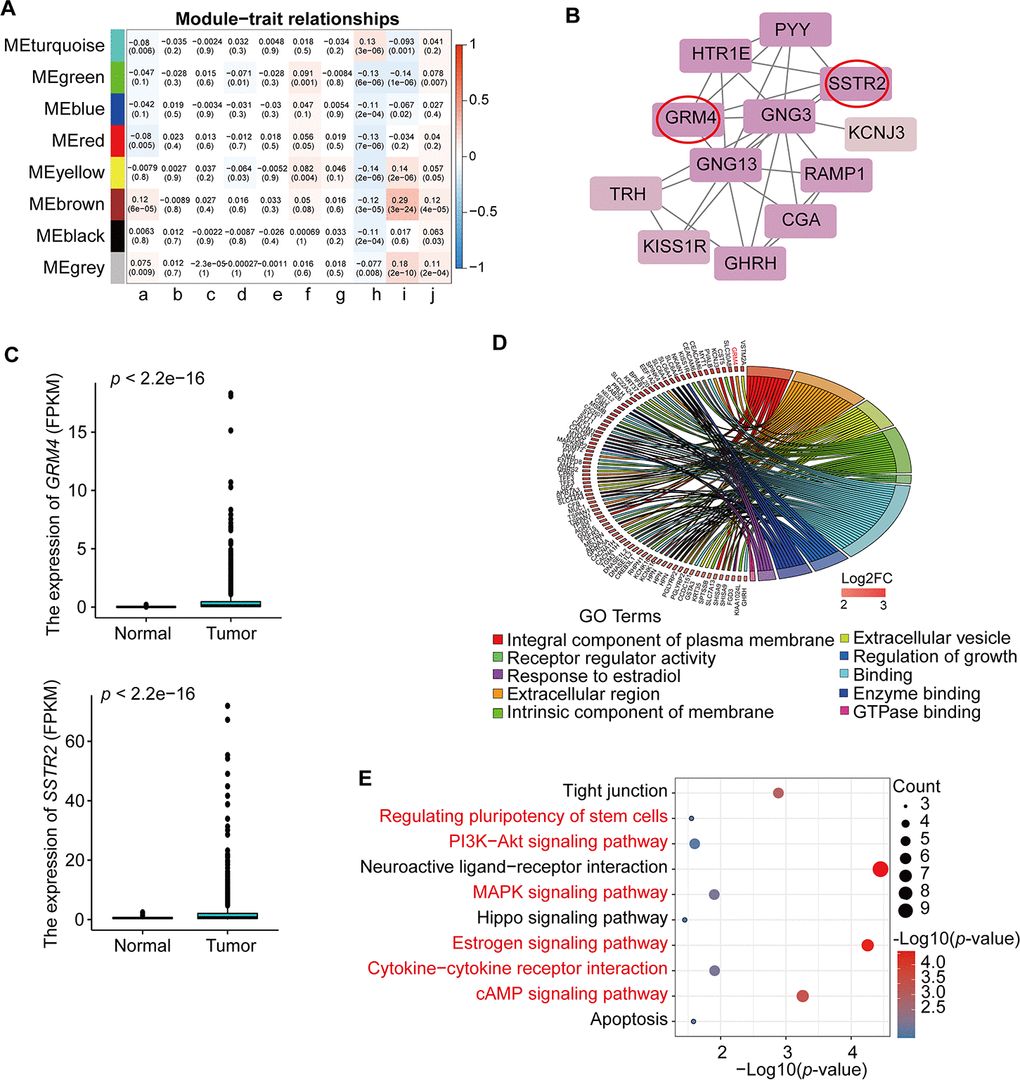
Figure 3.Identification and analysis of key module and hub genes. (A) Analysis of module-trait relationships of BRCA based on TCGA data; a. age at initial pathologic diagnosis, b. pathologic_M, c. pathologic_N, d. pathologic_T, e. tumor stage I, f. additional pharmaceutical therapy, g. radiation therapy, h. vital status, i. days to new tumor event after initial treatment, j. days to death. TNM = tumor, node, metastasis (classification). (B) PPI analysis and identification of hub genes involved in the co-expression Brown module using STRING database and MCODE plug-in in Cytoscape. The genes in the red circle are the hub genes. (C) Expression of GRM4 and SSTR2 in BRCA from TCGA database. (D) GO enrichment in the co-expression Brown module. The red gene is the hub gene of PPI. (E) KEGG pathway enrichment in the co-expression Brown module. Red pathways are common with total DEmRNAs.