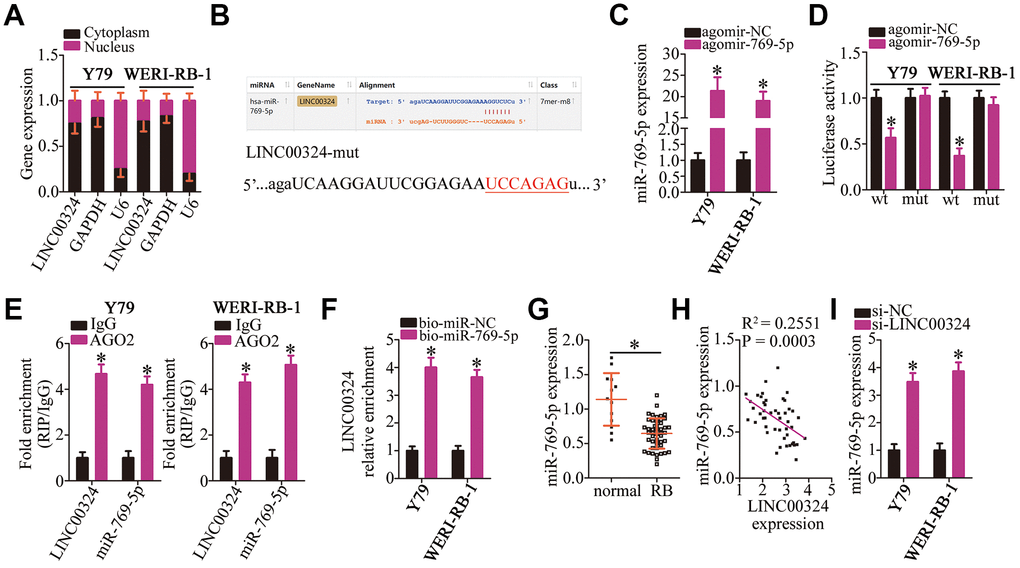
Figure 3.LINC00324 acts as a sponge on miR-769-5p in RB cells. (A) RNA was extracted from cytoplasmic and nuclear fractions, and then subjected to RT-qPCR to characterize the distribution of LINC00324 inside Y79 and WERI-RB-1 cells. (B) The miR-769-5p–binding sequences in LINC00324 were predicted using starBase 3.0. The designed mutant binding site is also shown. (C) Either agomir-769-5p or agomir-NC was transfected into Y79 and WERI-RB-1 cells. RT-qPCR was conducted at 48 h post-transfection to quantitate miR-769-5p expression. *P < 0.05 vs. group agomir-NC. (D) Luciferase activity was measured using the luciferase reporter assay in Y79 and WERI-RB-1 cells cotransfected with either LINC00324-wt or LINC00324-mut and either agomir-769-5p or agomir-NC. *P < 0.05 vs. the agomir-NC group. (E) The interaction between LINC00324 and miR-769-5p in Y79 and WERI-RB-1 cells was detected via the RIP assay. LINC00324 and miR-769-5p expression was measured by RT-qPCR. *P < 0.05 vs. the IgG group. (F) Y79 and WERI-RB-1 cells were transfected with bio-miR-769-5p or bio-miR-NC and their lysates were incubated with streptavidin-coupled beads to form bio-miRNA-lncRNA complexes. The LINC00324 enrichment was analyzed by means of RT-qPCR analysis. *P < 0.05 vs. the bio-miR-NC group. (G) The relative expression of miR-769-5p in 47 RB tissue samples and 13 normal retinal tissue samples was determined using RT-qPCR, and was normalized to that of U6. *P < 0.05 vs. normal retinal tissue samples. (H) The correlation between the expression of LINC00324 and miR-769-5p was investigated by Spearman’s correlation analysis; R2 = 0.2551, P = 0.0003. (I) After transfection with either si-LINC00324 or si-NC, the expression of miR-769-5p in Y79 and WERI-RB-1 cells was determined via RT-qPCR. *P < 0.05 vs. the si-NC group.