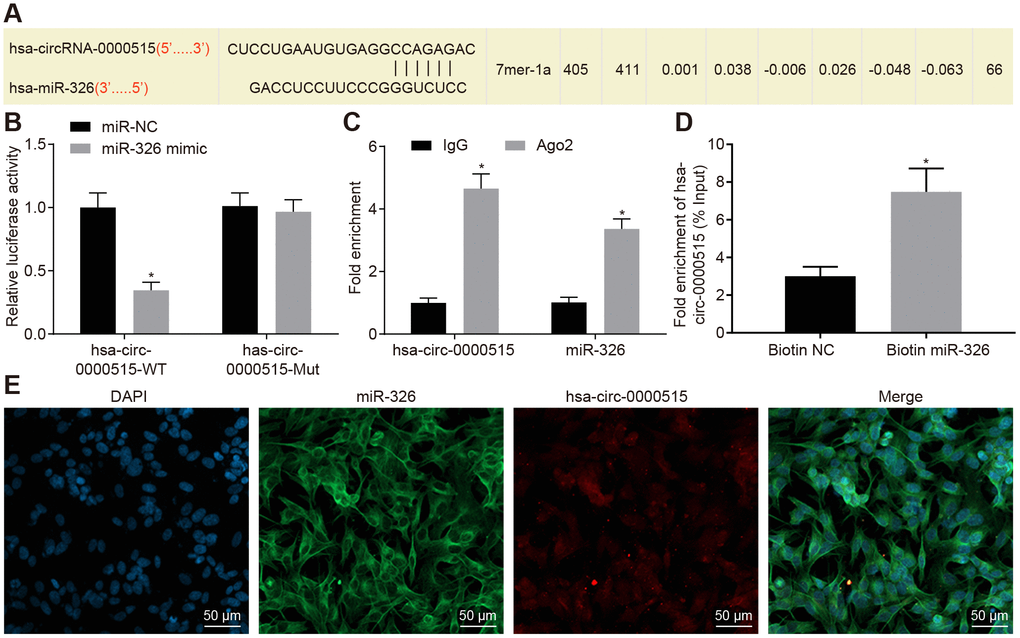
Figure 4.Hsa_circ_0000515 specifically binds to miR-326. (A) the miR-326 binding site in hsa_circ_0000515 predicted by bioinformatics websites (Starbase V2.0 and Circinteractome); (B) the relationship between hsa_circ_0000515 and miR-326 confirmed by dual luciferase reporter assay, *p < 0.05 vs. the si-NC group, (C) hsa_circ_0000515 and miR-326 co-immunoprecipitated with Ago2 revealed by RIP assay; (D) hsa_circ_0000515 pulled-down with biotin-labeled miR-326 by RNA pull-down assay, *p < 0.05 vs. the biotin-NC group; (E) FISH analysis of the localization of hsa_circ_0000515 and miR-326 in cervical cancer cells (200 ×). *p < 0.05 vs. the IgG or biotin-NC group. Measurement data were expressed as mean ± standard deviation. Unpaired t test was used to compare data with normal distribution and equal variance between two groups. The cell experiment is repeated three times independently.