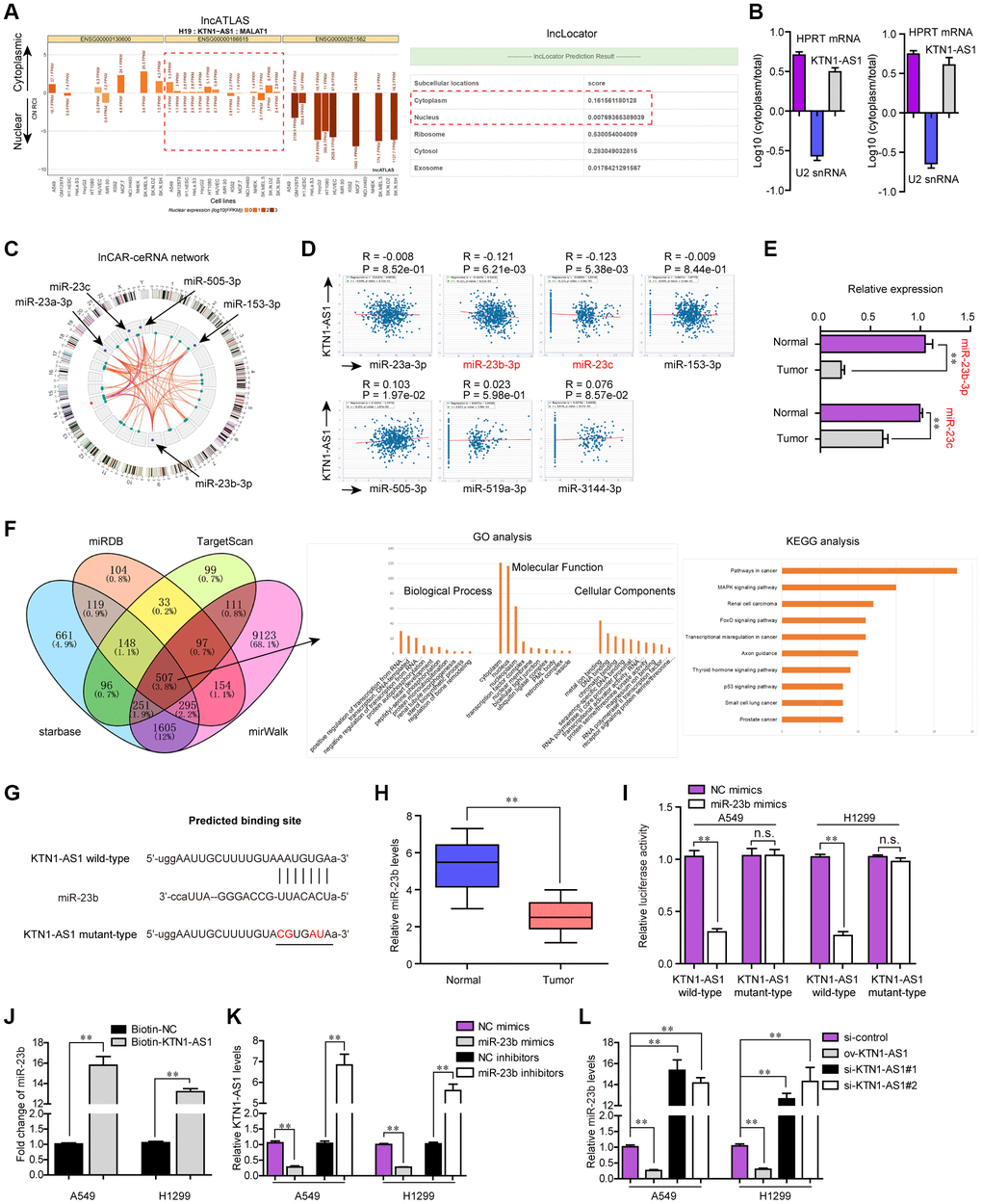
Figure 6.miR-23b was directly targeted by KTN1-AS1. (A) The subcellular localization of KTN1-AS1 was predicted by lncATLAS and lncLocator. (B) Subcellular fractionation assays. (C) The ceRNA network of KTN1-AS1 was analyzed by lnCAR. (D) starBase program analyzed the expressing correlation between KTN1-AS1 and miRNAs. (E) qPCR analysis detected the expression of miR-23b and miR-23c. (F) The intersection of the results from miRDB, TargetScan, starBase and miRWalk prediction. The commonly predicting genes were also used fo GO and KEGG analysis. (G) The predicting binding site between KTN1-AS1 and miR-23b. (H) qPCR assessed the miR-23b levels in 127 NSCLC tissues. (I) Relative luciferase activity detection. (J) RNA-pull down. (K, L) qPCR analysis detected the levels of KTN1-AS1 and miR-23b, respectively. * P < 0.05, **P < 0.01.