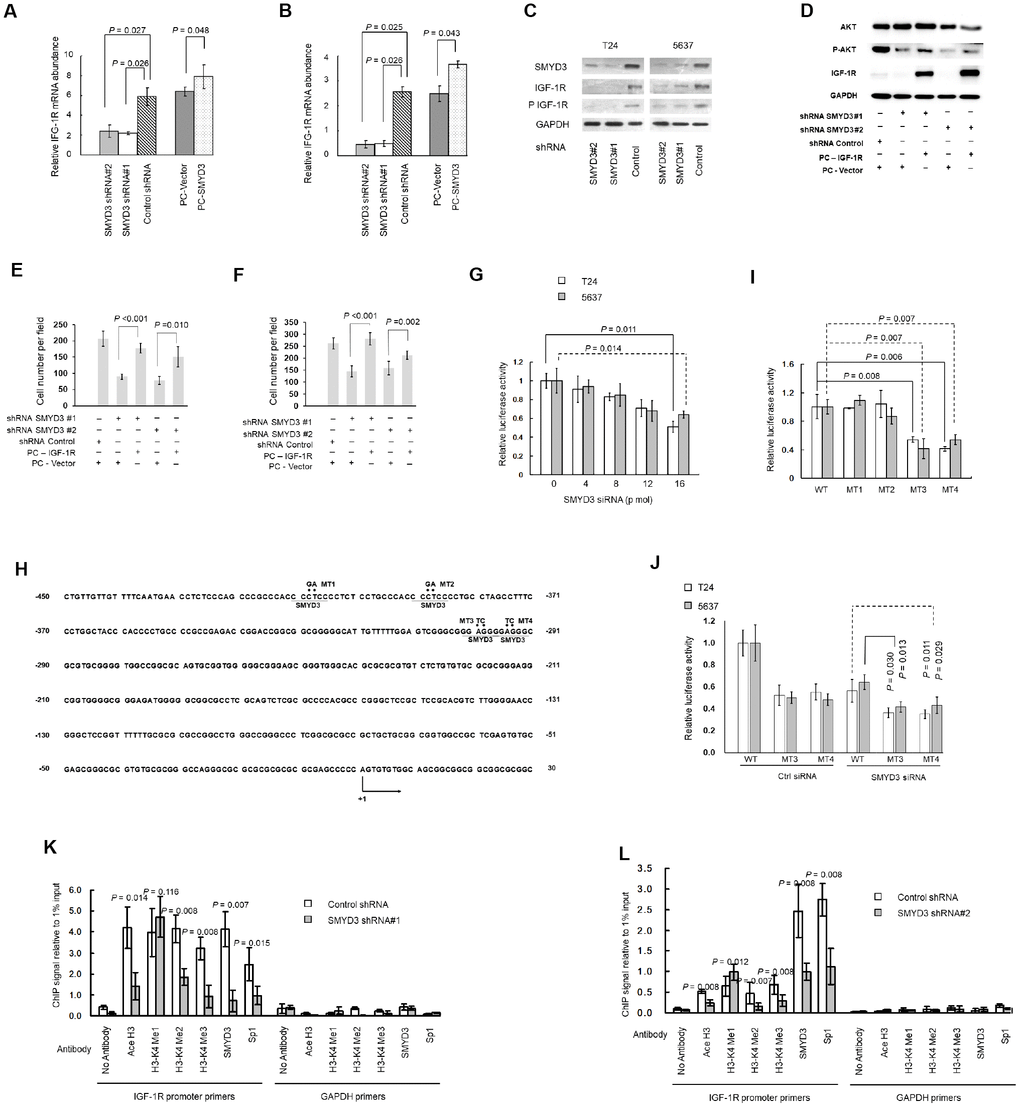
Figure 4.SMYD3 induces IGF-1R transcription through promoter chromatin remodeling. (A, B) RT-PCR of IGF-1R transcription in T24 (A) and 5637 (B) cells transfected with SMYD3 shRNA/ control shRNA or PC-vector/PC-SMYD3. The data were normalized to the mRNA abundance of β-actin. Error bars correspond to standard deviations. Wilcoxon signed-rank tests for the paired samples were used to calculate the two-sided P values based on six independent experiments. (C) Western blot analysis of SMYD3, IGF-1R, and phosphorylated IGF-1R (P-IGF-1R) expression in BC cells, the same transfections and blot (GAPDH and SMYD3) as Figure 2C. (n=3). (D) Western blot analysis of AKT, P-AKT and IGF-1R protein expression in T24 cells 48 h post transfected with indicated vectors. (E) Transwell migration assays of T24 cells 48 h post transfected with indicated vectors, performed as Figure 3E. (F) Transwell invasion assays of T24 cells 48 h post transfected with indicated vectors, performed as Figure 3F. (G) Increasing doses of SMYD3 siRNA were co-transfected with wild-type (WT) pGL3-IGF-1R-LUC plasmid and pRL-TK into T24 or 5637 cells. A luciferase activity assay was performed 48 h after transfection. Six independent transfections were performed. Error bars correspond to standard deviations. A Wilcoxon signed-rank test for the paired samples was used to calculate the two-sided P value. (H) Sequence of the IGF-1R core promoter region. Four potential SMYD3 binding sites are underlined. The sequence that was mutated in the transcriptional activity analysis of cis-acting elements (MT1–MT4) is indicated by dots, and substitutions are given above. The first nucleotide upstream of the transcription start site is indicated by +1; the arrow indicates the first nucleotide of the first exon. (I) WT or SMYD3 motif mutant (MT1–MT4) IGF-1R promoter activity in T24 and 5637 cells. Six independent transfections were performed. Error bars correspond to standard deviations. Wilcoxon signed-rank tests were used to calculate the two-sided P values. (J) SMYD3 siRNA was co-transfected with WT or mutant reporter plasmid (MT3 or MT4) into BC cells. Luciferase activity assay was performed 48 hours after transfection. Three independent experiments were performed in duplicate. Error bars: Standard deviations. Wilcoxon signed-rank tests were used to calculate the two-sided P values. (K, L) Quantitative ChIP assay for H3-K4 tri/di/monomethylation, H3 acetylation, and Sp1 and SMYD3 occupancy at the IGF-1Rpromoter in T24 (K) and 5637 (L) cells expressing SMYD3 shRNA or control shRNA. Omission of antibodies (No Antibody) was included throughout the entire experimental procedure, in addition to PCR amplification of the unrelated GAPDH gene, as an appropriate control. The data shown are from three independent experiments in triplicate. Mean values of ChIP signals are normalized to 1% input. Input control was from non-immunoprecipitated total chromatin DNA. Error bars correspond to standard deviations. Wilcoxon signed-rank tests were performed to calculate the two-sided P values. Ace H3, acetylated Histone 3; H3-K4 Me1, monomethylated H3-K4; H3-K4 Me2, dimethylated H3-K4; H3-K4 Me3, trimethylated H3-K4.