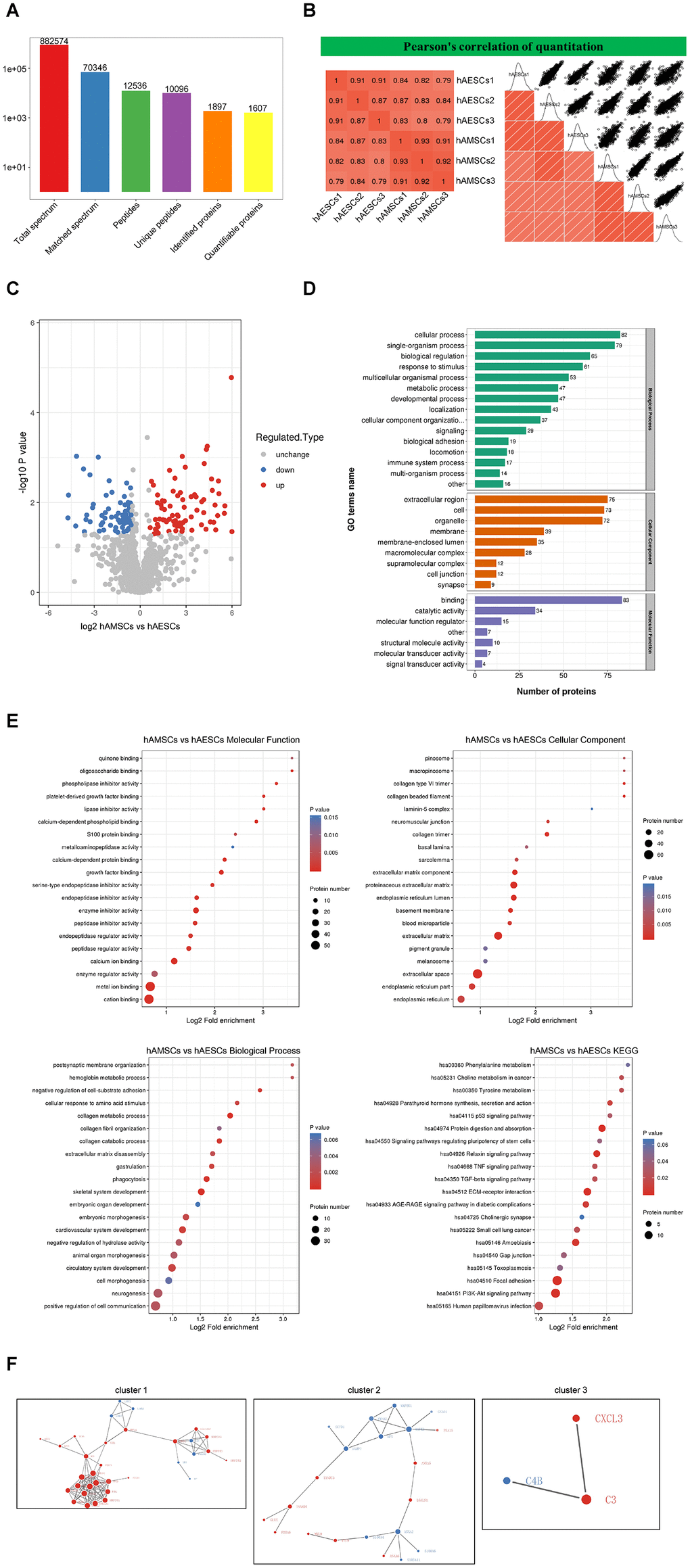
Figure 5.Mass spectrometry and bioinformatic analyses of proteins in hAMSCs-CM (condition media of hAMSCs) and hAECs-CM (condition media of hAECs). (A) Basic summary of mass spectrometric data. (B) A heat map compares Pearson correlation coefficients of proteins in hAMSCs-CM and hAECs-CM samples. A value closer to -1 indicates negative correlation, whereas a value closer to +1 indicates positive correlation. A value of 0 denotes no correlation. (C) Quantitative volcano map of differentially expressed proteins in conditioned media. The horizontal axis shows the Log2 relative expression value of the proteins, whereas, the vertical axis denotes the logarithmic p-value. The red dots indicate significantly up-regulated proteins and the blue dots indicate significantly down-regulated proteins. (D) Top differentially expressed proteins in the conditioned media classified according to three GO domains: biological process, molecular function and cellular component. (E) A directed acyclic graph shows GO and KEGG enrichment analyses of differentially expressed proteins in conditioned media. The circles indicate the GO classification of the differential expressed protein; the red color indicates highly significant protein (P<0.01), the yellow color indicates significant protein (P<0.05), and the blue color indicates no significance. The line with an arrow indicates the upper and lower levels of relationship according to the GO classification, and the circle size denotes the degree of enrichment. (F) The protein-protein interaction network map of the differentially expressed proteins in the conditioned media. The circles indicate differentially expressed proteins. The blue color denotes down-regulated proteins and the red color denotes up-regulated proteins. The size of the circle shows the number of proteins interacting with each other. A larger circle denotes a protein with several interacting partners.