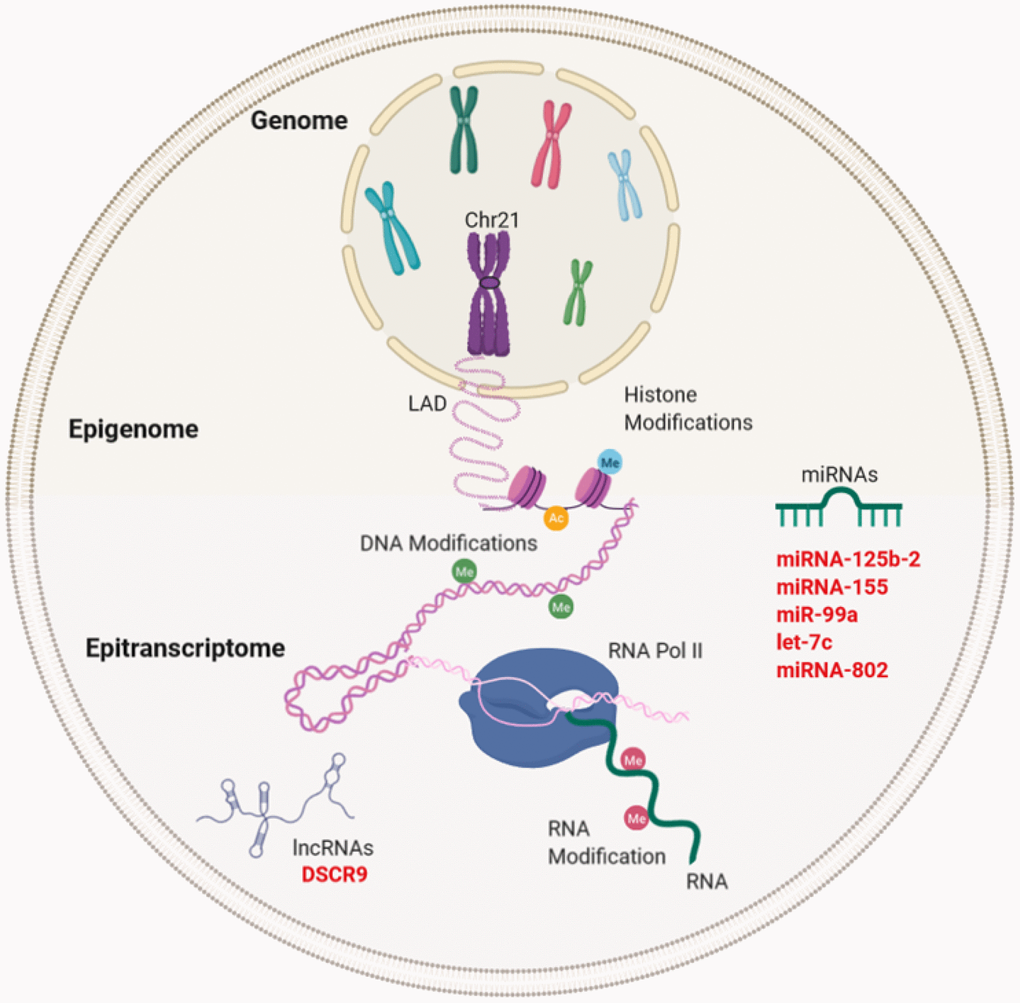
Figure 3.Gene expression regulation and examples of non-coding RNA in Down syndrome (DS) and Alzheimer's disease (AD). Gene regulation occurs through the genome, epigenome, and epitranscriptome. Beyond the DNA sequence, chromosomes are regulated by their locations or territories in the nucleus. The presence of an extra chromosome can alter the chromatin structure, ultimately affecting the transcription of the entire genome. At the epigenetic level, gene expression is regulated by reversible modifications of histones within nucleosomes that include methylation, acetylation, phosphorylation, ubiquitination, and sumoylation. Chemical modifications in RNA regulate the fate of transcription through a network of methyltransferases (writers), demethylases (drafts) and specific RNA reading proteins. The regulation of expression by ncRNAs can be affected at different levels. In red letters, the miRNAs and lncRNA encoded in Chr21 linked to DS and AD (miRNA-125b-a, miRNA-155, miR-99a, let-7c, and miRNA-802) and lncRNA (DSCR9) are highlighted.