Introduction
Glioblastoma multiform (GBM) is a cancer of the brain with a dismal outcome and a five-year survival rate of < 10% [1]. Its location in the brain, recurrence, and tendency to infiltrate areas adjacent to the primary tumor are some of the major features contributing to its aggressiveness. Current treatments rely on surgical resection, radio therapy and temozolomide (TMZ) administration [2], which extends patient survival. However, therapy resistance in tumor cells is a major limiting factor in therapeutic success [3, 4]. It is therefore important to advance our understanding of resistance mechanisms for the development of new much needed therapies.
Aging contributes to cancer [5, 6] and is a factor associated with poor survival in GBM animal models [7] and patients [8, 9]. In particular, epi-genetic changes affecting gene expression strongly correlated with age in low grade glioma [10]. The molecular mechanisms linking aging and GBM are still vague and were proposed to be related to the gain of function of tumor associated fibroblasts [11] and functional decline of the immune system [7]. Nevertheless, the molecular details linking aging to GBM aggressiveness are still vague.
Long noncoding RNA (lncRNA) are regulatory RNA molecules known to play important roles in cancer such as promoting resistance to therapy [12, 13]. The lncRNA TP73-AS1 is a gene neighbor of the transcription factor (TF) p73, a member of the p53 TF family [14] known to play important roles in aging [15–17], cancer [18–23] and brain development [24–27] by regulating gene expression at the transcriptional and translational levels [28, 29]. LncRNA function by diverse mechanisms including by regulating gene expression in cis [30] however, TP73-AS1 does not regulate p73 in GBM stem cells [31] and to best of our knowledge was not found to regulate p73 in other biological scenarios.
TP73-AS1 is a negative prognostic factor in several tumor types [32]. In glioma, the expression of TP73-AS1 was shown to be associated with poor patient outcome and, importantly, with aging [33]. In GBM cancer stem cells, TP73-AS1 was found to promote TMZ resistance by facilitating the expression of the TMZ detoxifying enzyme, ALDH1A1 [31]. In accordance, TP73-AS1 is clinically relevant in GBM and its high expression in GBM tumors is associated with poor patient outcome. Furthermore, TP73-AS1 is highly expressed in the more aggressive IDH WT and EGFR amplified GBM tumors [31]. Interestingly, TP73-AS1 is relevant and functional in other brain tumors including medulloblastoma [34] and astrocytoma [35, 36]. Nevertheless, how the expression of TP73-AS1 is regulated in the context of TMZ treatment is not known, nor is it known if the expression of TP73-AS1 is associated with aging in brain.
Here we found that TP73-AS1 is highly expressed in the aging brain, and under pathological conditions, its expression is increased. We discovered that the TF Yin Yang 1 (YY1) regulates TP73-AS1 expression. YY1 was known to contribute to TMZ resistance by promoting the expression of DNA repair genes [37], and to be a part of the transcriptional program of the aging brain [38]. Importantly, YY1 directly activates the transcription of TP73-AS1 upon TMZ treatment. Together, these findings bring forth a new mode of regulation implicated in TMZ resistance in GBM and an intriguing link between aging and GBM.
Materials and Methods
RNA extraction, cDNA synthesis, and quantitative reverse transcription polymerase chain reaction (qRT-PCR)
RNA from cultured cells was purified using the NUCLEOSPIN RNA PLUS (Macherey-Nagel). From a starting amount of 100–200 ng of RNA, cDNA was generated using the cDNA synthesis kit (E6300S, BioLabs). Briefly, 20 μl of the cDNA synthesis reaction was subjected to the following conditions: 5 min at 25° C, 60 min at 42° C, and 5 min at 80° C. The qPCR conditions were as follows: initialization step at 95° C for 10 min, 46 cycles of the denaturation step at 60° C for 30 s, annealing step at 60° C for 30 s, and elongation step at 72° C for 1 s, followed by the final elongation at 40° C for 10 min to ensure that any remaining single-stranded DNA is fully extended. Forward and reverse primers were designed to span different exons whenever possible. All primers were purchased from Integrated DNA Technologies (IDT). qRT-PCR was performed using an iCycler (Bio-Rad Laboratories) with a threshold cycle number determined with the use of iCycler software version. Reactions were performed in triplicate and threshold cycle numbers were averaged. The results were normalized to L32.
RT-PCR primers used in this study
TP73-AS1: probe #29, primer1 ctccggacactgtgttttctc, primer2 tcttttaaggcggccatatc;
L32: probe #33, primer1 gcacactgactacagccttga, primer2 taccc aggtttggaggtgtg;
YY1: probe #79: primer1 ttggagagaactcacctcctg, primer2 gccgagttatccctgaacat;
ZEB1: probe #68: primer1 gccaacagaccagacagtgtt, primer2 tttcttgcccttcctttctg;s.
Plasmids
psPAX2 was a gift from Didier Trono (Addgene plasmid # 12260). pMD2.G was a gift from Didier Trono (Addgene plasmid # 12259). pHAGE TRE dCas9-KRAB was a gift from Rene Maehr and Scot Wolfe (Addgene plasmid # 50917). pLKO.1-puro U6 sgRNA BfuAI large stuffer was a gift from Scot Wolfe (Addgene plasmid # 52628). Rennila and luciferase (pGL 4.23) vectors were a kind gift from Gabriel Leprivier. The two parts of the promoter (824BP and 1124BP) were cloned into the Firefly luciferase using TEDA method [39] between XhoI and HindIII restriction enzyme sites. The mutated YY1 binding site promoter was cloned using standard site directed mutagenesis. YY1 over expression vector was described in [38]. We used pLKO.1-puro U6 sgRNA BfuAI large stuffer. pLKO.1-puro U6 sgRNA BfuAI large stuffer was a gift from Scot Wolfe (Addgene plasmid # 52628). The primer sequences used were as follows:
scramble/control: ACCGCGCCAAACGTGCCCTGACGG
YY1 g#1: GGAGACCATCGAGACCACAG;
YY1 g#2: CGACACCCTCTACATCGCCA.
Generation of stable cell lines for gene knockdown
Lentiviruses were generated using our standard protocol and LentiX cells. Cells were grown in DMEM, 10% FBS, and transfected using CalFectin (Signagen) and plasmids at a ratio of 1:2:3 (PAX2; pMD2.G; transfer vector). The media was changed after 24 h and the viruses were collected after 48 h. The collected viruses were stored at −80° C. G7 and SHSY5Y cells were grown to 80% confluence on 6-cm dishes, then infected with a vector ratio of 1:10 (cas9, gRNA), and left for 24 h in a 37° C incubator. Following infection, the lentivirus containing medium was removed and replaced by fresh medium and incubated at 37° C with the selection antibiotic using puromycin (1 μg/ml) and G418 (1500 μg/ml).
CRIPSRi
The depletion of the YY1 was done using the CRISPRi method combined with viral infection. gRNA were designed to target the promoter region of the YY1 gene. Knockdown induction was done using doxycycline at a concentration of 2 μg/ml (Sigma-Aldrich, D3072). Gene knockdown was measured using qRT-PCR.
TMZ treatment
YY1 KD was induced by addition of doxycycline for 10 days. Luciferase Activity of the different TP73-AS1 promoter parts was measured by luciferase reporter gene assay (Promega, E1500) in SHSY5Y and HEK293 cell lines. The SHSY5Y cells were transient transfected (Lipojet, BioConsult SL100468) with the reporter and control plasmids, and were cultured in DMEM medium with 10% fetal bovine serum. 24h later, some wells were treated with DMSO and others with TMZ (750uM) for 2 days. Luciferase activity was measured and normalized by measuring co-transfected Renilla plasmid using the Dual-Luciferase® Reporter Assay System, Promega.
Western blot
Western blots were performed as in [40]. In short, lysates were mixed with sample loading buffer 5X (250 mM Tris·HCl pH 6.8; 10% SDS; 30% Glycerol; 10 mM DTT; 0.05% (w/v) Bromophenol Blue), followed by 10 minutes incubation at 96° C and spin down. Samples were then loaded to SDS-PAGE followed by transferring to nitrocellulose membrane. Membranes were then blocked by soaking in milk (5% skim milk; Tris buffer saline x1 contains 0.05% TWEEN20 (TBST)) for 1 hour. Membranes were then covered with primary antibody against YY1 solution (abcam ab109237) for 1 hour in room temperature or overnight in 4° C, followed by washing with TBST and incubation with secondary antibodies (Cell signaling technology,7074S) for 1 hour in room temperature.
Results
TP73-AS1 is highly expressed in the aging brain
To learn more about how the expression of TP73-AS1 is regulated, and given that its expression is increased during aging in glioma [33], we first asked if TP73-AS1 expression is associated with natural aging in the brain and interrogated the Cotman dataset [42], using R2 software [45] and 60-year-old brains as a cutoff. We found that TP73-AS1 is highly expressed in the aged brain and this is the case in most brain regions to which data is available (Figure 1A). In accord, the expression of TP73-AS1 in the brain is correlated with aging (Figure 1B). Together, these data suggest that TP73-AS1 is highly expressed in the aging brain.
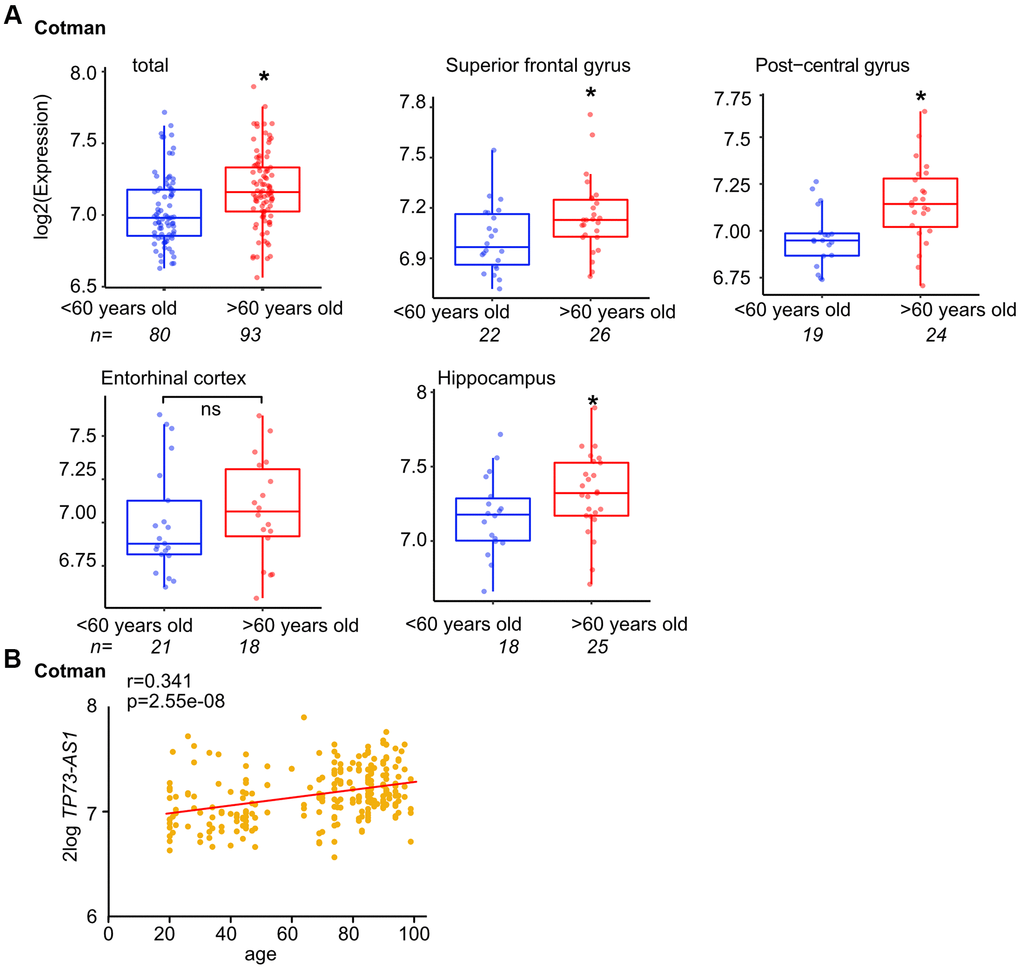
Figure 1. TP73-AS1 is highly expressed in the aging brain. (A) The levels of TP73-AS1 in the old vs. young brain are shown. Data were obtained from the R2 website and the indicated dataset (GSE48350). (B) The correlation between the expression of TP73-AS1 and age in the brain was determined using R2 and indicated dataset (GSE48350).
We next asked if TP73-AS1 expression is also linked to pathological aging in the human brain. To this end, we interrogated the Cotman dataset and found that TP73-AS1 is highly expressed in Alzheimer’s brain and that this is the case in most tested brain regions (Figure 2A). We confirmed these findings using a second dataset, the Salomon dataset [41] (Figure 2B) which supported our conclusion. While in the hippocampus (Cotman dataset), the median temporal and superior frontal gyrus (Salomon dataset) TP73-AS1 expression was trending higher in Alzheimer’s vs. normal brain, these differences did not reach statistical significance, possibly due to a lower number of samples as compared with the total which was obtained by pooling of all samples from different brain regions.
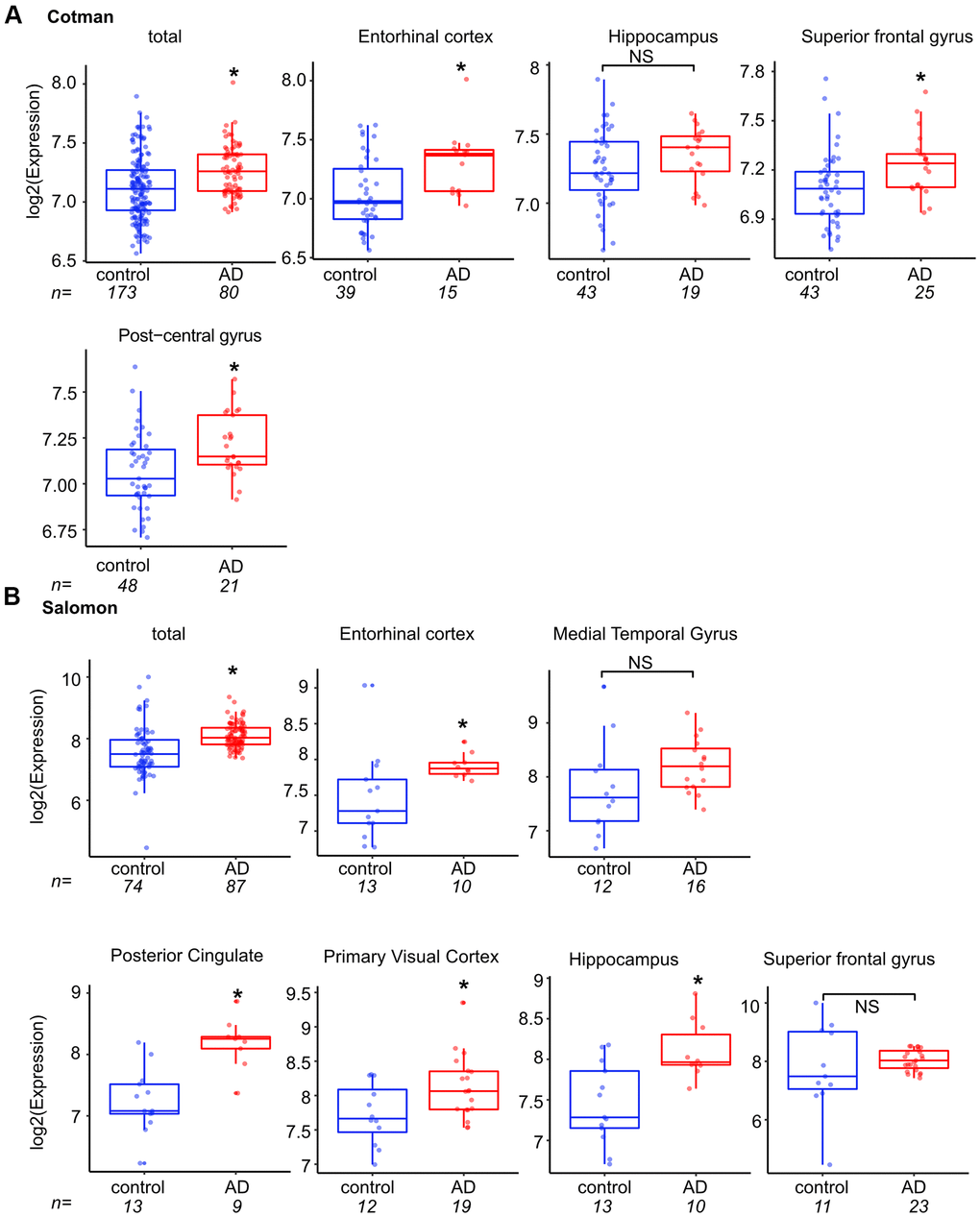
Figure 2. TP73-AS1 is highly expressed in the pathological aging brain. (A) The levels of TP73-AS1 in Alzheimer’s (AD) vs. normal brain are shown. Data were obtained from R2 and the Cotman dataset (GSE48350). In both (A, B) panels, asterisks indicate statistically significant differences as calculated by the two-sample Wilcoxon test (*p < 0.05; NS, non-significant). (B) The levels of TP73-AS1 in Alzheimer’s vs. normal brain are shown. Data were obtained from R2 and the Salomon dataset (GSE5281).
The high expression TP73-AS1 in the aging brain and our previous findings that TP73-AS1 is highly expressed in GBM tumors, lead us to ask if the age of the patient is associated with TP73-AS1 expression in GBM tumors. We analyzed TP73-AS1 expression in GBM tumors of old and young patients using 60-year-old as a cutoff. Indeed, TP73-AS1 is up-regulated in GBM tumors of older patients (Supplementary Figure 1A). In addition, we calculated the correlation between TP73-AS1 expression and patient’s age and found that they are positively correlated (Supplementary Figure 1B).
Interestingly, we recently found that the TF YY1 and its gene targets are a prominent part of the pathological aging brain transcriptional program [38] and were intrigued as to whether YY1 is a TP73-AS1 TF as well.
TP73-AS1 is induced by TMZ
Previously, we found that TP73-AS1 protects GBM stem cells from TMZ, an alkylating agent, by promoting the expression of detoxifying genes such as ALDH1A1 [31], which neutralize aldehydes and other toxic molecules in the cell [46, 47]. We asked if TP73-AS1 is induced by TMZ, as is expected from a gene promoting a detoxification transcriptional response. We treated two GBM cancer stem cell models, G7 and G26 [48], and a neuroblastoma model, SHY5Y, with TMZ (750 μM for 48 hours), after which we measured the expression of TP73-AS1 using qRT-PCR (Figure 3A). We found that TP73-AS1 is induced in TMZ treated cells.
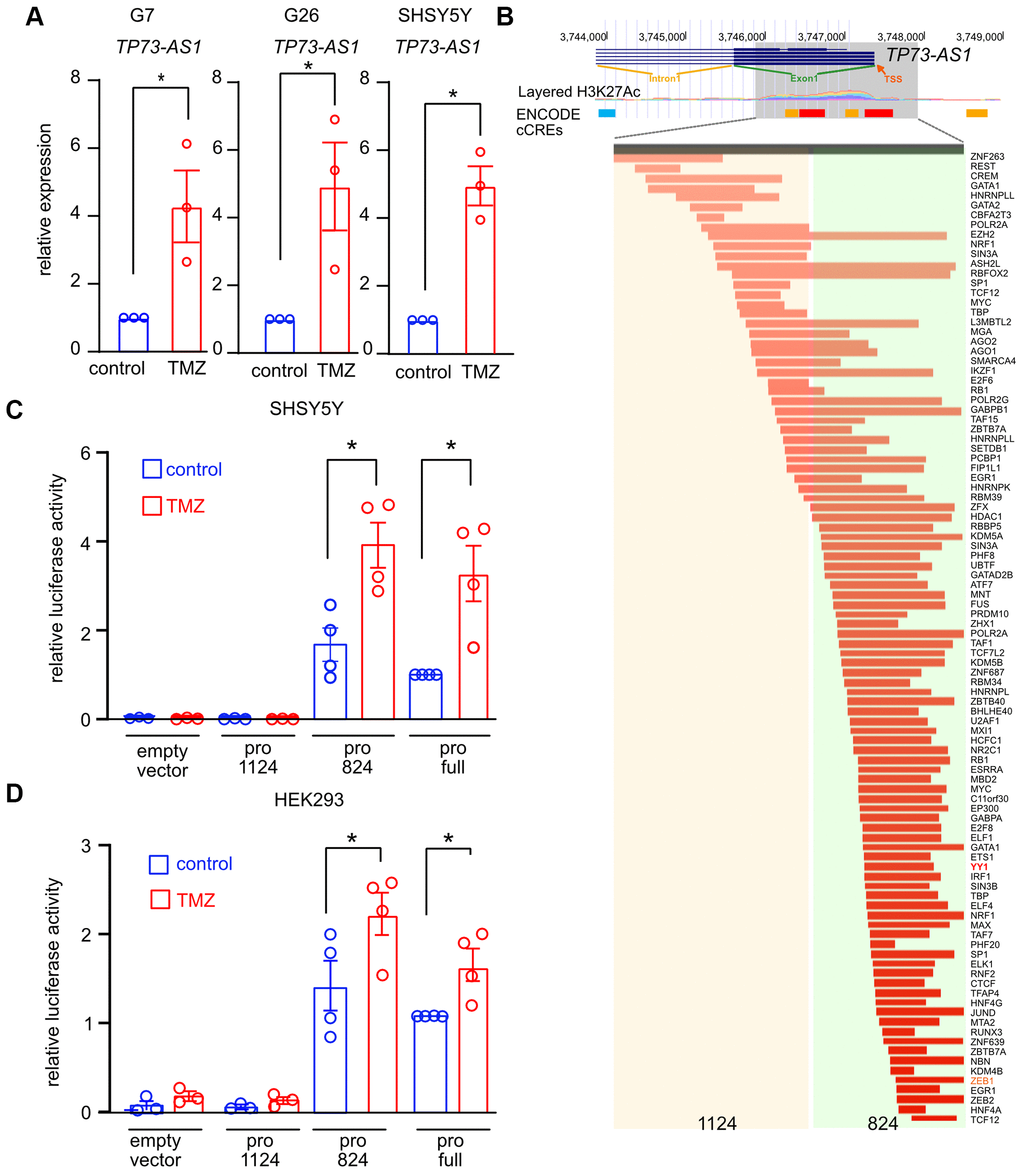
Figure 3. TP73-AS1 is induced by TMZ. (A) TP73-AS1 levels in the indicated cell lines were measured using qRT-PCR. * p<0.05 in the two-tailed t-test. (B) The genomic region of the putative TP73-AS1 promoter was identified using the UCSC browser. H3K27 acetylation and TF ChIP-SEQ data obtained from ENCODE are shown. Transcriptional start site (TSS), intron 1, and exon 1 are shown. The two promoter regions used in this study are highlighted. (C) Promoter activity assay was performed in the indicated cell lines using the dual luciferase assay. * p<0.05 in the two tailed t-test. (D) Promoter activity assay was performed in the indicated cell lines using the dual luciferase assay. * p<0.05 in the two tailed t-test.
We next asked if TP73-AS1 expression is induced by a TF binding to its promoter. The sequence we defined as the promoter was chosen using ENCODE ChIP-seq data obtained using antibodies against histone K27 acetylation and specific TFs [49–52] (Figure 3B). To monitor promoter activity, we used the dual luciferase system. In short, the putative promoter sequence is cloned upstream to Firefly luciferase and cells are transfected with the vector harboring the promoter along with the transfection control vector encoding Renilla luciferase. Firefly and Renilla luciferase activity are measured, and the ratio between the two measurements reflects promoter activity. Transfected cells were treated with TMZ (750 μM for 48 hours) after which we measured Firefly/Renilla luciferase activity. We found that the full promoter is active in basal conditions and is further induced upon TMZ treatment (Figure 3C, 3D).
Based on the ENCODE TF binding data [49–52], we divided the promoter into two regions and cloned each region upstream to Firefly luciferase (Figure 3B). We transfected cells with each construct and measured Firefly and Renilla luciferase activity in basal and TMZ treated conditions (Figure 3C, 3D). We found that the 1124-promoter region was inactive in TMZ treated and non-treated cells, while the 824-promoter region was active in non-treated cells and induced upon TMZ treatment. Interestingly, YY1 was found to bind the 824-TP73-AS1 promoter region (ENCODE data; Figure 3B). We, therefore, concluded that TP73-AS1 is a TMZ responsive gene and that the TF activating TP73-AS1 upon TMZ treatment is most likely to be one of the TFs mapped to the 824-region.
YY1 promotes the expression of TP73-AS1 upon TMZ treatment
To identify the TF activating TP73-AS1 upon TMZ treatment, we asked if there are specific TFs binding sites enriched in the list of genes upregulated during TMZ treatment, assuming that TP73-AS1 is part of a TMZ transcriptional program whose genes may be regulated by a common TF. To this end, we analyzed the list of genes up regulated by TMZ in G7 GBM stem cells using data found in our previous publication [31] and oPPOSUM [53–55], and identified several TF consensus sites enriched in the promoters of these genes, including YY1 and ZEB1 (Figure 4A and Supplementary Table 1), both of which were found to be physically associated with the 824-promoter region (Figure 3B). Interestingly, both YY1 and ZEB1 are co-expressed with TP73-AS1 in GBM stem cells (Figure 4B). This is not the case in bulk GBM tumor data (Supplementary Figure 2A), possibly due to TP73-AS1 function in GBM stem cells, a sub-population of cells in bulk tumor [56]. Note, the r and p values are more significant in the case of YY1. To study the correlation of expression between TP73-AS1 and YY1 in normal aging we used GTEx data [43] and found a positive correlation in the different brain regions (Supplementary Figure 2B).
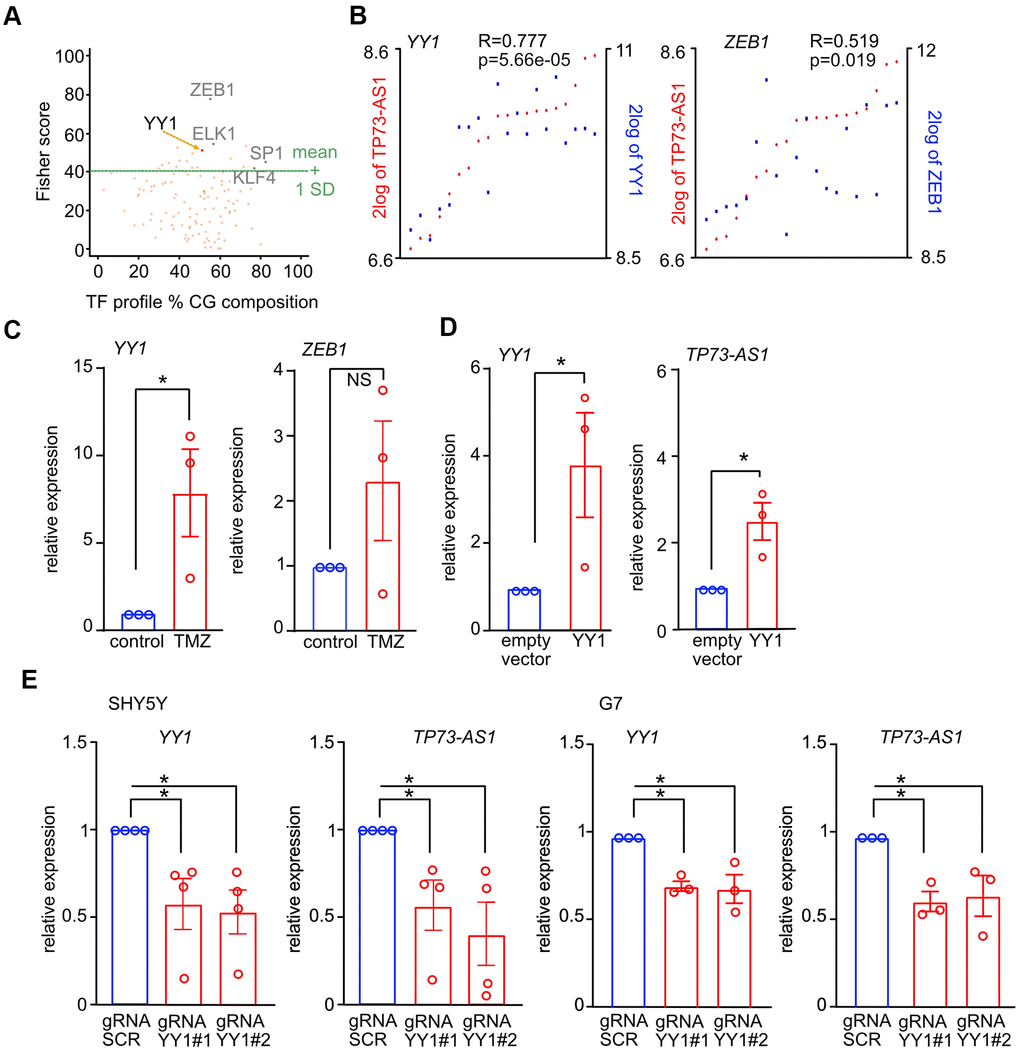
Figure 4. YY1 promotes the expression of TP73-AS1 upon TMZ treatment. (A) TF binding sites statistically enriched in the list of genes up-regulated upon TMZ treatment in G7 cells were found using oPPOSUM (full list in Supplementary Table 1). Key TFs known to play a role in GBM are indicated. (B) Co-expression between the indicated TFs and TP73-AS1 in GBM stem cells is shown. Data obtained using R2 and the Pollard dataset (GSE15209) [48]. (C) The levels of the indicated transcripts in SHY5Y cells treated or not with TMZ were measured using qRT-PCR. * p<0.05 in the two-tailed t-test; NS, not significant. (D) The levels of the indicated transcripts in SHY5Y cells transfected with the indicated constructs were measured using RT-qPCR. * p<0.05 in the two-tailed t-test; NS, not significant. (E) The indicated cell lines were engineered to express KREB-dCAS9 and gRNAs targeting YY1. Cells were treated with DOX for 10 days, to induce KREB-dCAS9. The levels of the indicated transcripts were measured using qRT-PCR. * p<0.05 in the two-tailed t-test; NS, not significant.
We measured YY1 and ZEB1 expression in TMZ treated and control cells and found that the expression of YY1 but not ZEB1 was significantly induced upon TMZ treatment (Figure 4C). Note, we tested if YY1 is also induced at the protein level and found that this is not the case (Supplementary Figure 3). We therefore conclude that the YY1 gene responds to TMZ treatment and that YY1 putative function is not due to increased expression.
Having found that YY1 responds to TMZ and that YY1 target genes are part of the aging brain transcriptional program [38], we hypothesized that YY1 plays a major role in promoting TP73-AS1 expression upon TMZ treatment. We next asked if YY1 is involved in TP73-AS1 up regulation upon TMZ treatment. We over expressed YY1 in SHSY5Y cells and measured TP73-AS1 levels using RT-qPCR (Figure 4D). YY1 promoted TP73-AS1 expression. Next, we used CRISPRi to down regulate YY1 expression, with two independent gRNAs, and measured TP73-AS1 levels using qRT-PCR in basal conditions (Figure 4E). We found that YY1 depletion led to reduced TP73-AS1 expression and therefore concluded that YY1 is a major TP73-AS1 regulator. Indeed, the expression of TP73-AS1 and YY1 is correlated in both human pathological aging (AD) brain datasets (Supplementary Figure 2C).
In our previous work studying TP73-AS1 in GBM or medulloblastoma, we asked if the expression of TP73-AS1 and p73 are correlated, as one would expect if they were to regulate each other in cis, and the answer was negative [31, 34]. To address the possibility that during aging TP73-AS1 regulates p73 (or vice versa), we performed a co-expression analysis and found that there is significant co-expression in one dataset (Salomon) but not the other (Cotman) (Supplementary Figure 2D). This discrepancy could be due to the higher number of samples in the former. Considering that the role of p73 in the aging brain is under debate [57], we decided to focus on YY1 as a putative regulator of TP73-AS1.
YY1 directly promotes TP73-AS1 expression and promoter activation
To investigate the interaction between YY1 and the promoter of TP73-AS1, we over expressed YY1 and measured the 824-promoter activity using luciferase activity assay (Figure 5A). We found that YY1 over expression increased 824-promoter activity. Importantly, CRISPRi knock down of YY1 prevented the induction of TP73-AS1 upon TMZ treatment (Figure 5B).
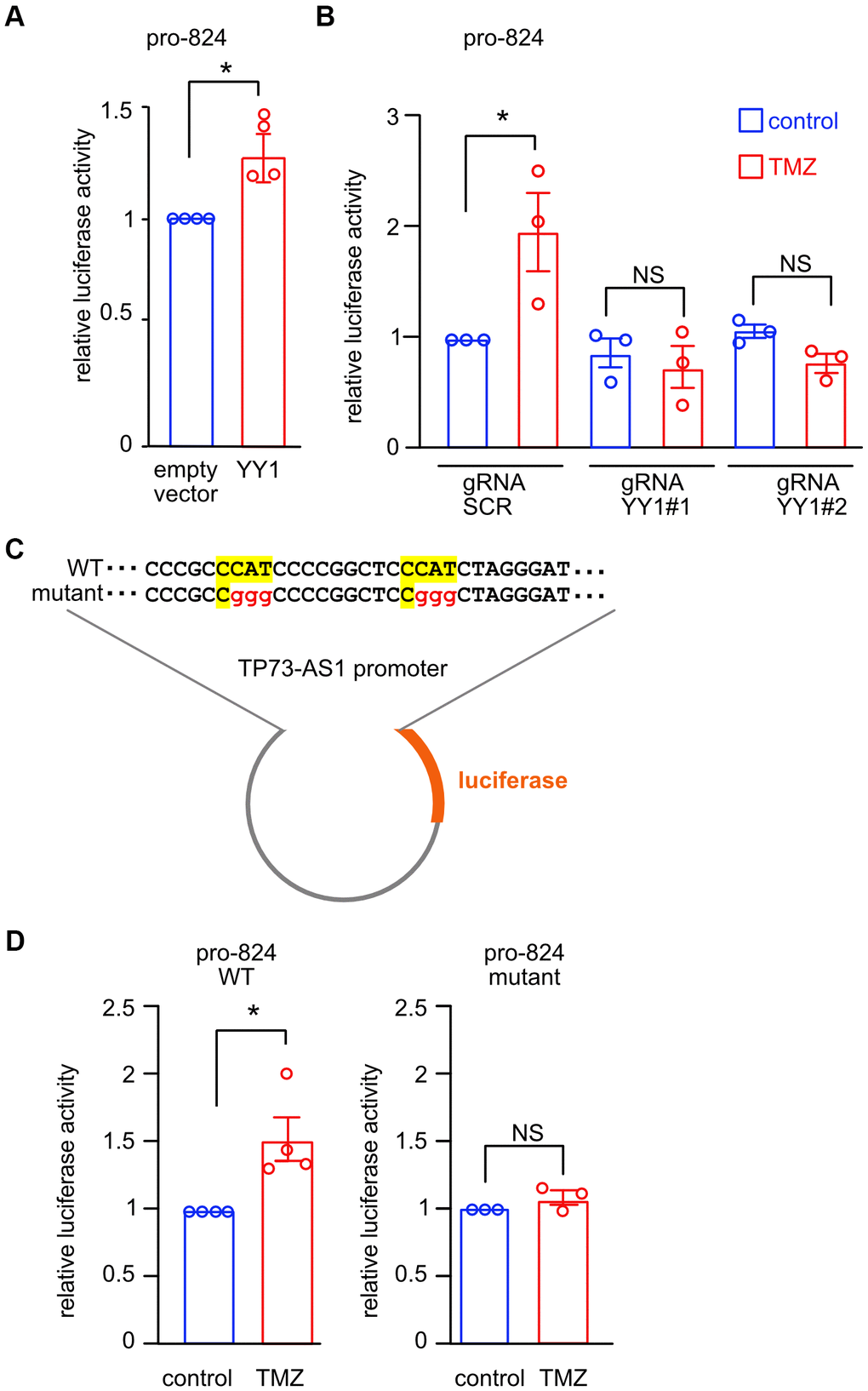
Figure 5. YY1 directly promotes TP73-AS1 promoter activation. (A) The indicated promoter activity was measured in SHY5Y cells over expressing YY1 or empty vector. * p<0.05 in the two-tailed t-test; NS, not significant. (B) CRISPRi SHY5Y cells were treated with DOX for 10 days to induce YY1 KD. Cells were treated with TMZ (750 μM for 48 hours). The activity of TP73-AS1-promoter-824 was measured using the dual luciferase assay. * p<0.05 in the two-tailed t-test; NS, not significant. (C) Schematic representation of the WT or mutant YY1 binding sites within the TP73-AS1 promoter luciferase construct. (D) SHY5Y cells expressing WT or mutant promoter were treated with TMZ or DMSO control for 48 hours after which promoter activity was measured using dual luciferase assay. * p<0.05 in the two-tailed t-test; NS, not significant.
We identified two canonical YY1 binding sites in the 824-promoter using Jaspar [31] and mutated the two binding sites to study their importance for promoter activation upon TMZ treatment (Figure 5C). We treated cells, expressing the wild type or mutant 824-promoter, with TMZ (750 μM for 48 hours), after which we measured Firefly/Renilla luciferase activity (Figure 5D). TMZ treatment led to increased promoter activation of the wild type but not the mutant 824-promoter. Together, these data strongly suggest that YY1 promotes TP73-AS1 induction upon TMZ treatment by binding to and activating the 824-promoter region.
Discussion
GBM aggressiveness is tightly linked to temozolomide resistance [58]. The lncRNA TP73-AS1 protects GBM stem cells from TMZ toxicity [31]. Previous studies of TP73-AS1 expression in brain tumors focused on genetic and epigenetic mechanisms to explain its aberrant expression. In oligodendroglial tumors, reduced TP73-AS1 expression was identified and correlated with a deletion of the p36.31–p36.32 region on chromosome 1, where TP73-AS1 resides, and with TP73-AS1 promoter hyper methylation [35]. In a study aimed at identifying GBM epigenetic subgroups, downregulation of TP73-AS1 expression and hypermethylation of its promoter were identified as defining features of the IDH/G-CIMP+ subgroup [59]. Here, we found that YY1 promotes TP73-AS1 expression and the activity of its promoter upon TMZ treatment, thus adding a mechanistic explanation to how TP73-AS1 is induced in this context. Such a TF-lncRNA axis promoting TMZ resistance has been previously found. The lncRNA MALAT1 is positively regulated by the TFs NF-kappaB and p53 which upon TMZ treatment, bind specific sequences within the MALAT1 promoter and activate it, increasing MALAT1 levels and, consequently, TMZ resistance [34].
Aging contributes to glioma incidence and aggressiveness and interestingly, the expression pattern of TP73-AS1 is associated with key features linked to aging and glioma aggressiveness. These include high expression of TP73-AS1 in EGFR amplified and IDH-wild type tumors [10]. These associations are in line with our findings that TP73-AS1 expression correlates with aging and aggressiveness providing a possible molecular link explaining how aging contributes to GBM aggressiveness.
YY1 is a TF shown to carry out pro-tumorigenic functions [60] including promoting the expression of c-MYC by directly binding its promoter [61] or inhibiting the tumor suppressor activity of p53 upon DNA damage [62, 63]. YY1 was also shown to promote the expression of the stemness factor KLF4 [64] and has been implicated in promoting the expression of stemness factors in cancer stem cells [65]. Interestingly, the levels and activity of the transcription factor YY1 and its gene targets decline with age in T-cells [66] and increase with age in brain [38]. Moreover, higher mRNA levels of YY1 were found in Alzheimer diseased brain and YY1 was defined as a “master regulator” in Alzheimer disease [67]. Furthermore, in GBM stem cells, YY1 was shown to promote stemness, TMZ resistance and tumorigenicity, through yet unknown mechanisms [37].
Here we found that YY1 induces the expression of TP73-AS1 upon TMZ treatment, thus providing a possible explanation for the protective functions of YY1, in light of the known function of TP73-AS1 in promoting TMZ resistance in GBM stem cells [31]. Although YY1 transcript level increase in response to TMZ treatment, and directly induces the expression of TP73-AS1, YY1 protein levels were not induced. Reasonable explanation for the upregulation at the mRNA levels but not the protein level, is that TMZ treatment may induce a cellular stress leading to inhibition of translation [68]. Nevertheless, YY1 is modified and regulated post-translationally. For example, YY1 is marked for degradation by SMURF2 E3 ubiquitin ligase [69], is tyrosine phosphorylated by the SRC kinases family [70] or subjected to acetylation and deacetylation [70]. Considering YY1 regulates multiple genes, the contribution of TP73-AS1 to a given YY1 biological function is yet to be determined.
Aberrant YY1 function in aging brain was recently reported and attributed to reduced expression of its binding partner, SIRT6 [38]. Reduced SIRT6 expression, occurring during aging, leads to changes in the expression of genes which are regulated by YY1, many of which are involved in pathological aging. It is therefore possible that TP73-AS1 is a YY1 target in aging brain.
In conclusion, we show that TP73-AS1 levels increase in pathological and natural aging brain and upon TMZ treatment, and that YY1 directly activates the TP73-AS1 promoter to induce its expression. These findings provide a plausible explanation for how the expression of TP73-AS1 is regulated, and an interesting molecular link between aging and GBM.
Author Contributions
Conception and design: Gal Mazor, Deborah Toiber; Barak Rotblat; Collection and assembly of data: Gal Mazor, Dmitri Smirnov, Hila BenDavid, Ekaterina Khrameeva; Data analysis and interpretation: Gal Mazor, Dmitri Smirnov, Hila Ben David, Ekaterina Khrameeva; Manuscript writing: Gal Mazor, Deborah Toiber, Ekaterina Khrameeva, Barak Rotblat; All authors read and approved the final manuscript.
Conflicts of Interest
The authors declare that they have no conflicts of interest.
Funding
BR: This research was supported by the Israel Science Foundation (grant No. 1436/19), by the Israeli Cancer Association (grant #20180012) and the NIBN.
DT: This work was supported by The David and Inez Myers foundation and by the Ministry of Science and Technology, Israel.
EK: This work was supported by RFBR, project number 20-34-70077.
References
- 1. Reifenberger G, Wirsching HG, Knobbe-Thomsen CB, Weller M. Advances in the molecular genetics of gliomas - implications for classification and therapy. Nat Rev Clin Oncol. 2017; 14:434–52. https://doi.org/10.1038/nrclinonc.2016.204 [PubMed]
- 2. Stupp R, Mason WP, van den Bent MJ, Weller M, Fisher B, Taphoorn MJ, Belanger K, Brandes AA, Marosi C, Bogdahn U, Curschmann J, Janzer RC, Ludwin SK, et al, and European Organisation for Research and Treatment of Cancer Brain Tumor and Radiotherapy Groups, and National Cancer Institute of Canada Clinical Trials Group. Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med. 2005; 352:987–96. https://doi.org/10.1056/NEJMoa043330 [PubMed]
- 3. Lee SY. Temozolomide resistance in glioblastoma multiforme. Genes Dis. 2016; 3:198–210. https://doi.org/10.1016/j.gendis.2016.04.007 [PubMed]
- 4. Wheeler DA, Takebe N, Hinoue T, Hoadley KA, Cardenas MF, Hamilton AM, Laird PW, Wang L, Johnson A, Dewal N, Miller V, Piñeyro D, Castro de Moura M, et al. Molecular Features of Cancers Exhibiting Exceptional Responses to Treatment. Cancer Cell. 2021; 39:38–53.e7. https://doi.org/10.1016/j.ccell.2020.10.015 [PubMed]
- 5. de Magalhães JP. How ageing processes influence cancer. Nat Rev Cancer. 2013; 13:357–65. https://doi.org/10.1038/nrc3497 [PubMed]
- 6. Ben-Zion Berliner M, Yerushalmi R, Lavie I, Benouaich-Amiel A, Tsoref D, Hendler D, Goldvaser H, Sarfaty M, Rotem O, Ulitsky O, Siegal T, Neiman V, Yust-Katz S. Central nervous system metastases in breast cancer: the impact of age on patterns of development and outcome. Breast Cancer Res Treat. 2021; 185:423–32. https://doi.org/10.1007/s10549-020-05959-x [PubMed]
- 7. Ladomersky E, Zhai L, Gritsina G, Genet M, Lauing KL, Wu M, James CD, Wainwright DA. Advanced age negatively impacts survival in an experimental brain tumor model. Neurosci Lett. 2016; 630:203–08. https://doi.org/10.1016/j.neulet.2016.08.002 [PubMed]
- 8. Ladomersky E, Scholtens DM, Kocherginsky M, Hibler EA, Bartom ET, Otto-Meyer S, Zhai L, Lauing KL, Choi J, Sosman JA, Wu JD, Zhang B, Lukas RV, Wainwright DA. The Coincidence Between Increasing Age, Immunosuppression, and the Incidence of Patients With Glioblastoma. Front Pharmacol. 2019; 10:200. https://doi.org/10.3389/fphar.2019.00200 [PubMed]
- 9. Chaichana KL, Chaichana KK, Olivi A, Weingart JD, Bennett R, Brem H, Quiñones-Hinojosa A. Surgical outcomes for older patients with glioblastoma multiforme: preoperative factors associated with decreased survival. Clinical article. J Neurosurg. 2011; 114:587–94. https://doi.org/10.3171/2010.8.JNS1081 [PubMed]
- 10. Chatsirisupachai K, Lesluyes T, Paraoan L, Van Loo P, de Magalhães JP. An integrative analysis of the age-associated multi-omic landscape across cancers. Nat Commun. 2021; 12:2345. https://doi.org/10.1038/s41467-021-22560-y [PubMed]
- 11. Song H, Fu X, Wu C, Li S. Aging-related tumor associated fibroblasts changes could worsen the prognosis of GBM patients. Cancer Cell Int. 2020; 20:489. https://doi.org/10.1186/s12935-020-01571-7 [PubMed]
- 12. Bester AC, Lee JD, Chavez A, Lee YR, Nachmani D, Vora S, Victor J, Sauvageau M, Monteleone E, Rinn JL, Provero P, Church GM, Clohessy JG, Pandolfi PP. An Integrated Genome-wide CRISPRa Approach to Functionalize lncRNAs in Drug Resistance. Cell. 2018; 173:649–64.e20. https://doi.org/10.1016/j.cell.2018.03.052 [PubMed]
- 13. Zottel A, Šamec N, Videtič Paska A, Jovčevska I. Coding of Glioblastoma Progression and Therapy Resistance through Long Noncoding RNAs. Cancers (Basel). 2020; 12:1842. https://doi.org/10.3390/cancers12071842 [PubMed]
- 14. Amelio I, Melino G. Similar Domains for Different Regulations of p53 Family. Structure. 2018; 26:1047–49. https://doi.org/10.1016/j.str.2018.07.003 [PubMed]
- 15. Agostini M, Niklison-Chirou MV, Catani MV, Knight RA, Melino G, Rufini A. TAp73 promotes anti-senescence-anabolism not proliferation. Aging (Albany NY). 2014; 6:921–30. https://doi.org/10.18632/aging.100701 [PubMed]
- 16. Rufini A, Niklison-Chirou MV, Inoue S, Tomasini R, Harris IS, Marino A, Federici M, Dinsdale D, Knight RA, Melino G, Mak TW. TAp73 depletion accelerates aging through metabolic dysregulation. Genes Dev. 2012; 26:2009–14. https://doi.org/10.1101/gad.197640.112 [PubMed]
- 17. Lopriore P, Capitanio N, Panatta E, Di Daniele N, Gambacurta A, Melino G, Amelio I. TAp73 regulates ATP7A: possible implications for ageing-related diseases. Aging (Albany NY). 2018; 10:3745–60. https://doi.org/10.18632/aging.101669 [PubMed]
- 18. Tomasini R, Tsuchihara K, Wilhelm M, Fujitani M, Rufini A, Cheung CC, Khan F, Itie-Youten A, Wakeham A, Tsao MS, Iovanna JL, Squire J, Jurisica I, et al. TAp73 knockout shows genomic instability with infertility and tumor suppressor functions. Genes Dev. 2008; 22:2677–91. https://doi.org/10.1101/gad.1695308 [PubMed]
- 19. Flores ER, Sengupta S, Miller JB, Newman JJ, Bronson R, Crowley D, Yang A, McKeon F, Jacks T. Tumor predisposition in mice mutant for p63 and p73: evidence for broader tumor suppressor functions for the p53 family. Cancer Cell. 2005; 7:363–73. https://doi.org/10.1016/j.ccr.2005.02.019 [PubMed]
- 20. Du W, Jiang P, Mancuso A, Stonestrom A, Brewer MD, Minn AJ, Mak TW, Wu M, Yang X. TAp73 enhances the pentose phosphate pathway and supports cell proliferation. Nat Cell Biol. 2013; 15:991–1000. https://doi.org/10.1038/ncb2789 [PubMed]
- 21. Landré V, Antonov A, Knight R, Melino G. p73 promotes glioblastoma cell invasion by directly activating POSTN (periostin) expression. Oncotarget. 2016; 7:11785–802. https://doi.org/10.18632/oncotarget.7600 [PubMed]
- 22. Amelio I, Markert EK, Rufini A, Antonov AV, Sayan BS, Tucci P, Agostini M, Mineo TC, Levine AJ, Melino G. p73 regulates serine biosynthesis in cancer. Oncogene. 2014; 33:5039–46. https://doi.org/10.1038/onc.2013.456 [PubMed]
- 23. Amelio I, Inoue S, Markert EK, Levine AJ, Knight RA, Mak TW, Melino G. TAp73 opposes tumor angiogenesis by promoting hypoxia-inducible factor 1α degradation. Proc Natl Acad Sci USA. 2015; 112:226–31. https://doi.org/10.1073/pnas.1410609111 [PubMed]
- 24. Amelio I, Panatta E, Niklison-Chirou MV, Steinert JR, Agostini M, Morone N, Knight RA, Melino G. The C terminus of p73 is essential for hippocampal development. Proc Natl Acad Sci USA. 2020; 117:15694–701. https://doi.org/10.1073/pnas.2000917117 [PubMed]
- 25. Yang A, Walker N, Bronson R, Kaghad M, Oosterwegel M, Bonnin J, Vagner C, Bonnet H, Dikkes P, Sharpe A, McKeon F, Caput D. p73-deficient mice have neurological, pheromonal and inflammatory defects but lack spontaneous tumours. Nature. 2000; 404:99–103. https://doi.org/10.1038/35003607 [PubMed]
- 26. Agostini M, Tucci P, Killick R, Candi E, Sayan BS, Rivetti di Val Cervo P, Nicotera P, McKeon F, Knight RA, Mak TW, Melino G. Neuronal differentiation by TAp73 is mediated by microRNA-34a regulation of synaptic protein targets. Proc Natl Acad Sci USA. 2011; 108:21093–98. https://doi.org/10.1073/pnas.1112061109 [PubMed]
- 27. Niklison-Chirou MV, Steinert JR, Agostini M, Knight RA, Dinsdale D, Cattaneo A, Mak TW, Melino G. TAp73 knockout mice show morphological and functional nervous system defects associated with loss of p75 neurotrophin receptor. Proc Natl Acad Sci USA. 2013; 110:18952–57. https://doi.org/10.1073/pnas.1221172110 [PubMed]
- 28. Marini A, Rotblat B, Sbarrato T, Niklison-Chirou MV, Knight JR, Dudek K, Jones C, Bushell M, Knight RA, Amelio I, Willis AE, Melino G. TAp73 contributes to the oxidative stress response by regulating protein synthesis. Proc Natl Acad Sci USA. 2018; 115:6219–24. https://doi.org/10.1073/pnas.1718531115 [PubMed]
- 29. Rotblat B, Agostini M, Niklison-Chirou MV, Amelio I, Willis AE, Melino G. Sustained protein synthesis and reduced eEF2K levels in TAp73-\- mice brain: a possible compensatory mechanism. Cell Cycle. 2018; 17:2637–43. https://doi.org/10.1080/15384101.2018.1553341 [PubMed]
- 30. Toiber D, Leprivier G, Rotblat B. Erratum: Long noncoding RNA: noncoding and not coded. Cell Death Discov. 2017; 3:17035. https://doi.org/10.1038/cddiscovery.2017.35 [PubMed]
- 31. Mazor G, Levin L, Picard D, Ahmadov U, Carén H, Borkhardt A, Reifenberger G, Leprivier G, Remke M, Rotblat B. The lncRNA TP73-AS1 is linked to aggressiveness in glioblastoma and promotes temozolomide resistance in glioblastoma cancer stem cells. Cell Death Dis. 2019; 10:246. https://doi.org/10.1038/s41419-019-1477-5 [PubMed]
- 32. Zhong Y, Zhao M, Yu Y, Li Q, Wang F, Wu P, Zhang W, Miao L. Prognostic value and therapeutic potential of the long noncoding RNA TP73-AS1 in cancers: A systematic review and meta-analysis. Sci Rep. 2020; 10:9053. https://doi.org/10.1038/s41598-020-65726-2 [PubMed]
- 33. Zhang B, Li Q, Wu B, Zhang S, Li L, Jin K, Li S, Li K, Wang Z, Lu Y, Xia L, Sun C. Long non-coding RNA TP73-AS1 is a potential immune related prognostic biomarker for glioma. Aging (Albany NY). 2021; 13:5638–49. https://doi.org/10.18632/aging.202490 [PubMed]
- 34. Varon M, Levy T, Mazor G, Ben David H, Marciano R, Krelin Y, Prasad M, Elkabets M, Pauck D, Ahmadov U, Picard D, Qin N, Borkhardt A, et al. The long noncoding RNA TP73-AS1 promotes tumorigenicity of medulloblastoma cells. Int J Cancer. 2019; 145:3402–13. https://doi.org/10.1002/ijc.32400 [PubMed]
- 35. Pang JC, Li KK, Lau KM, Ng YL, Wong J, Chung NY, Li HM, Chui YL, Lui VW, Chen ZP, Chan DT, Poon WS, Wang Y, et al. KIAA0495/PDAM is frequently downregulated in oligodendroglial tumors and its knockdown by siRNA induces cisplatin resistance in glioma cells. Brain Pathol. 2010; 20:1021–32. https://doi.org/10.1111/j.1750-3639.2010.00405.x [PubMed]
- 36. Marine JC, Dawson SJ, Dawson MA. Non-genetic mechanisms of therapeutic resistance in cancer. Nat Rev Cancer. 2020; 20:743–56. https://doi.org/10.1038/s41568-020-00302-4 [PubMed]
- 37. Jia B, Liu W, Gu J, Wang J, Lv W, Zhang W, Hao Q, Pang Z, Mu N, Zhang W, Guo Q. MiR-7-5p suppresses stemness and enhances temozolomide sensitivity of drug-resistant glioblastoma cells by targeting Yin Yang 1. Exp Cell Res. 2019; 375:73–81. https://doi.org/10.1016/j.yexcr.2018.12.016 [PubMed]
- 38. Stein D, Mizrahi A, Golova A, Saretzky A, Venzor AG, Slobodnik Z, Kaluski S, Einav M, Khrameeva E, Toiber D. Aging and pathological aging signatures of the brain: through the focusing lens of SIRT6. Aging (Albany NY). 2021; 13:6420–41. https://doi.org/10.18632/aging.202755 [PubMed]
- 39. Xia Y, Li K, Li J, Wang T, Gu L, Xun L. T5 exonuclease-dependent assembly offers a low-cost method for efficient cloning and site-directed mutagenesis. Nucleic Acids Res. 2019; 47:e15. https://doi.org/10.1093/nar/gky1169 [PubMed]
- 40. Marciano R, Prasad M, Ievy T, Tzadok S, Leprivier G, Elkabets M, Rotblat B. High-Throughput Screening Identified Compounds Sensitizing Tumor Cells to Glucose Starvation in Culture and VEGF Inhibitors In Vivo. Cancers (Basel). 2019; 11:156. https://doi.org/10.3390/cancers11020156 [PubMed]
- 41. Liang WS, Dunckley T, Beach TG, Grover A, Mastroeni D, Walker DG, Caselli RJ, Kukull WA, McKeel D, Morris JC, Hulette C, Schmechel D, Alexander GE, et al. Gene expression profiles in anatomically and functionally distinct regions of the normal aged human brain. Physiol Genomics. 2007; 28:311–22. https://doi.org/10.1152/physiolgenomics.00208.2006 [PubMed]
- 42. Berchtold NC, Cribbs DH, Coleman PD, Rogers J, Head E, Kim R, Beach T, Miller C, Troncoso J, Trojanowski JQ, Zielke HR, Cotman CW. Gene expression changes in the course of normal brain aging are sexually dimorphic. Proc Natl Acad Sci USA. 2008; 105:15605–10. https://doi.org/10.1073/pnas.0806883105 [PubMed]
- 43. Carithers LJ, Ardlie K, Barcus M, Branton PA, Britton A, Buia SA, Compton CC, DeLuca DS, Peter-Demchok J, Gelfand ET, Guan P, Korzeniewski GE, Lockhart NC, et al, and GTEx Consortium. A Novel Approach to High-Quality Postmortem Tissue Procurement: The GTEx Project. Biopreserv Biobank. 2015; 13:311–19. https://doi.org/10.1089/bio.2015.0032 [PubMed]
- 44. Zhang J, Bajari R, Andric D, Gerthoffert F, Lepsa A, Nahal-Bose H, Stein LD, Ferretti V. The International Cancer Genome Consortium Data Portal. Nat Biotechnol. 2019; 37:367–69. https://doi.org/10.1038/s41587-019-0055-9 [PubMed]
- 45. R2: Genomics Analysis and Visualization Platform. http://r2.amc.nl.
- 46. Vasiliou V, Pappa A, Estey T. Role of human aldehyde dehydrogenases in endobiotic and xenobiotic metabolism. Drug Metab Rev. 2004; 36:279–99. https://doi.org/10.1081/dmr-120034001 [PubMed]
- 47. Calleja LF, Yoval-Sánchez B, Hernández-Esquivel L, Gallardo-Pérez JC, Sosa-Garrocho M, Marín-Hernández Á, Jasso-Chávez R, Macías-Silva M, Salud Rodríguez-Zavala J. Activation of ALDH1A1 by omeprazole reduces cell oxidative stress damage. FEBS J. 2021. [Epub ahead of print]. https://doi.org/10.1111/febs.15698 [PubMed]
- 48. Pollard SM, Yoshikawa K, Clarke ID, Danovi D, Stricker S, Russell R, Bayani J, Head R, Lee M, Bernstein M, Squire JA, Smith A, Dirks P. Glioma stem cell lines expanded in adherent culture have tumor-specific phenotypes and are suitable for chemical and genetic screens. Cell Stem Cell. 2009; 4:568–80. https://doi.org/10.1016/j.stem.2009.03.014 [PubMed]
- 49. ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature. 2012; 489:57–74. https://doi.org/10.1038/nature11247 [PubMed]
- 50. Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM, Haussler D. The human genome browser at UCSC. Genome Res. 2002; 12:996–1006. https://doi.org/10.1101/gr.229102 [PubMed]
- 51. Davis CA, Hitz BC, Sloan CA, Chan ET, Davidson JM, Gabdank I, Hilton JA, Jain K, Baymuradov UK, Narayanan AK, Onate KC, Graham K, Miyasato SR, et al. The Encyclopedia of DNA elements (ENCODE): data portal update. Nucleic Acids Res. 2018; 46:D794–801. https://doi.org/10.1093/nar/gkx1081 [PubMed]
- 52. Landt SG, Marinov GK, Kundaje A, Kheradpour P, Pauli F, Batzoglou S, Bernstein BE, Bickel P, Brown JB, Cayting P, Chen Y, DeSalvo G, Epstein C, et al. ChIP-seq guidelines and practices of the ENCODE and modENCODE consortia. Genome Res. 2012; 22:1813–31. https://doi.org/10.1101/gr.136184.111 [PubMed]
- 53. Ho Sui SJ, Fulton DL, Arenillas DJ, Kwon AT, Wasserman WW. oPOSSUM: integrated tools for analysis of regulatory motif over-representation. Nucleic Acids Res. 2007; 35:W245–52. https://doi.org/10.1093/nar/gkm427 [PubMed]
- 54. Ho Sui SJ, Mortimer JR, Arenillas DJ, Brumm J, Walsh CJ, Kennedy BP, Wasserman WW. oPOSSUM: identification of over-represented transcription factor binding sites in co-expressed genes. Nucleic Acids Res. 2005; 33:3154–64. https://doi.org/10.1093/nar/gki624 [PubMed]
- 55. Kwon AT, Arenillas DJ, Hunt RW, Wasserman WW. oPOSSUM-3: Advanced Analysis of Regulatory Motif Over-Representation Across Genes or ChIP-Seq Datasets. G3 (Bethesda). 2012; 2:987–1002. https://doi.org/10.1534/g3.112.003202 [PubMed]
- 56. Sachamitr P, Ho JC, Ciamponi FE, Ba-Alawi W, Coutinho FJ, Guilhamon P, Kushida MM, Cavalli FM, Lee L, Rastegar N, Vu V, Sánchez-Osuna M, Coulombe-Huntington J, et al. PRMT5 inhibition disrupts splicing and stemness in glioblastoma. Nat Commun. 2021; 12:979. https://doi.org/10.1038/s41467-021-21204-5 [PubMed]
- 57. Vardarajan B, Vergote D, Tissir F, Logue M, Yang J, Daude N, Ando K, Rogaeva E, Lee J, Cheng R, Brion JP, Ghani M, Shi B, et al. Role of p73 in Alzheimer disease: lack of association in mouse models or in human cohorts. Mol Neurodegener. 2013; 8:10. https://doi.org/10.1186/1750-1326-8-10 [PubMed]
- 58. Hegi ME, Diserens AC, Gorlia T, Hamou MF, de Tribolet N, Weller M, Kros JM, Hainfellner JA, Mason W, Mariani L, Bromberg JE, Hau P, Mirimanoff RO, et al. MGMT gene silencing and benefit from temozolomide in glioblastoma. N Engl J Med. 2005; 352:997–1003. https://doi.org/10.1056/NEJMoa043331 [PubMed]
- 59. Sturm D, Witt H, Hovestadt V, Khuong-Quang DA, Jones DT, Konermann C, Pfaff E, Tönjes M, Sill M, Bender S, Kool M, Zapatka M, Becker N, et al. Hotspot mutations in H3F3A and IDH1 define distinct epigenetic and biological subgroups of glioblastoma. Cancer Cell. 2012; 22:425–37. https://doi.org/10.1016/j.ccr.2012.08.024 [PubMed]
- 60. Meliala IT, Hosea R, Kasim V, Wu S. The biological implications of Yin Yang 1 in the hallmarks of cancer. Theranostics. 2020; 10:4183–200. https://doi.org/10.7150/thno.43481 [PubMed]
- 61. Riggs KJ, Saleque S, Wong KK, Merrell KT, Lee JS, Shi Y, Calame K. Yin-yang 1 activates the c-myc promoter. Mol Cell Biol. 1993; 13:7487–95. https://doi.org/10.1128/mcb.13.12.7487-7495.1993 [PubMed]
- 62. Grönroos E, Terentiev AA, Punga T, Ericsson J. YY1 inhibits the activation of the p53 tumor suppressor in response to genotoxic stress. Proc Natl Acad Sci USA. 2004; 101:12165–70. https://doi.org/10.1073/pnas.0402283101 [PubMed]
- 63. Verheul TC, van Hijfte L, Perenthaler E, Barakat TS. The Why of YY1: Mechanisms of Transcriptional Regulation by Yin Yang 1. Front Cell Dev Biol. 2020; 8:592164. https://doi.org/10.3389/fcell.2020.592164 [PubMed]
- 64. Morales-Martinez M, Valencia-Hipolito A, Vega GG, Neri N, Nambo MJ, Alvarado I, Cuadra I, Duran-Padilla MA, Martinez-Maza O, Huerta-Yepez S, Vega MI. Regulation of Krüppel-Like Factor 4 (KLF4) expression through the transcription factor Yin-Yang 1 (YY1) in non-Hodgkin B-cell lymphoma. Oncotarget. 2019; 10:2173–88. https://doi.org/10.18632/oncotarget.26745 [PubMed]
- 65. Kaufhold S, Garbán H, Bonavida B. Yin Yang 1 is associated with cancer stem cell transcription factors (SOX2, OCT4, BMI1) and clinical implication. J Exp Clin Cancer Res. 2016; 35:84. https://doi.org/10.1186/s13046-016-0359-2 [PubMed]
- 66. Ye Z, Li G, Kim C, Hu B, Jadhav RR, Weyand CM, Goronzy JJ. Regulation of miR-181a expression in T cell aging. Nat Commun. 2018; 9:3060. https://doi.org/10.1038/s41467-018-05552-3 [PubMed]
- 67. Aubry S, Shin W, Crary JF, Lefort R, Qureshi YH, Lefebvre C, Califano A, Shelanski ML. Assembly and interrogation of Alzheimer’s disease genetic networks reveal novel regulators of progression. PLoS One. 2015; 10:e0120352. https://doi.org/10.1371/journal.pone.0120352 [PubMed]
- 68. Leprivier G, Rotblat B, Khan D, Jan E, Sorensen PH. Stress-mediated translational control in cancer cells. Biochim Biophys Acta. 2015; 1849:845–60. https://doi.org/10.1016/j.bbagrm.2014.11.002 [PubMed]
- 69. Ramkumar C, Cui H, Kong Y, Jones SN, Gerstein RM, Zhang H. Smurf2 suppresses B-cell proliferation and lymphomagenesis by mediating ubiquitination and degradation of YY1. Nat Commun. 2013; 4:2598. https://doi.org/10.1038/ncomms3598 [PubMed]
- 70. Wang GZ, Goff SP. Regulation of Yin Yang 1 by Tyrosine Phosphorylation. J Biol Chem. 2015; 290:21890–900. https://doi.org/10.1074/jbc.M115.660621 [PubMed]



