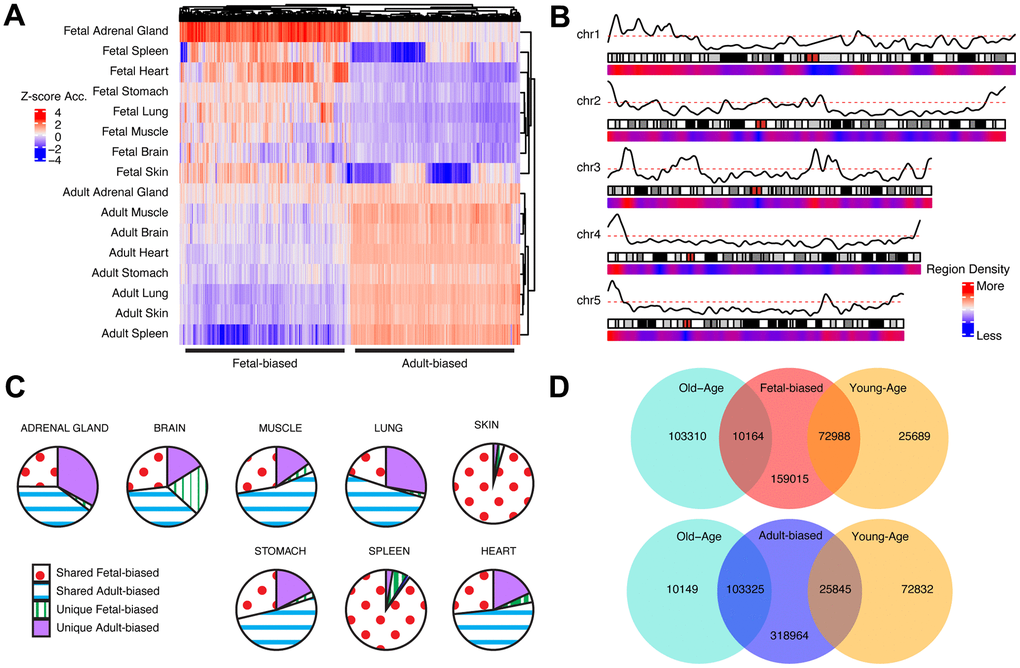
Figure 1.Cross-tissue accessibility. (A) Representative heatmap of Dnase-I accessibility for regions significantly different between fetal/adult tissues. Color scale indicates magnitude of chromatin accessibility signal (see Supplementary Methods). Horizontal lines denote defined fetal-biased (left) and adult-biased regions. (B) Genomic distribution of regions changing accessibility in fetal and adult comparison. Red/blue: density of defined differentially-accessible regions. Solid black line: relative proportion of regions more accessible in adult (top) or fetal (bottom) tissues. First five autosomes shown (see Supplementary Figure 2). (C) The proportion of defined altered-accessibility regions between adult and fetal samples for indicated tissues which are unique to that tissue, or captured in the pan-tissue set. (D) Overlaps between regions defined as differentially-accessible in fetal/adult comparison and those defined in the young/old-age comparison. Directionality in accessibility change is significantly shared (see Supplementary Table 1). Related content can be found in Supplementary Information, Supplementary Figures 1–6 and Supplementary Tables 1, 2.