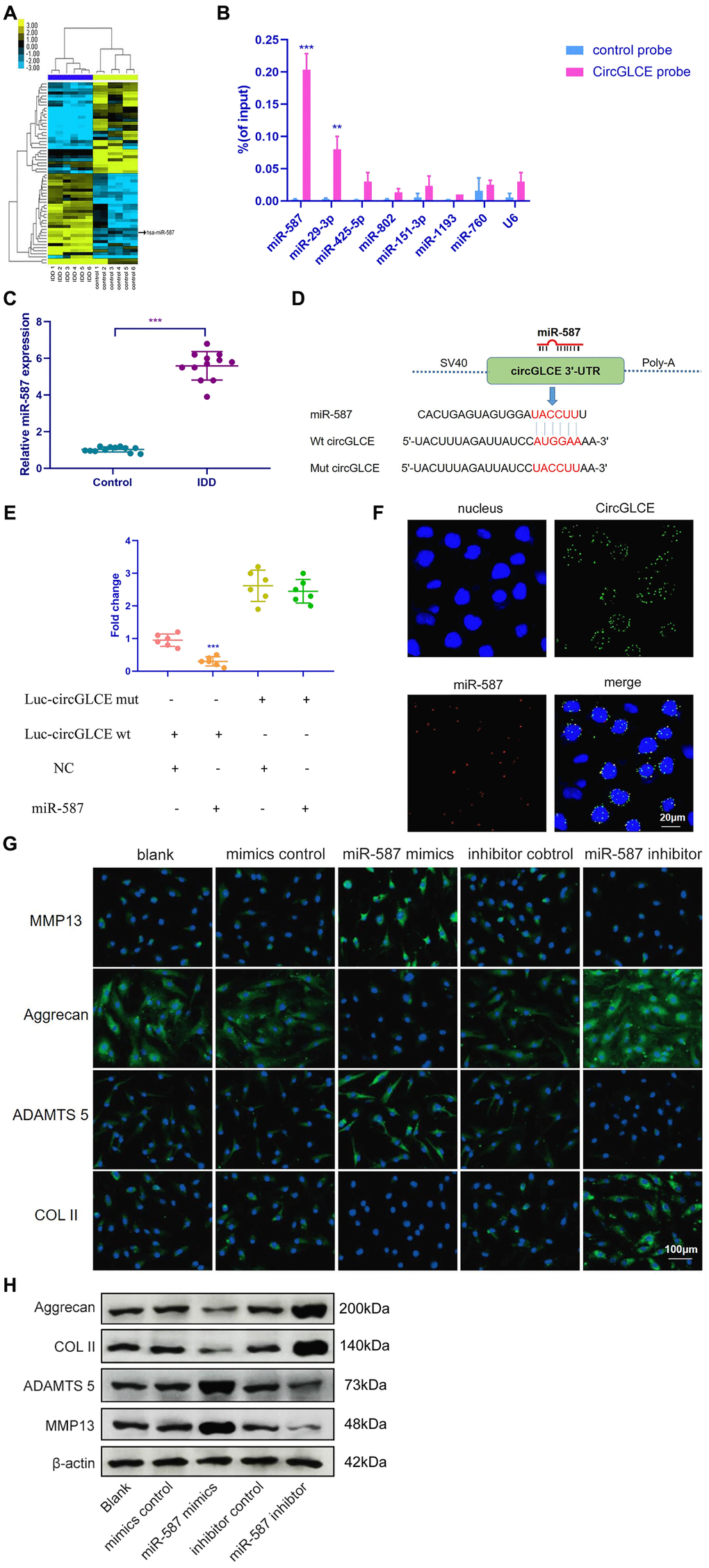
Figure 3.CircGLCE serves as a miR-587 sponge in NP cells. (A) Differentially expressed miRNAs in degenerative NP samples were identified with high-throughput sequencing. (B) Following RNA pull-down experiments performed with a CircGLCE probe, RT-qPCR indicated significantly high miR-587 levels. The relative levels of CircGLCE were normalized to the input levels (***P < 0.001, one-way ANOVA coupled with Tukey’s post hoc test). (C) RT-qPCR demonstrated that miR-587 was upregulated in IDD (n=12, **P < 0.01, and ***P < 0.001; unpaired two-tailed Student’s t test). (D) The binding sites in CircGLCE for miR-587 were analysed by a bioinformatic approach. (E) NP cells were co-transfected with miR-587 mimics/negative controls and luciferase reporter constructs for wild-type or mutant CircGLCE. Dual luciferase assays demonstrated binding between miR-587 and CircGLCE (n=6, *** P<0.001, unpaired two-tailed Student’s t test). (F) FISH showed the colocalization of CircGLCE and miR-587 in NP cells. The miR-587 probe was labelled with Alexa Fluor 488, the CircGLCE probe was tagged with Cy3, and the nuclei were stained with DAPI. (G) The ECM enzymes were effectively regulated by overexpressing or knocking down miR-587. *** P<0.001. (H) Western blot analysis (n=3) confirmed the results in (G). Data are the mean ± SEM.