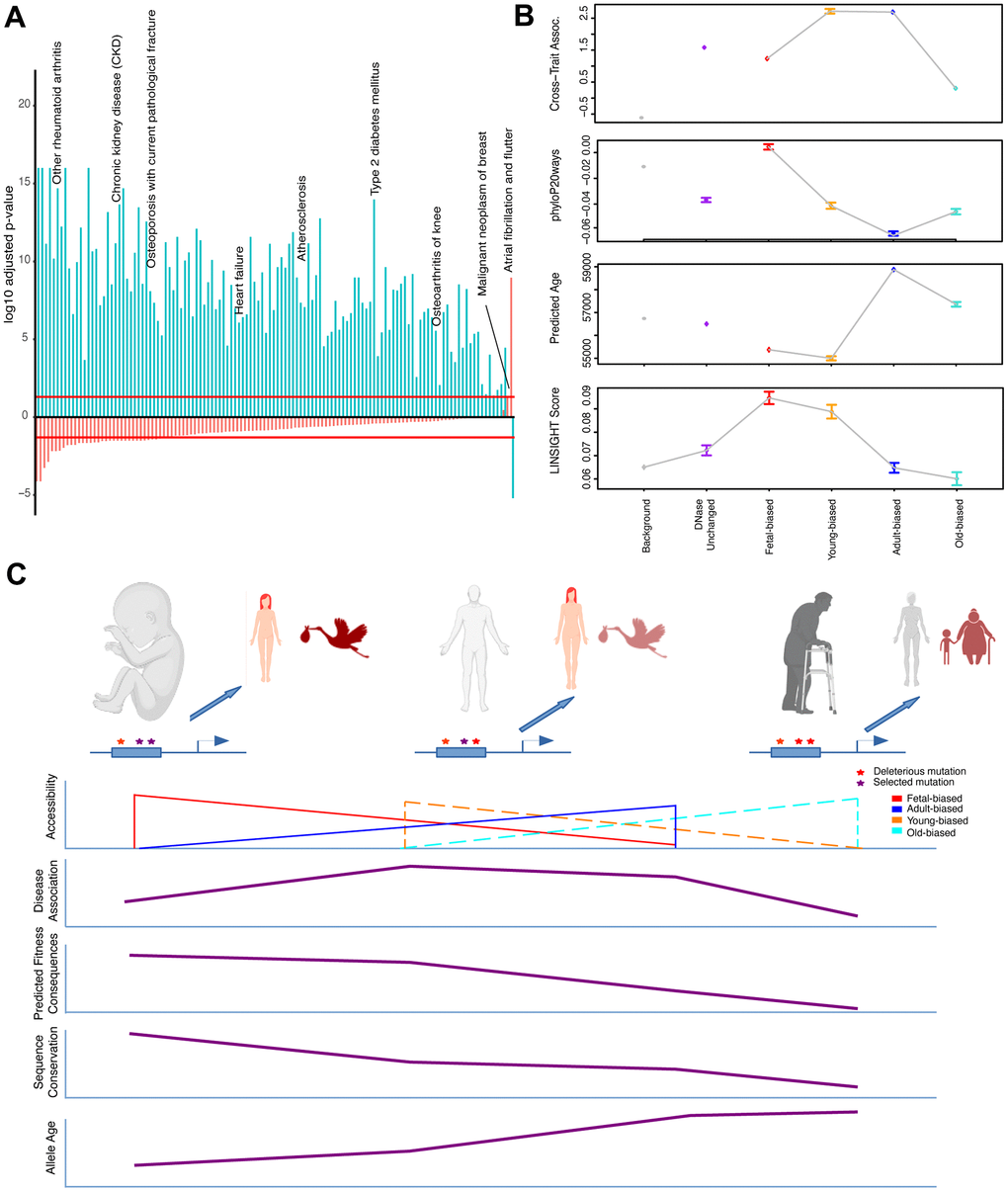
Figure 3.Epigenetic context and heritable disease associations. (A) Adjusted p-values for hypergeometric tests showing enrichment/depletion (positive/negative) for GWAS variants nearby regions increasing (blue) or decreasing (red) accessibility across adult tissues for a number of age-associated diseases (see Supplementary Table 4). (B) Cross-set disease associations, and additional per-SNP metrics, for variants nearby developmental and age-altered region sets along with unaltered DNase sites and variants not nearby accessible regions. See Supplementary Table 4 and Methods. (C) Model for the effects of epigenetic context on disease association and sequence evolution. (Top): Example enhancer elements more accessible in fetal, adult, and old-adult tissues (left-right) which have been modified by mutations. (Left): Deleterious mutations disrupting regulation in development stand to have the biggest impact on fitness, while having a moderate effect on tissue homeostasis. (Middle): Mutations disrupting regulation in young-adult tissues have a moderate impact on fitness, but a larger effect on tissue homeostasis (particularly over adulthood). (Right): Mutations disrupting regulation in old-adult tissues have weak impacts on fitness, and a weak effect on tissue homeostasis (which has already deteriorated with age). (Bottom): Illustrating patterns of accessibility, disease association, sequence constraint and variant allele age for these sets of regions changing accessibility over time.