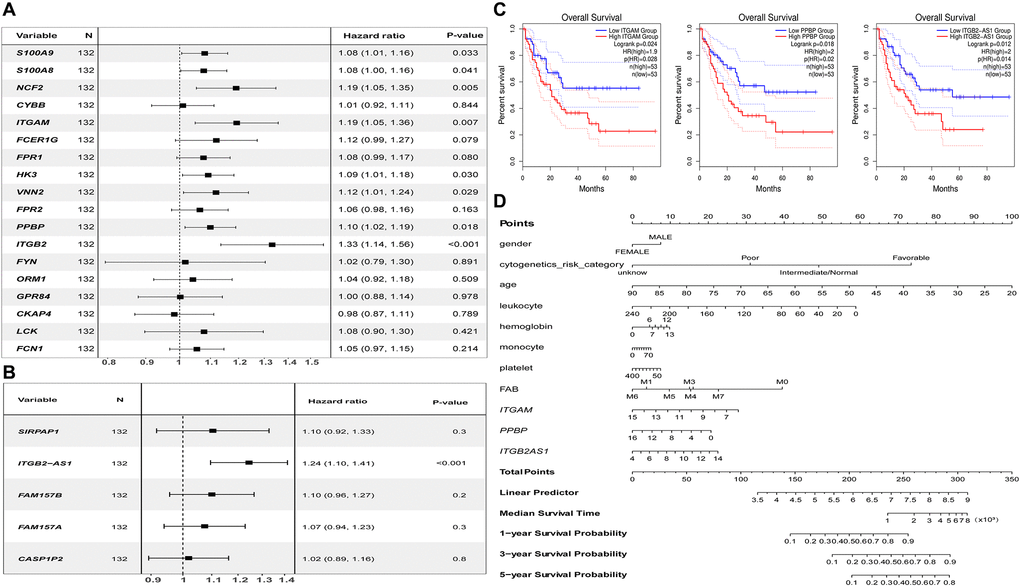
Figure 8.Prognostic analysis results of essential mRNAs and co-expressed DElncRNAs. (A) The result of univariate Cox regression analysis showed that the key genes such as S100A8 (HR:1.119, 95% CI:1.01–1.16), S100A9 (HR:1.08, 95% CI:1.00–1.16), NCF2 (HR:1.19, 95% CI:1.05–1.35), ITGAM (HR:1.19, 95% CI:1.05–1.36), HK3 (HR:1.09, 95% CI:1.01–1.08), VNN2 (HR:1.12, 95% CI:1.01–1.24), PPBP (HR:1.10, 95% CI:1.02–1.19), and ITGB2 (HR:1.33, 95% CI:1.14–1.56) both have significant impact on the prognosis of AML patients (P < 0.05). (B) The development of the research showed that the expression of DElnRNAs as ITGB2-AS1 (HR:1.24, 95% CI:1.10–1.41) has a significant impact on the prognosis of AML patients (P < 0.05). (C) The results of K–M survival analysis showed that AML patients with high expression of ITGAM, PPBP, and ITGB2-AS1 had a poor prognosis (P < 0.05). (D) The Nomogram was established based on the clinical information of TCGA-LAML. The points for 11 factors (gender, cytogenetics risk category, age, leukocyte, hemoglobin, monocyte, platelet, FAB classification, and the expression level of ITGAM, PPBP, or ITGB2-AS1) were listed in the Nomogram. The score for each factor in the Nomogram was read out by drawing a straight line from the predictor to the point axis, and then the survival rates of 1, 3, and 5 years could be estimated by adding the points corresponding to each factor in the bottom scale. Abbreviations: DElncRNAs: differentially expressed lncRNAs; HR: hazard ratio; CI: confidence interval.