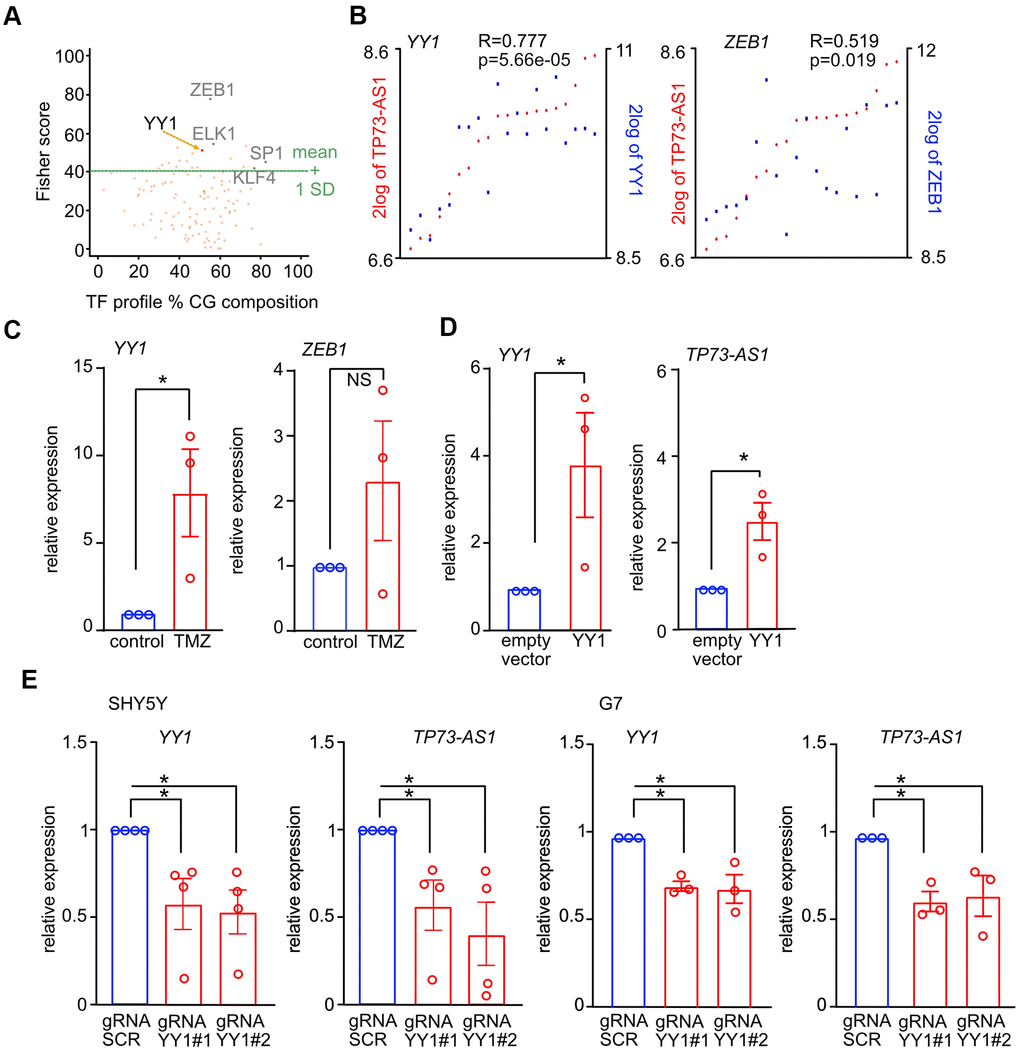
Figure 4.YY1 promotes the expression of TP73-AS1 upon TMZ treatment. (A) TF binding sites statistically enriched in the list of genes up-regulated upon TMZ treatment in G7 cells were found using oPPOSUM (full list in Supplementary Table 1). Key TFs known to play a role in GBM are indicated. (B) Co-expression between the indicated TFs and TP73-AS1 in GBM stem cells is shown. Data obtained using R2 and the Pollard dataset (GSE15209) [48]. (C) The levels of the indicated transcripts in SHY5Y cells treated or not with TMZ were measured using qRT-PCR. * p<0.05 in the two-tailed t-test; NS, not significant. (D) The levels of the indicated transcripts in SHY5Y cells transfected with the indicated constructs were measured using RT-qPCR. * p<0.05 in the two-tailed t-test; NS, not significant. (E) The indicated cell lines were engineered to express KREB-dCAS9 and gRNAs targeting YY1. Cells were treated with DOX for 10 days, to induce KREB-dCAS9. The levels of the indicated transcripts were measured using qRT-PCR. * p<0.05 in the two-tailed t-test; NS, not significant.