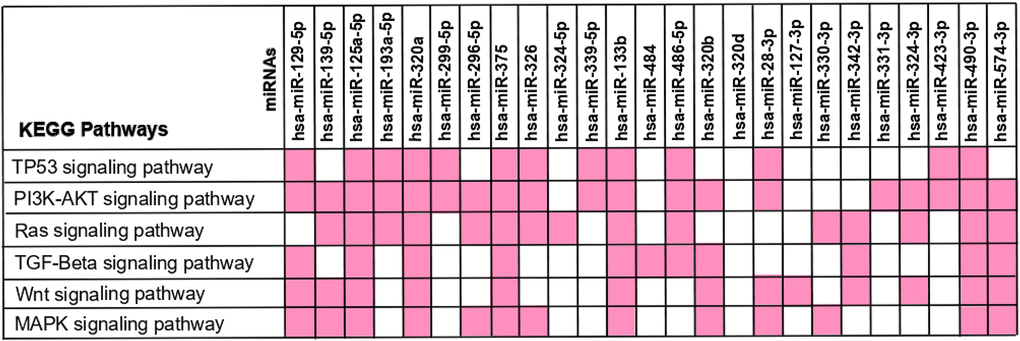
Figure 6.KEGG pathway analysis – interaction between the 25 differently expressed miRNAs and the frequently altered pathways in colorectal cancer. The 173 miRNA-target genes were used to perform the enrichment pathway analysis using KEEG at DAVID database. The miRNA-target genes found to be involved in CRC were crossed with the genes intervening in each signalling pathway in order to identify the miRNAs affecting each pathway. The miRNAs interaction with each KEGG Pathway is reported in pink.