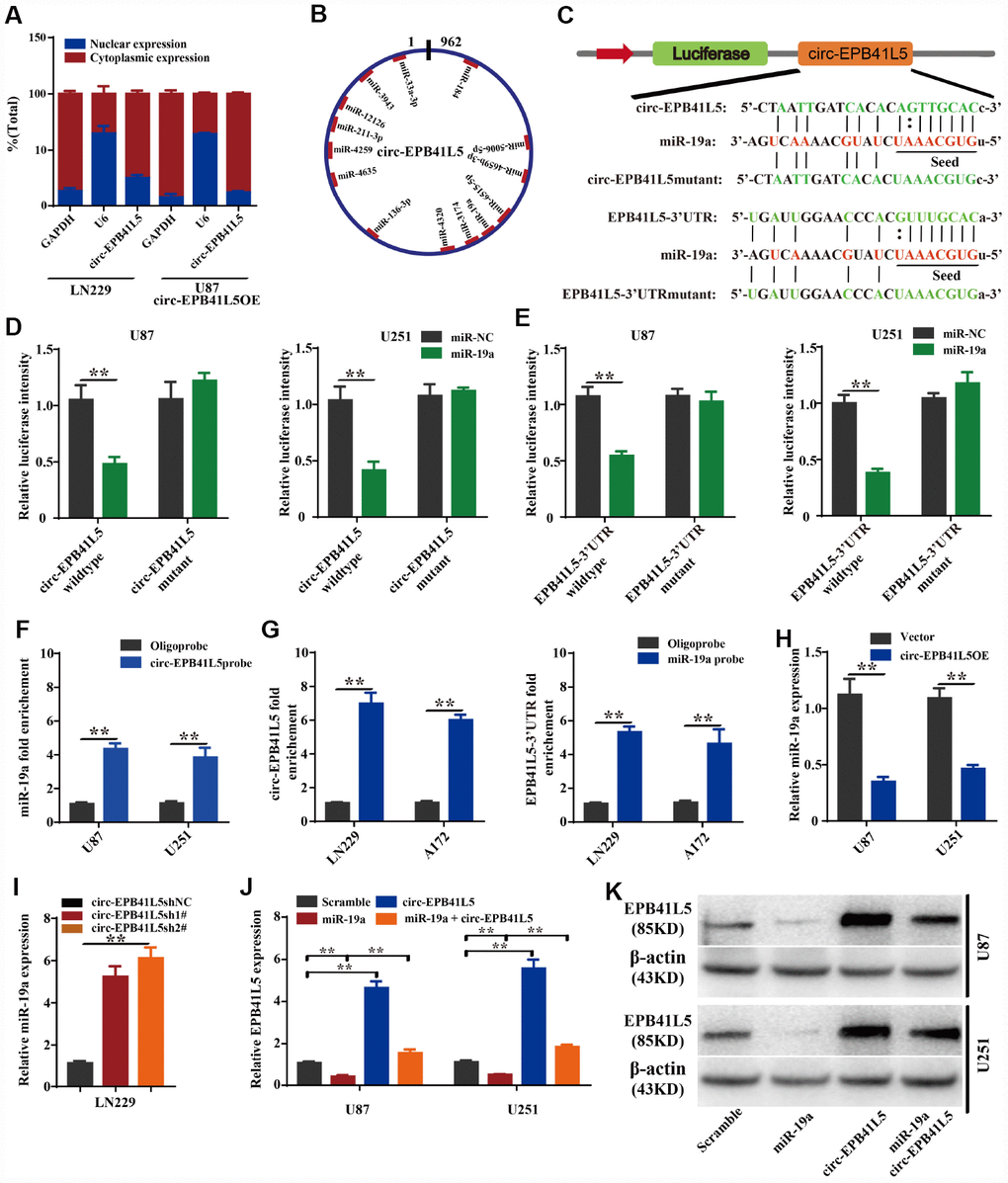
Figure 5.circ-EPB41L5 functions as a sponge of miR-19a to regulate the expression of EPB41L5. (A) Nuclear cytoplasm separation assay compared the abundance of circ-EPB41L5 in the nucleus and cytoplasm. Fractionation of LN229 and U87 cells followed by qRT-PCR. U6 RNA acted as a positive control for gene expression. (B) Schematic representation of the predicted binding sites of miRNAs related to circ-EPB41L5. (C) Schematic representation of circ-EPB41L5 or EPB41L5 3’-UTR wild-type (WT) and mutant (Mut) luciferase reporter vectors and miR-19a binding sites. (D, E) The relative luciferase activities were detected after co-transfection with miR-NCs or miR-19a mimics and circ-EPB41L5 or EPB41L5 3′-UTR WT and Mut luciferase reporter vectors in U87 and U251 cells. (F) Lysates from U87 and U251 cells were subjected to biotinylation-circ-EPB41L5 pulldown assay; qRT-PCR detected the expression of miR-19a. (G) Lysates from A172 and LN229 cells were subjected to biotinylation-miR-19a pulldown assay; qRT-PCR detected the expression of circ-EPB41L5 or EPB41L5-3’UTR. (H, I) qRT-PCR detected the expression of miR-19a in glioma cells transfected with circ-EPB41L5sh or circ-EPB41L5 overexpression plasmids. (J, K) qRT-PCR and WB assays detected the expression of EPB41L5 in glioma cells transfected with miR-19a mimics or circ-EPB41L5 vector. The data are the means±SEM of three experiments, *P<0.05, **P<0.01.