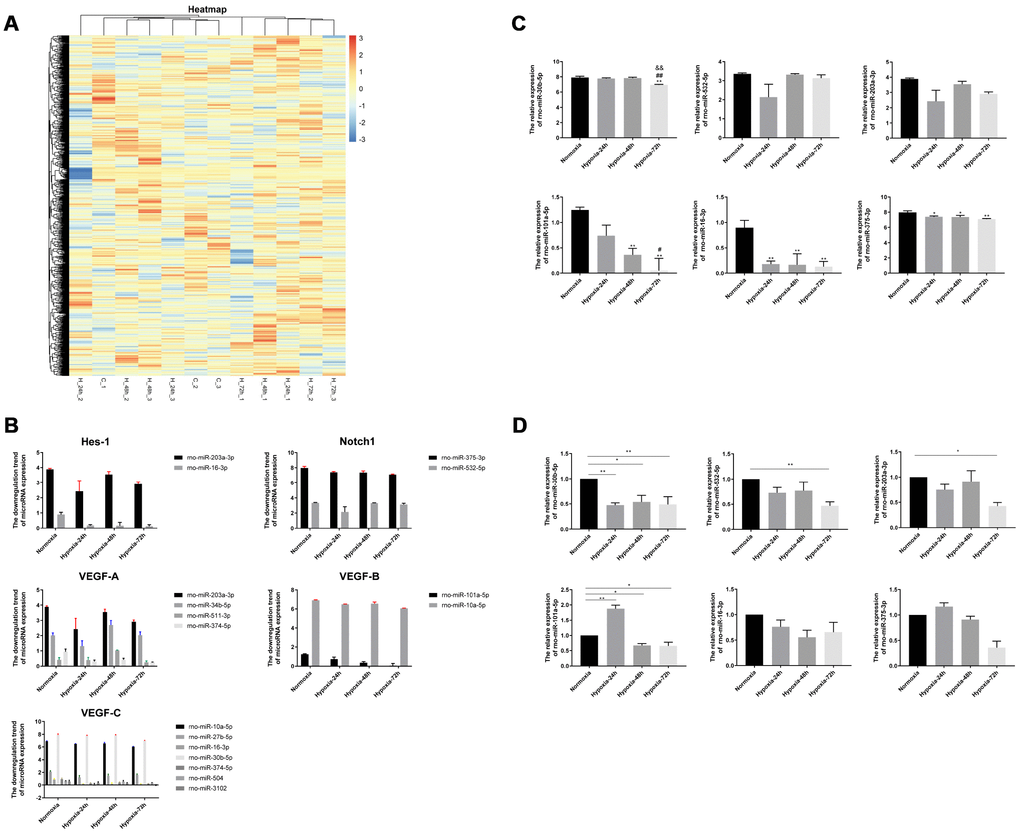
Figure 3.Screening and validation of miRNAs associated with hypoxic exposure. (A) Heatmap of miRNA microarray data. Unmonitored hierarchical clustering analysis was conducted for differentially expressed genes induced by hypoxia (24,48 and 72h) in the rat lung. A total of 57 miRNAs showed > 1.5 fold change relative to normoxic, control lung tissue samples; blue indicates downregulation. (B) Downregulation of miRNAs targeting Hes-1, Notch1, VEGF-A, VEGF-B, and VEGF-C by hypoxia. (C) Screening of 6 selected differentially expressed miRNAs associated with the VEGF/Notch pathway. (D) Verification of selected miRNAs expression in rat lung by qRT-PCR. Values are expressed as fold change ± SEM relative to control. *P < 0.05, **P < 0.01 compared with the normoxic control group; #P <0.05, ##P < 0.01 compared with the 24-h hypoxia group; &&P < 0.01 compared with the 48-h hypoxia group.