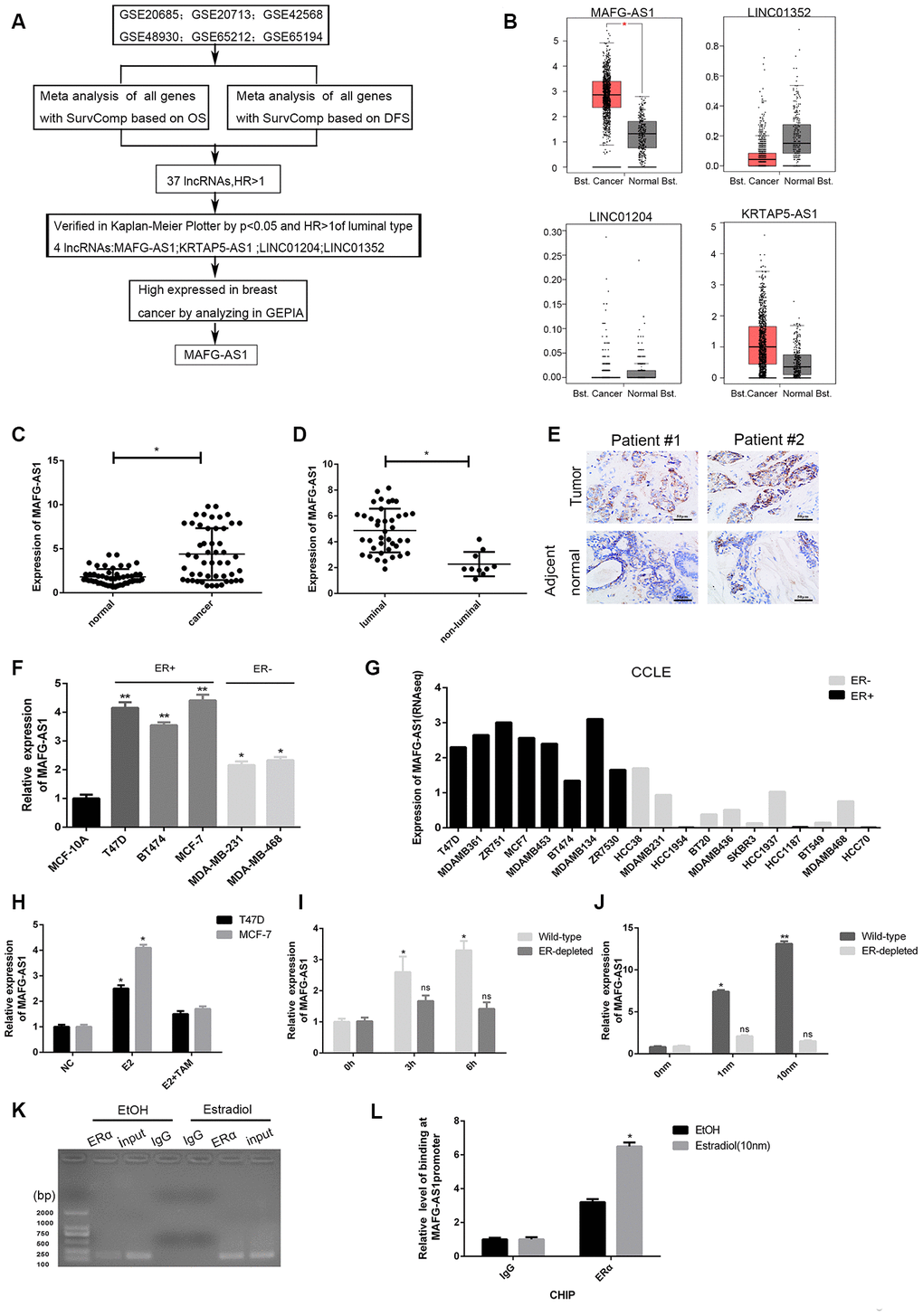
Figure 1.Identification of lncRNAs associated with poor survival and related to ER+ breast cancer. (A) The flow chart of identifying target lncRNA using bioinformatics methods. (B) The expression level plots of four lncRNAs in cancer and normal tissues of breast cancer recorded in GEPIAE (http://gepia.cancer-pku.cn/index.html). Only the expression of MAFG-AS1 in breast cancer (n=1085) and adjacent normal tissues (n=291) is significantly different (p<0.05). (C) Expression of MAFG-AS1 in 50 breast tumors compared with para-carcinoma normal tissues. (D) Expression of MAFG-AS1 was higher in luminal breast cancer (n=40) than in non-luminal breast cancer (n=10)(*p<0.05). (E) Representative ISH (in situ hybridization) detection images of MAFG-AS1 expression in matched normal and primary tumors from two ER positive patients are shown. Scale bars, 50um. (F) Relative MAFG-AS1 expression in 5 breast cancer cell lines and 1 normal breast cell line. Error bars represent mean ±SD for triplicate experiments, *p <0.05,**p<0.01. (G) MAFG-AS1 was specifically highly expressed in ER+ breast cancer cell lines, as determined by analyzing the RNA-seq data from CCLE (Cancer Cell Line Encyclopedia). (H) qPCR expression of MAFG-AS1 8 h following addition of DMSO vehicle, 10 nM estrogen with or without tamoxifen (1uM) in MCF-7 and T47D cell lines. (I) The expression of MAFG-AS1 in Wild-type and ERα-depleted T47D cells by different time same concentration. (J) qRT-PCR analysis of the expression of MAFG-AS1 in Wild-type and ERα-depleted T47D cells by same time different concentrations. (K–L) Gel imaging and qPCR-based ChIP analysis of the MAFG-AS1 promoter following ChIP for ERα following 12hr estradiol or DMSO vehicle stimulation. Expression normalized to IgG pulldown. Error bars represent mean±SD.