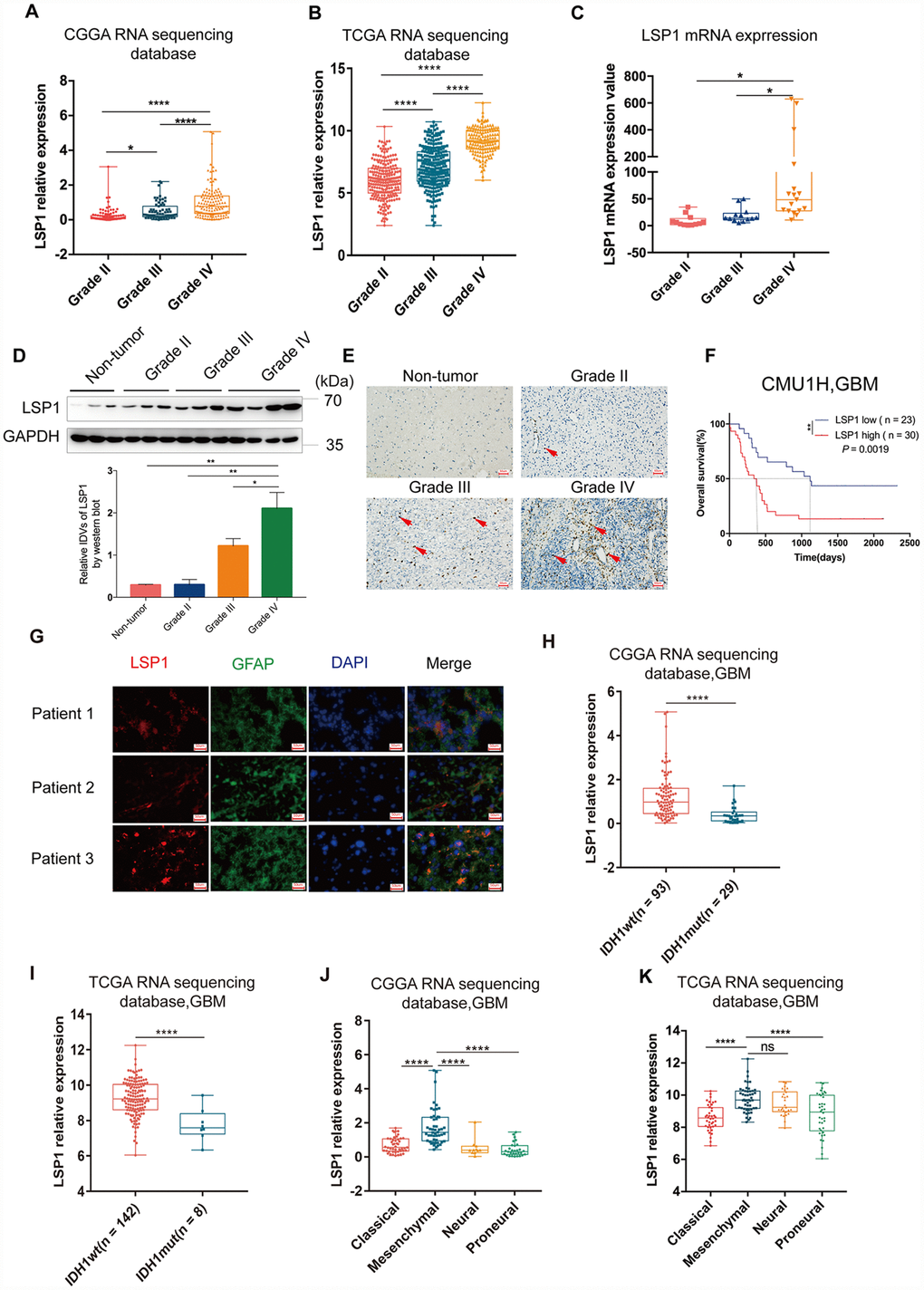
Figure 2.The analyses of LSP1 expression according to WHO grades, GBM subtypes and IDH1 mutant status. (A and B) LSP1 expression was significantly increased in GBM in comparison with WHO grade II and WHO grade III glioma (A, CGGA RNA sequencing dataset, Grade II n = 105; Grade III n = 67; Grade IV n = 138; B, TCGA RNA sequencing dataset, Grade II n = 224; Grade III n = 243; Grade IV n = 155; with one-way ANOVA). (C) qPCR analysis of LSP1 expression in clinical glioma samples (Grade II n = 11; Grade III n = 10; Grade IV n = 21; with one-way ANOVA). (D) Representative western blot images (upper panel) and analyses of LSP1 (lower panel) expressed in clinical tissues (Non-tumor n = 3; Grade II n = 3; Grade III n = 3; Grade IV n = 3; with one-way ANOVA). (E) Representative IHC images of LSP1 staining in clinical non-tumor and glioma samples (200X, scale bar = 50μm). (F) Kaplan-Meier curve evaluating the association of LSP1 expression with the prognosis of GBM patients (LSP1 high vs. low, P = 0.0019; with log-rank test). (G) Representative IF co-staining images of LSP1 and GFAP in clinical GBM samples (200X, scale bar = 50μm). (H and I) LSP1 expression was significantly upregulated in IDH1 wild type GBM (H, CGGA RNA sequencing dataset; I TCGA RNA sequencing dataset; with t test). (J and K) The expression analysis of LSP1 in four subtypes of GBM (J, CGGA RNAseq, Classical n = 47, Mesenchymal n = 50, Neural n = 11, Proneural n = 30; K, TCGA RNAseq, Classical n = 40, Mesenchymal n = 50, Neural n = 27, Proneural n = 38; with one-way ANOVA). ns, *, ** and **** indicate no significance, P < 0.05, P < 0.01, and P < 0.0001, respectively. CGGA, Chinese Glioma Genome Atlas; TCGA, The Cancer Genome Atlas; GBM, glioblastoma multiforme; LSP1, lymphocyte specific protein 1; GFAP, glial fibrillary acidic protein; DAPI, 4’,6-diamidino-2-phenylindole; IDH1, isocitrate dehydrogenase 1.