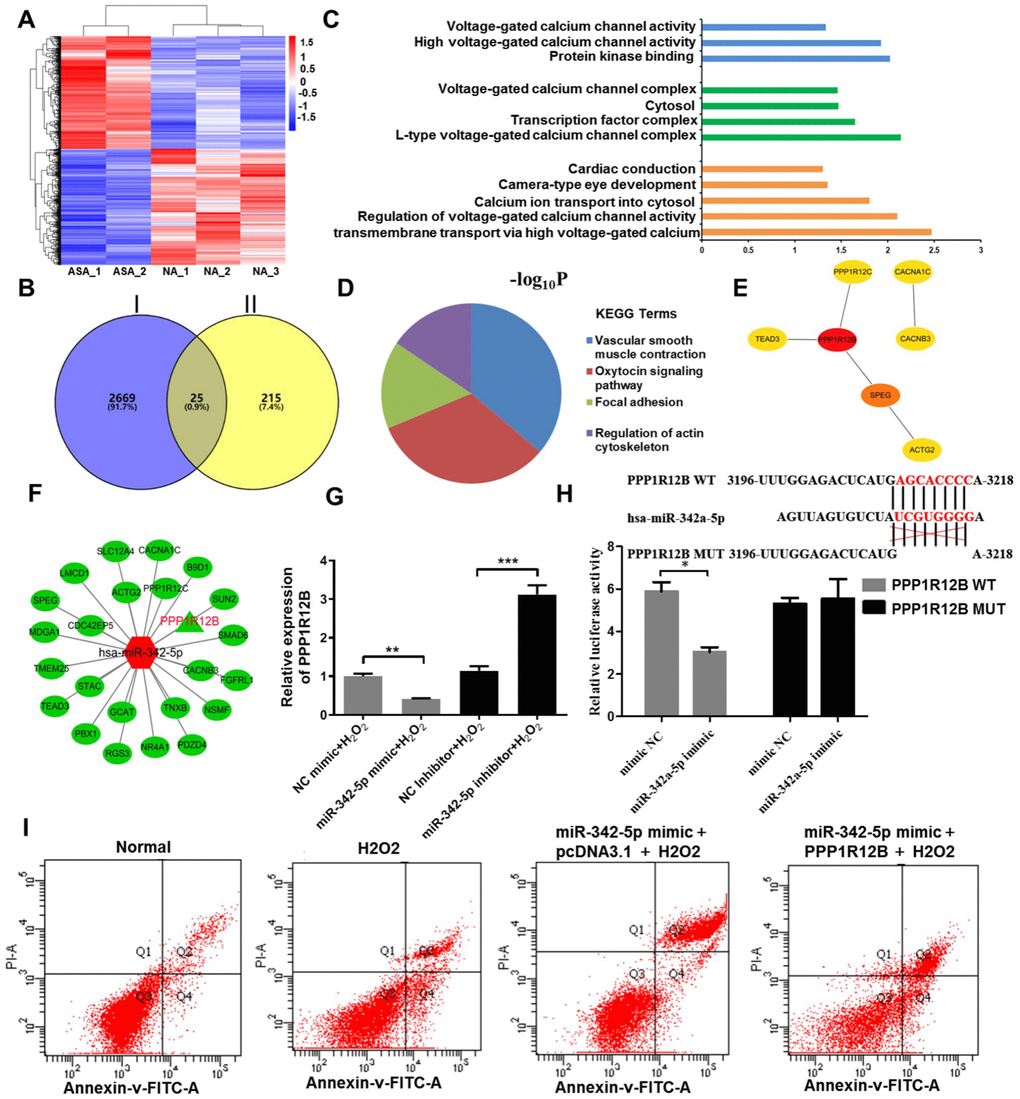
Figure 4.PPP1R12B was identified as a target of miR-342-5p according to bioinformatics analyses and test verification. (A) R cluster analysis of the differentially expressed mRNAs obtained from RNA-sequencing analysis. The red presents upregulated mRNAs and the blue indicates downregulated mRNAs. (B) The downregulated mRNAs in RNA-sequencing and the common genes from three databases (Miranda, PITA and RNAhybrid) were overlapped to obtain twenty-five genes. Circle I stands for the downregulated mRNAs in RNA-sequencing. Circle II represents the common genes obtained from Miranda, PITA and RNAhybrid databases. (C) GO terms and (D) KEGG signaling pathway enrichment analysis of twenty-five overlapped genes. (E) The hub genes extracted from twenty-five overlapped genes by Hubba plug-in according to degree score. (F) The network interaction between miR-342-5p and twenty-five overlapped genes was visualized by Cytoscape. The red shape node represents upregulated in atherosclerosis and the green shape node suggests downregulated in atherosclerosis. PPP1R12B as aimed mRNA is highlighted in red. (G) The effect of miR-342-5p on PPP1R12B expression in HUVECs’ lesion model was detected by qRT-PCR analysis. **, P<0.01, ***, P<0.001. (H) The target gene of miR-342a-5p, PPP1R12B, was identified by dual luciferase reporter gene assay. *, P<0.05. (I) HUVECs were co-transfected with miR-342-5p mimic and PPP1R12B overexpression vector, while pcDNA 3.1 was set as the control group, followed by treating with 1500 uM of H2O2. Their apoptotic rates were then analyzed by flow cytometry assay. miR-342-5p, microRNA-342-5p; mRNA, messenger RNA; GO, Gene ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; HUVECs, human umbilical vein endothelial cells; qRT-PCR, quantitative real-time polymerase chain reaction; NC, negative control.