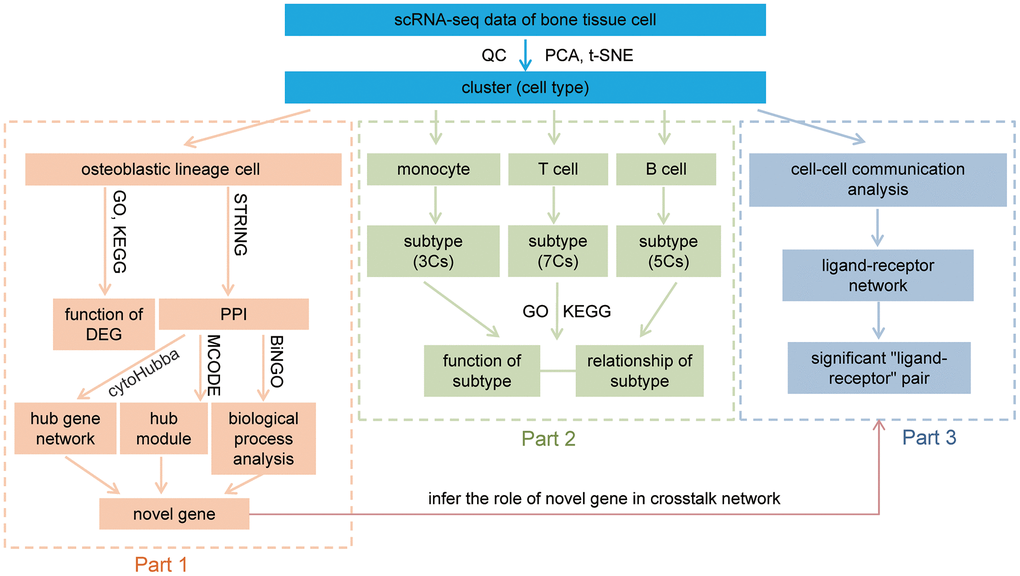
Figure 9.Workflow of this study. After QC, dimension reduction, and clustering of the data, we identify nine cell types in our data. The downstream analysis was divided into three parts. Part 1, analysis of osteoblastic lineage cells, functional analyses of osteoblastic lineage cells and identify novel bone metabolism-related gene. Part 2, revealing distinct subtypes in monocytes, T cells and B cells, and discussion their relationship with bone metabolism. Part 3, constructing the communication networks of human femoral head tissue cells, and inferring the role of novel metabolism-related gene in crosstalk network. QC: quality control; PCA: principal-component analysis; t-SNE: t-Distributed Stochastic Neighbor Embedding; GO: gene ontology enrichment analysis; KEGG: Kyoto Encyclopedia of Genes and Genomes enrichment analysis; DEG: differentially expressed gene; PPI: protein-protein interaction; MCODE: Molecular Complex Detection; Cs: clusters.