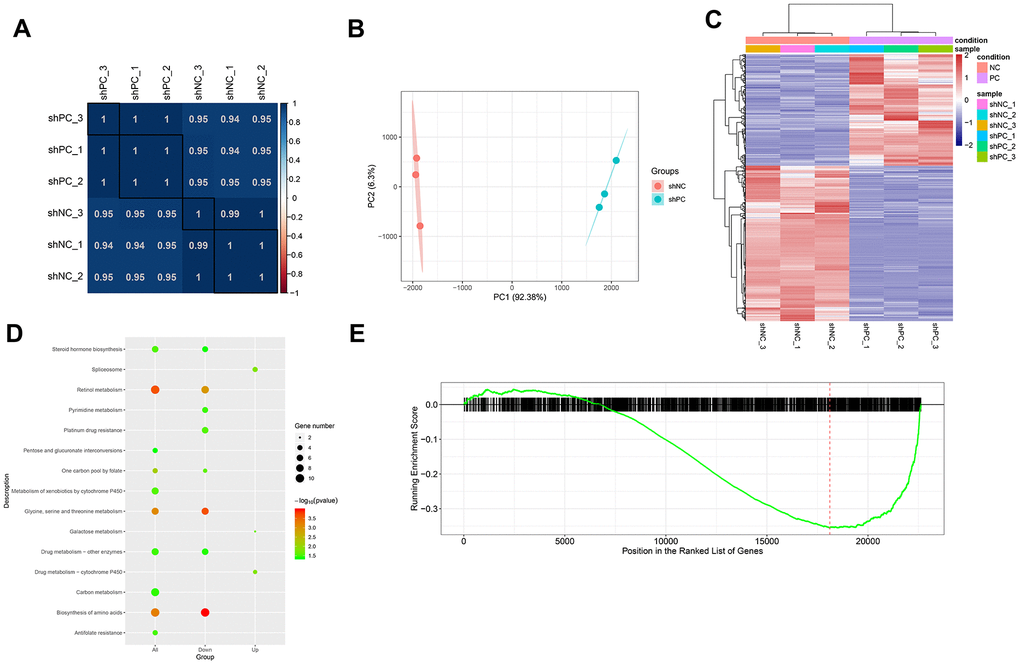
Figure 4.Differentially expressed genes and pathways enrichment analysis after PC knockdown based on transcriptome sequencing. (A) Sample correlation based on differential gene expression. The correlation between samples was analyzed using Pearson’s correlation coefficient based on gene expression values. There were significant positive correlations between samples. (B) Principal component analysis results. The different colored dots represent the sample group under the condition. (C) Heatmap of differentially expressed genes between shPC and shNC samples. Two-dimension clustering analysis results were visualized using heatmaps for differentially expressed genes from PC-knockdown samples compared to the normal control group. The gene expression profiles were significantly different between groups. Red represents high expression levels while blue represents low expression levels. (D) KEGG pathway enrichment analysis for differentially expressed genes. The top ten pathway terms ranked by p-value were visualized using dot plot. The vertical axis represents KEGG pathways and the horizontal axis shows differentially expressed genes. A category with a smaller p-value represents a more significant difference. (E) Gene set enrichment analysis results. The red line refers to the highest enrichment score.