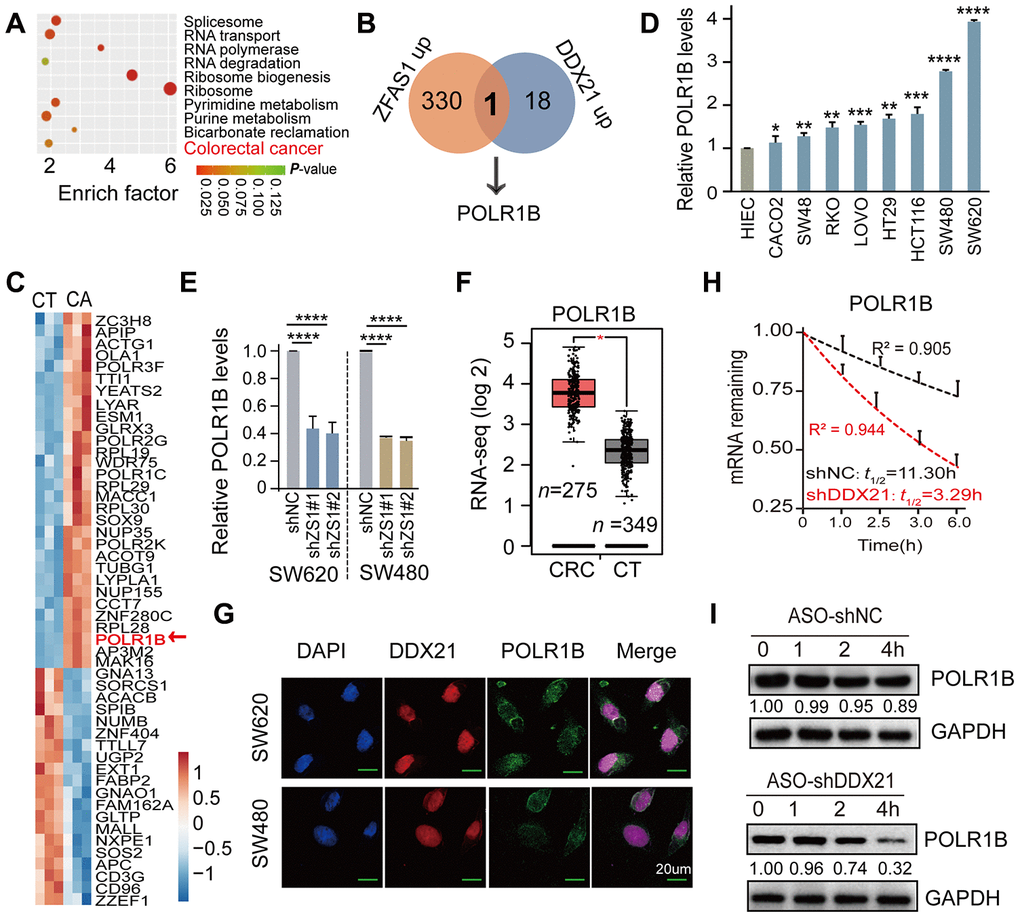
Figure 5.POLR1B is a critical target of lncRNA ZFAS1 interacting with DDX21. (A) KEGG and GO analysis enriched the co-expression target genes and the top 10 cellular function components affected by lncRNA ZFAS1 and DDX21. (B) The intersection of co-expression downstream genes between lncRNA-ZFAS1 up-regulation and DDX21 up-regulation. (C) The hierarchical clustering heat map showing the DDX21 related target genes in CRC patient tissues and their matched paired adjacent-tumor samples (n = 3). Red in heat map denotes up-regulation. Blue denotes down-regulation. (D) The expression of POLR1B in CRC cells including SW480, SW620, HCT116, SW48, CACO2, LOVO, HT29, and RKO cells and in normal intestinal epithelial HIEC cell assayed by the qPCR method. GAPDH was selected as an internal control. (E) The mRNA expression of POLR1B and POLR1A after lncRNA ZFAS1 knockdown in SW620 and SW480 cells by qPCR assay. (F) RNA-seq data showing the log 2 gene expression of POLR1B in CRC patients tissues (n = 275) and the normal controls (n = 375) based on the TCGA dataset (http://gepia.cancer-pku.cn/). (G) Co-localization of POLR1B protein and DDX21 protein detected by IF assays in SW620 and SW480 cells. Scale bar = 20μm. (H) RNA stability assay detected the POLR1B mRNA decay after knockdown DDX21 expression. (I) The protein expression levels of POLR1B were measured by translation inhibition assays treated with CHX. Data were shown as mean ± s.d.. Two-tailed Student’s t-tests were used. *P <0.05; **P <0.01; ***P <0.001; ****P <0.0001.