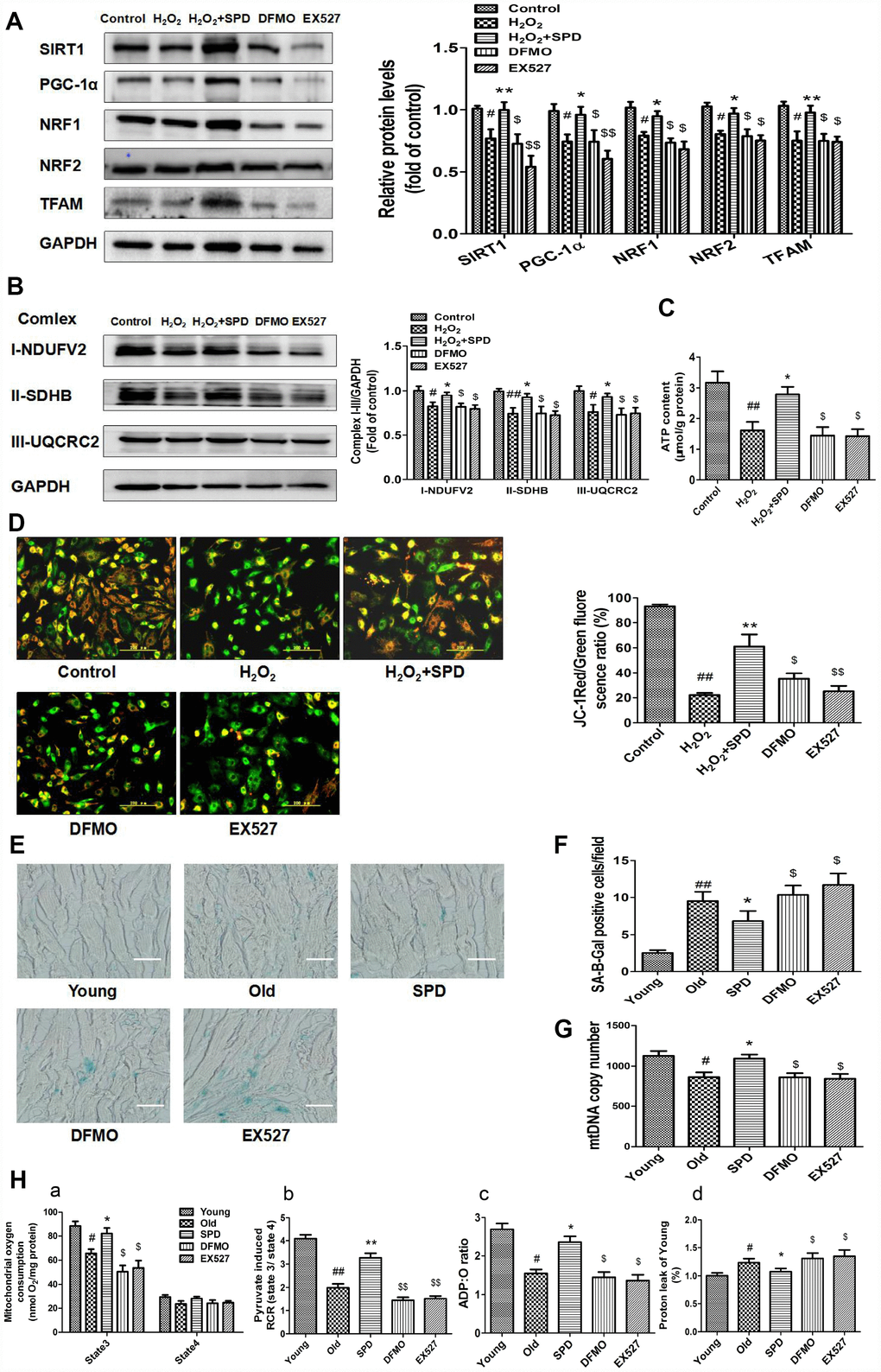
Figure 5.Inhibition of polyamine biogenesis and SIRT1 activity attenuates SPD-induced mitochondrial biogenesis and functional improvement in aging cardiomyocytes. For in vitro studies, NRMCs and H9C2 cells were cultured as follows: normal culture (Control), H2O2 treatment-induced aging (H2O2), H2O2 plus SPD (H2O2 + SPD), H2O2 plus SPD and DFMO (DFMO), or H2O2 plus SPD and EX527 (EX527). (A) Representative immunoblot bands for SIRT1, PGC-1α, NRF1, NRF2, and TFAM, and quantification of protein expression in NRMCs. GAPDH was used as loading control (n = 4). (B) Representative immunoblot bands for OXPHOS complexes I (NDUFV2), II (SDHB), and III (UQCRC2), and quantification of protein expression in NRMCs (n = 4). (C) ATP content measured by luminometry in NRMCs (n = 8). (D) Mitochondrial transmembrane potential (ΔΨm) detected by JC-1 fluorescence staining in H9C2 cells. Mean fluorescence intensity is displayed on the right of the graphs (n = 6). # P < 0.05 vs. Control, ## P < 0.01 vs. Control, * P < 0.05 vs. H2O2, ** P < 0.01 vs. H2O2, $ P < 0.05 vs. H2O2 + SPD, $$ P < 0.01 vs. H2O2 + SPD. For in vivo studies, the rats were divided into five groups: 1) young (3 months old), 2) old (24 months old), 3) SPD (24-months-old rats treated by SPD for 6 weeks), 4) DFMO (24-month-old rats treated with SPD and DFMO plus MGBG), and 5) EX527 (24-month-old rats treated with SPD and EX527). (E) Cardiac aging evaluated by SA-β-gal staining ex-vivo. (F) SA-β-gal staining quantification. Scale bars: 20 μm (n = 6). (G) Mitochondrial DNA (mtDNA) copy number detected by real-time PCR (n = 8). (H) Mitochondrial oxidative phosphorylation (OXPHOS) efficiency was evaluated based on mitochondrial State 3 and 4 oxygen consumption (a), respiratory control rate (RCR) (b), P/O ratio (c), and proton leakage (d); n = 8. # P < 0.05 vs. young, ## P < 0.01 vs. young, * P < 0.05 vs. old, ** P < 0.01 vs. old, $ P < 0.05 vs. SPD, $$ P < 0.01 vs. SPD. n = 6 for each group.