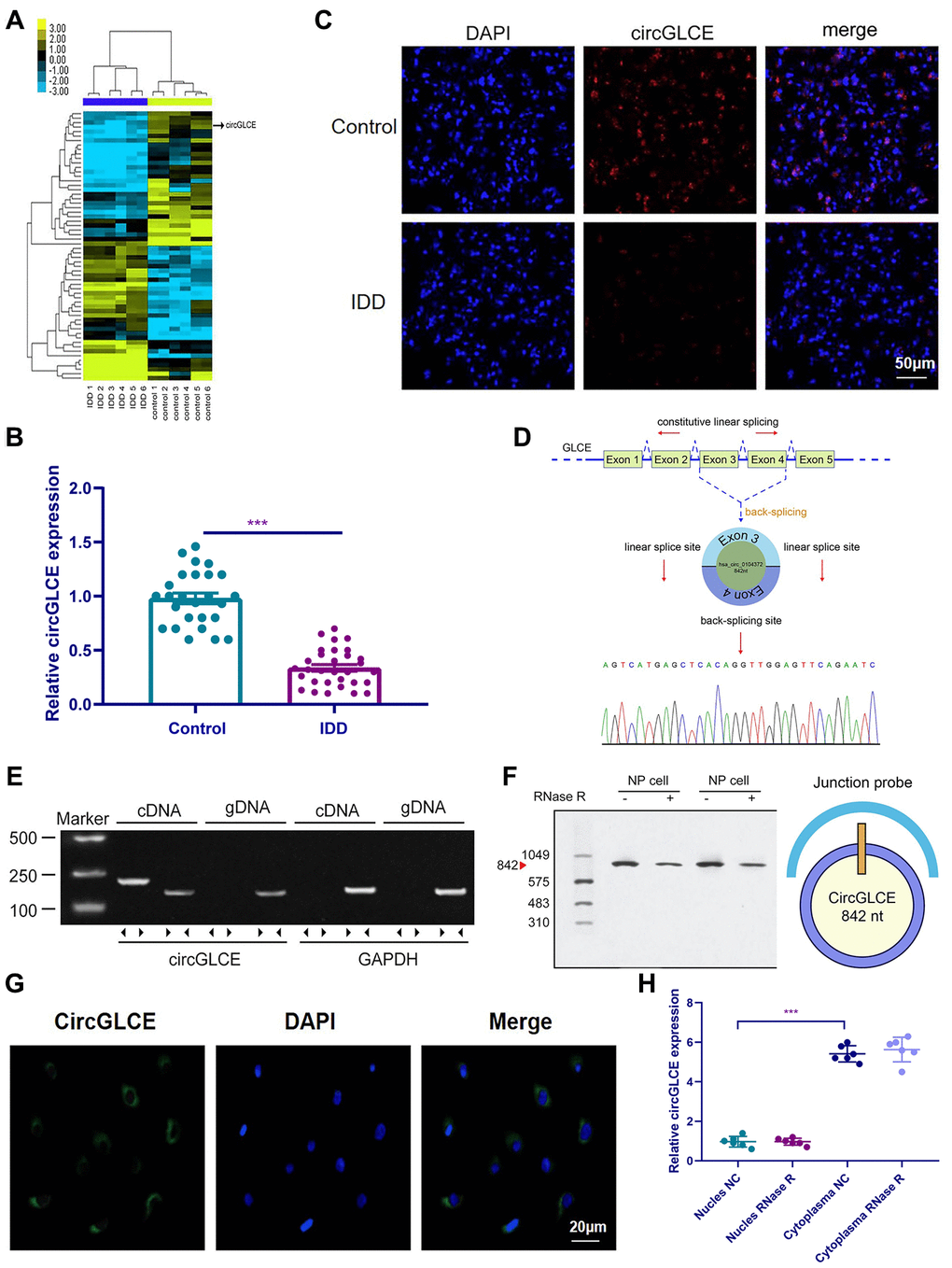
Figure 1.CircGLCE expression in IDD and normal NP tissue samples. (A) Heat map of all differentially expressed circRNAs between the IDD sample and the paired controls. (B) The comparison of CircGLCE expression between 31 degenerative NP samples and 26 controls, which was quantified with RT-qPCR (***P < 0.001, unpaired two-tailed Student’s t test). (C) The CircGLCE level was measured by FISH, and it also showed a significant decrease in the levels in degenerative NP samples compared to controls. (D) Schematic illustration showing the circularization of GLCE exons 3-4 from circular RNA. The Sanger sequencing of CircGLCE relied on the validation of content by RT-qPCR. (E) CircGLCE was amplified by divergent primers with cDNA. (F) CircGLCE showed RNase R resistance in northern blotting. The length of CircGLCE was 842 nt. The probe targeted the junction. (G) The FISH results demonstrated the presence of CircGLCE in the cytoplasm of NP cells. (H) The results of RT-qPCR confirmed the presence of CircGLCE in the cytoplasm of NP cells (n=6, *** P<0.001, unpaired two-tailed Student’s t test). Data are the mean ± SEM.