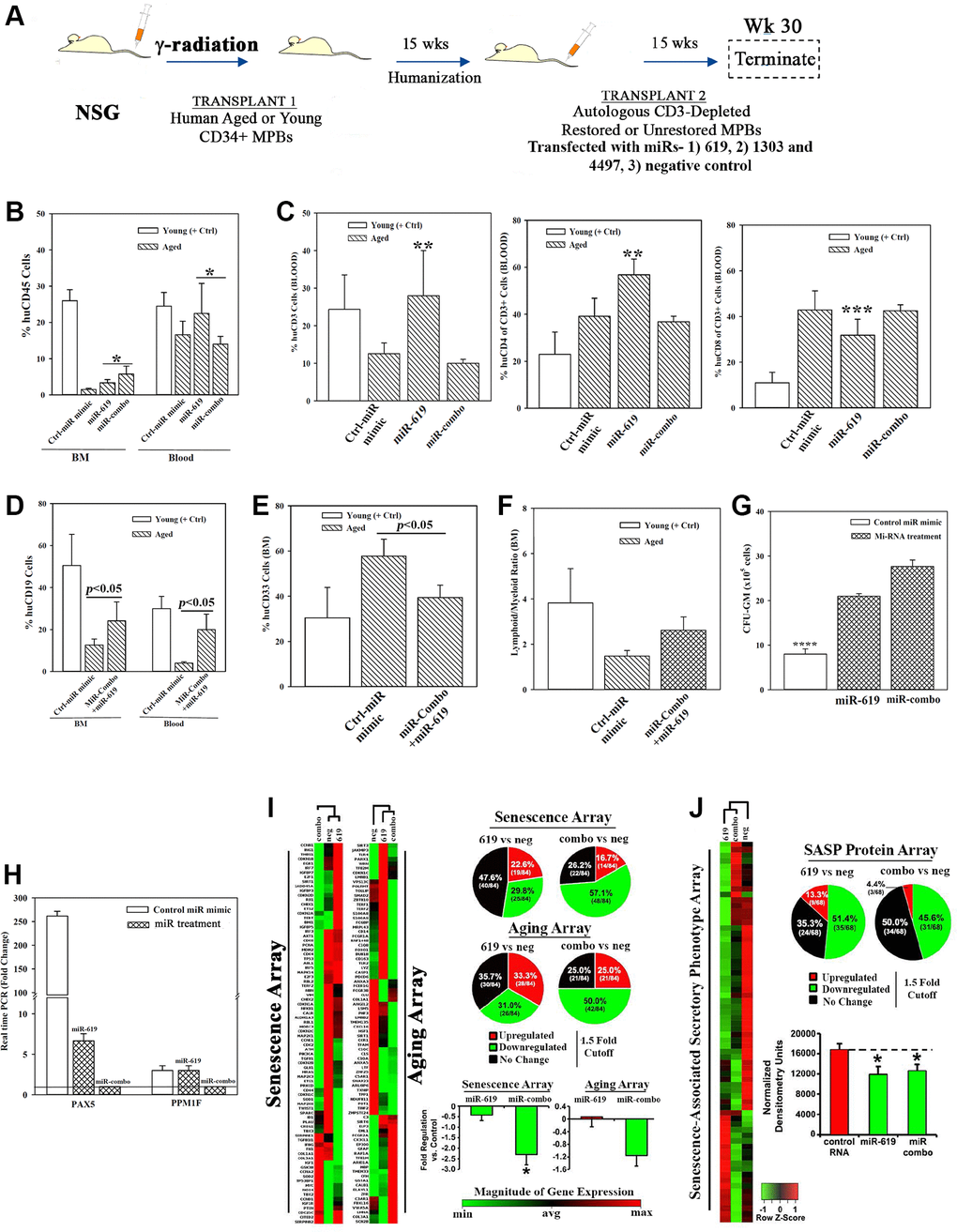
Figure 5.Hematopoietic restoration with miRNAs. (A) Restoration protocol with NSG was similar to Figure 2H. Chimeric mice were given autologous CD3-depleted aged MPBs, transfected with miR-619, miR-combo, −619, −1303 and −4497, or control (RNA mimic) and then cultured for 7 days, (n = 18). (B–E) Mice were analyzed for huCD45 in BM and blood (B); huCD3, CD4 and CD8 in blood (C); HuCD19 in blood and BM (D); HuCD33 in BM (E). Results presented as % mean cells ± SD (F) Lymphoid:Myeloid ratio (CD3++CD19+/CD33+) in BM. (G) CFU-GM with huCD45+ cells from femurs, mean ± SD. (H) qPCR for PAX5 and PPM1F with RNA from huCD45+cells from femurs. Fold change ± SD used 1 for the lowest value. (I, J) RNA from `H’ were analyzed in qPCR human senescence and aging arrays. Normalized results used 1.5-fold cutoff to classify up- or down-regulation, or no change (I). SASF analyses with plasma. Semi-quantitative densitometry used 1.5-fold cutoff, similar to I (J). Differential gene and protein expressions as heatmaps (left), pie charts (top) and bar graph (bottom), mean ± SD. *p ≤ 0.05 vs. control. See also Supplementary Figure 5.