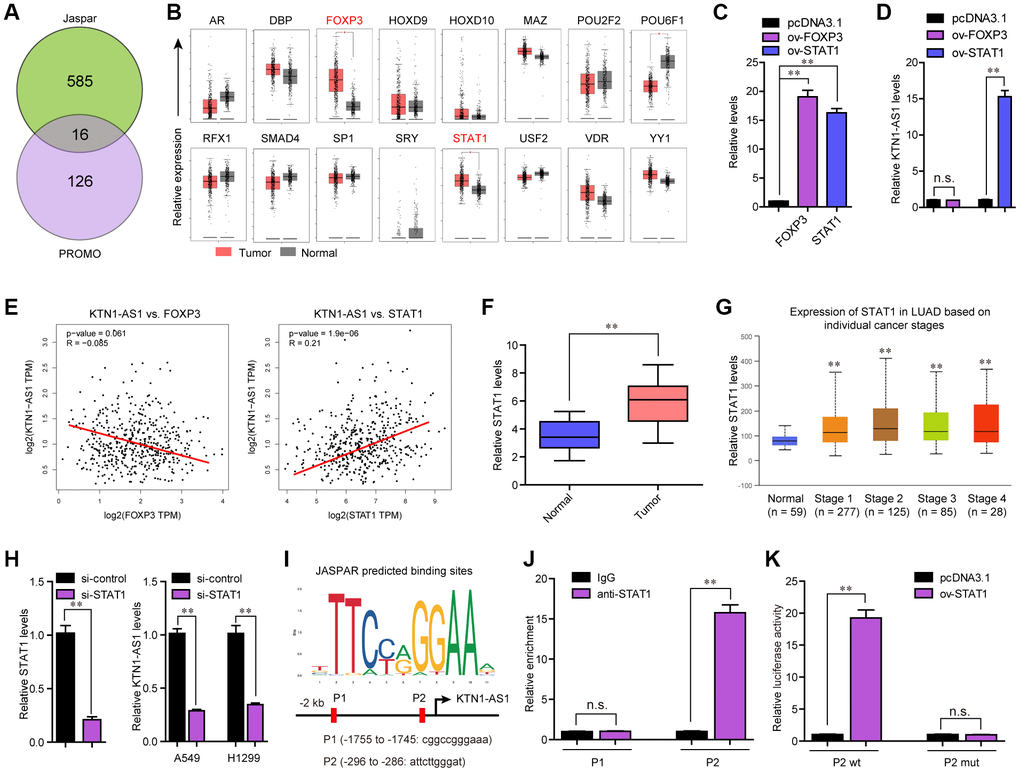
Figure 2.STAT1 stimulate KTN1-AS1 dysregulation in NSCLC. (A) Potential TFs were predicted by Jaspar and PROMO. The results of Jaspar and PROMO were intersected. (B) Expression of 16 TFs were analyzed by GEPIA using TCGA data. (C and D) qPCR detection for the determination of the association between overexpression of STAT1 and KTN1-AS1 expression. (E) The expressing correlation between KTN1-AS1 expression and FOXP3 and STAT1 expression was analyzed by GEPIA. (F) qPCR detected STAT1 expression in 127 NSCLC samples. (G) STAT1 expression across stages were analyzed by “UALCAN”. (H) KTN1-AS1 expression in A549 and H1299 cells after knockdown of STAT1 was determined by qRT-PCR. (I) Jaspar predicting potential binding sites between STAT1 and KTN1-AS1 promoter. (J) ChIP assay was performed to determine the affinity of STAT1 to KTN1-AS1 promoter. (K) Luciferase activity assays were applied to further confirm the binding of STAT1 to KTN1-AS1 promoter. * P < 0.05, **P < 0.01.