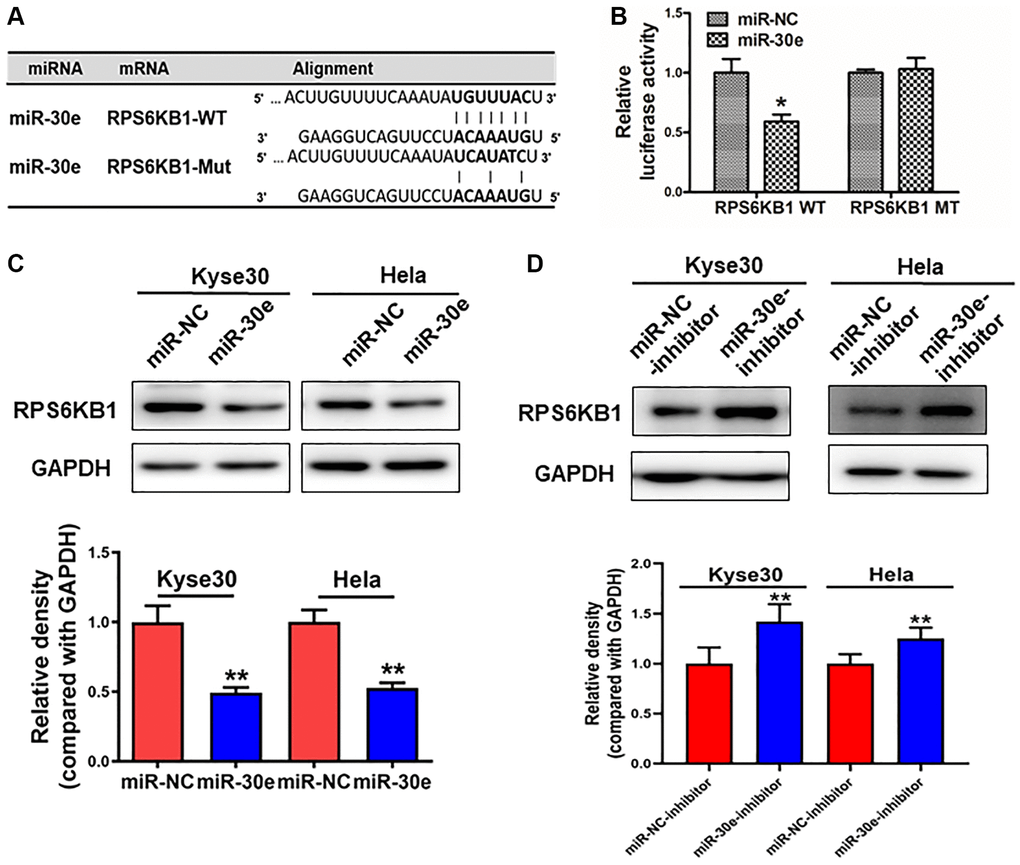
Figure 3.RPS6KB1 was a novel target of miR-30e. (A) The miR-30e binding site predicted in the 3′-UTR of RPS6KB1 mRNA. Mutant construct was generated at the seed sequence region of RPS6KB1 3′-UTR as indicated. The 3′-UTR fragment of RPS6KB1 mRNA containing wild-type (WT) or mutant (MT) of the miR-30e binding site was cloned into pmirGLO vector. (B) Kyse30 cells were transfected with pmirGLO reporter vector containing wild-type or mutant RPS6KB1 3′-UTR (indicated as RPS6KB1-WT and RPS6KB1-MT) with the combination of either miR-NC or miR-30e mimic. Luciferase activities were determined 24 h after the co-transfection. (C, D) The protein levels of RPS6KB1 in Kyse30 and Hela cells were analyzed by Western blotting assay. The density levels of RPS6KB1 were determined by ImageJ software, and relative fold changes were obtained by the ratios of RPS6KB1 to GAPDH levels. Data represent mean ±SD. of 3 replicates. *,**indicates significant difference at P < 0.05, and at P < 0.01, respectively.