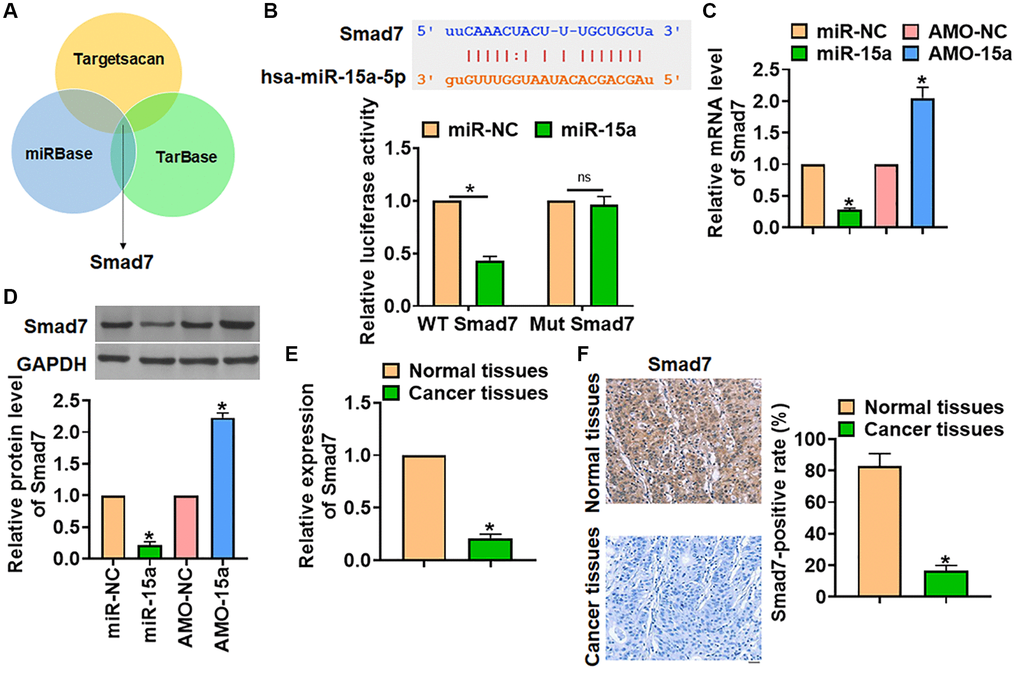
Figure 3.Smad7 was the target of miR-15a in glioma progression. (A) The miRBase, Targetscan and Tarbase were used to search the predicted target of miR-15a, and Smad7 has the has the greatest potential. (B) The binding bases between miR-15a and Smad7 (upper), and luciferase assay used to test the activity of WT Smad7 and Mut Smad7 (lower) in HEK193 cells. SHG139 cells were transfected with miR-15a or AMO-15a, and the mRNA (C) and protein (D) level of Smad7 was explored. (E) The mRNA expression of Smad7 in in adjacent normal tissues and cancer tissues of glioma was determined. (F) IHC staining used to examine the expression of Smad7 in different grades of glioma tissues, and relative Smad7 positive area was calculated. Scale bar, 30 μm. Data were expressed as mean ± SD.*P < 0.05.