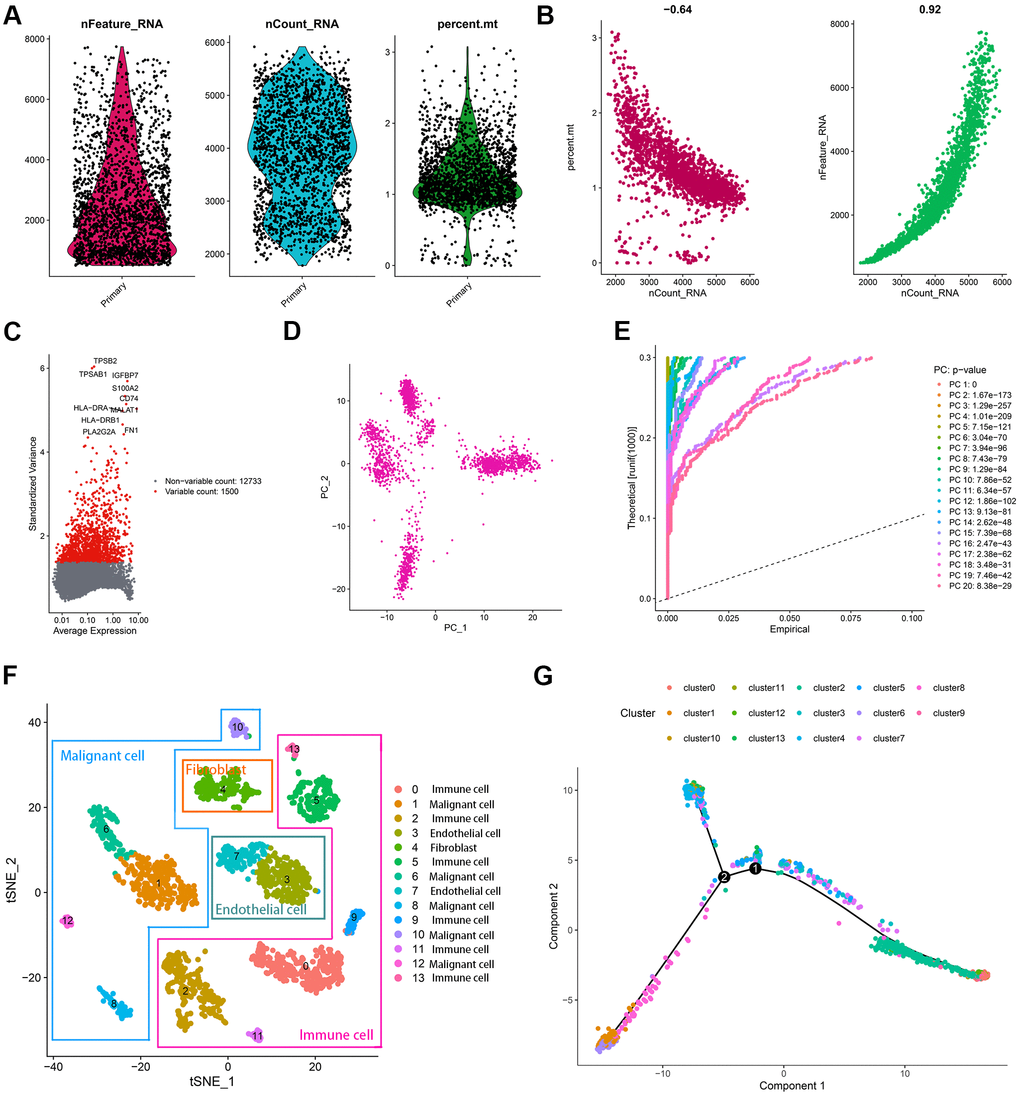
Figure 2.Characterization of scRNA-seq from 2075 cells. (A) Quality control plots of cell samples. (B) The sequencing depth was negatively correlated with the proportion of mitochondrial genes and positively associated with detected gene numbers. (C) 1500 variable genes across cell samples were identified. (D, E) PCA was conducted to reduce the dimension of data sets. (F) Cell samples were classified into 14 clusters with the t-SNE algorithm. (G) The trajectory analysis of 14 cell clusters.