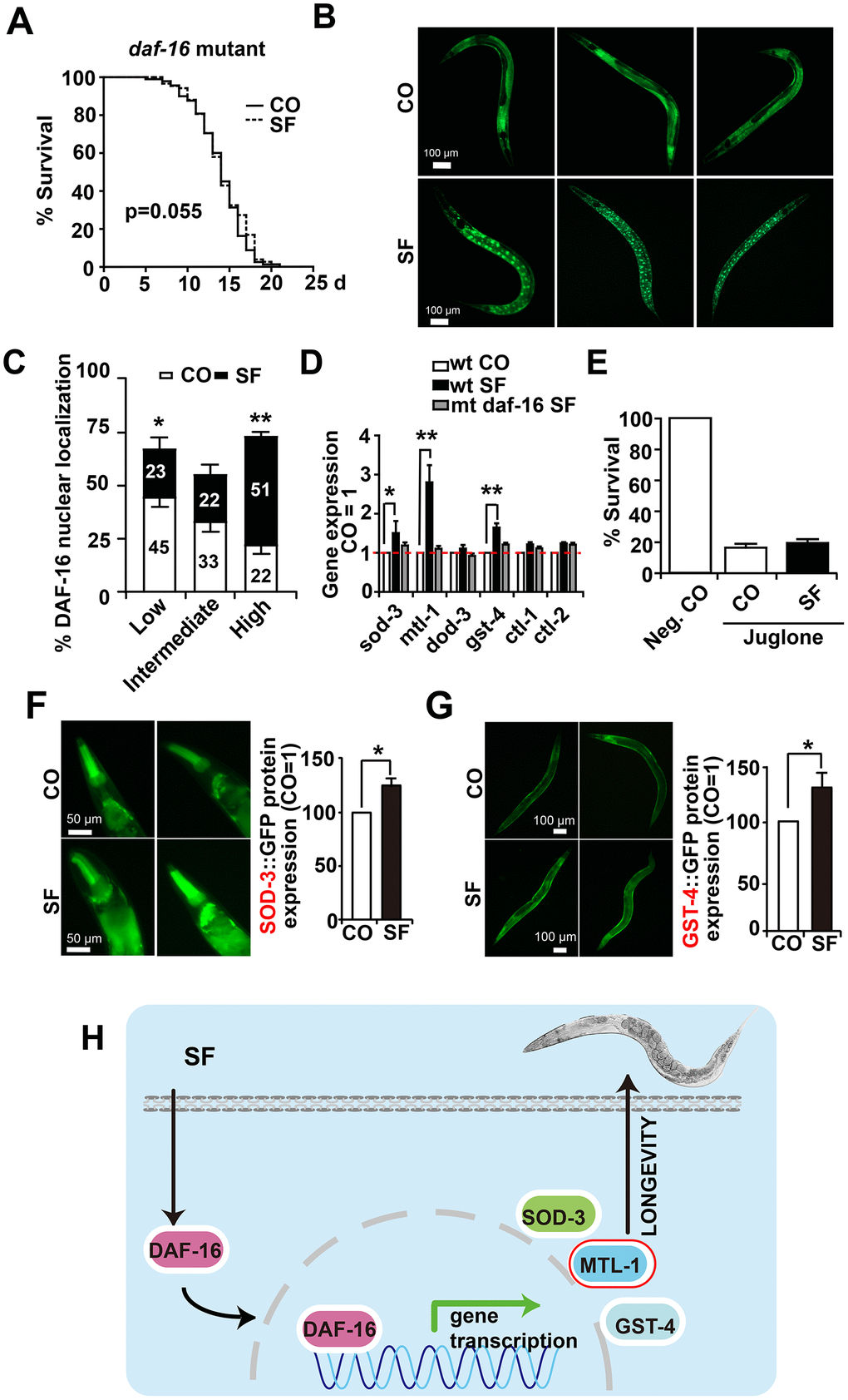
Figure 6.DAF-16 mediates sulforaphane-induced longevity and stress resistance. (A) Approximately 100 L4 larvae from the C. elegans CF1038 daf-16(mu86)I strain, carrying a mutant daf-16 gene (daf-16 mutant), were fed in the presence or absence (CO) of 100 μM sulforaphane (SF), and a Kaplan-Meier survival curve was generated as described in Figure. 1 and Materials and Methods. P=0.055, according to a log-rank test. (B) Adult TJ356 worms with a daf-16-GFP fusion gene were treated with sulforaphane for 3 days (SF) or were left untreated (CO), followed by the detection of green fluorescence in 20 worms per group by fluorescence microscopy. The bright green fluorescent dots indicate the nuclear localization of DAF-16. The scale bar indicates 100 μm. (C) The number of spots was counted and is represented in the diagram. The following scoring of the fluorescence intensity was applied: low (number of green fluorescent spots <50%), intermediate (number of green fluorescent spots ≈50%), and high (number of green fluorescent spots >50%). (D) Approximately 200 synchronized C. elegans L4 wild-type larvae received a regular diet (wt CO) or were fed in the presence of sulforaphane (wt SF). Additionally, L4 larvae of the daf-16(mu86)I strain, carrying a daf-16 mutation (mt daf-16), were fed in the presence of sulforaphane (mt daf-16 SF). Three days later, living worms were selected, and total RNA was extracted. A qRT-PCR assay with specific C. elegans primers (Table 3) was performed to detect the expression of the DAF-16-target genes sod-3, mtl-1, dod-3, gst-4, ctl-1, and ctl-2. The expression levels in untreated control worms were set as 1. (E) Approximately 100 synchronized L4 larvae of the daf-16(mu86)I C. elegans strain were fed OP50 bacteria in the absence (Neg. CO) or presence of sulforaphane (SF), in the presence or absence of a 120 μM concentration of the oxidative chemical juglone, as indicated. Fourteen days later, the surviving worms of each group were selected and counted. (F) L4 CF1553 worms with a SOD-3::GFP fusion gene were fed with sulforaphane for 3 days (SF) or were left untreated (CO), followed by the detection of green fluorescence in 20 worms per group by fluorescence microscopy, indicating a high expression of SOD-3 in the head and foregut of C. elegans. The fluorescence intensity was analyzed by imageJ, and the fluorescence of the control was set to 1. Representative images at 100× magnification are shown on the left, and the scale bar indicates 50 μm. (G) L4 CL2166 worms with a GST-4::GFP fusion gene were treated and analyzed as described above. Representative images at 100× magnification are shown on the left, and the scale bar indicates 100 μm. *P<0.05. (H) Cartoon to demonstrate that sulforaphane mediates the nuclear translocation of the DAF-16 transcription factor, where it binds to the promoter regions of its target genes and thereby activates SOD-3, MTL-1 and GST-4, which mediate the stress resistance and longevity of C. elegans. **P<0.01; *P<0.05.